Design
A randomized controlled cell experiment.
Time and settings
The experiment was completed at the Central Laboratory, the Affiliated Hospital of Qingdao University, China from August 2014 to June 2015.
Materials
HUMSCs were obtained from the National Research Center for Stem Cell Engineering and Technology, China.
Methods
Group Information
HUMSCs were cultured in DMEM/F12 (1:1) with 10% fetal bovine serum, 100 U/mL penicillin at 37 ℃ in a humidified atmosphere containing 5% CO2. The cells at passages 5-6 were used for the subsequent assays. HUMSCs cultured in osteogenic medium
(50 mg/L ascorbate-2 phosphate, 10-8 mol/L dexamethasone, 10 mmol/L β-glycerophosphate). Insulin (10-7 mmol/L) was added to the culture medium as the experimental group. The cells in the osteogenic medium without any treatment were used as controls. The medium was changed every 2 days.
Cell proliferation assay
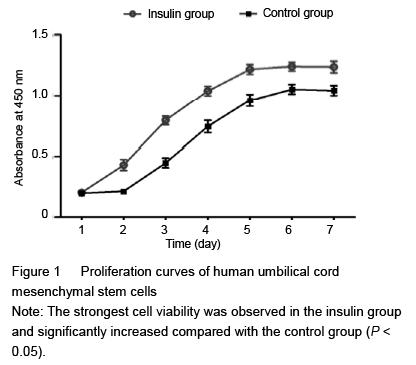
Cell counting kit-8 (CCK-8) was employed in this experiment to quantitatively evaluate HUMSC viability. Briefly, approximately 1×104 HUMSCs were seeded on each film placed in the 24-well plates for 12 hours, then the growth medium was replaced by 200 mL of osteogenic medium with insulin
(10-7 mmol/L) in the experimental group, and the medium in the control group was replaced only by 200 mL of osteogenic medium at 37 ℃ in a humidified atmosphere containing 5% CO2. Cells were harvested daily (6 wells/group) and enumerated for 7 consecutive days. Culture medium was removed every day and the cells were washed with PBS twice. Approximately 900 μL serum-free DMEM/F12 medium and 100 μL CCK-8 solutions were added, followed by incubation for 3 hours at
37 ℃. Supernatant was transferred to 96-well plates, and the absorbance value at 450 nm was determined using a microplate reader. Six parallel experiments in each sample were used to assess the cell viability.
Assay for the alkaline phosphatase (ALP) activity in HUMSCs
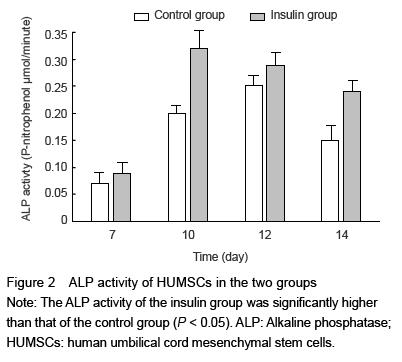
The ALP activity in HUMSCs was assessed at 7, 10, 12 and 14 days, using alkaline phosphatase activity colorimetric assay kit. The amount of p-nitrophenol was measured at 405 nm by STAKMAX™ microplate reader. Protein concentrations were measured using the Bio Rad protein assay kit with bovine serum albumin as a standard. The experiments were repeated three times.
Immunocytochemical staining of type I collagen protein
Following trypsinization, HUMSCs were seeded onto glass coverslips with a density of 5×103 cells per well in DMEM/F12 (1:1) with 10% fetal bovine serum, 100 U/mL penicillin at 37 ℃ in a humidified atmosphere containing 5% CO2. When 50%-70% confluence was reached, the medium was changed to the osteogenic medium with insulin or with no insulin. The medium was changed every
2 days. After 12 days, cells seeded on glass coverslips were washed with PBS and fixed with 4% formaldehyde (Unifix) for 15 minutes at room temperature. After fixation and washing in
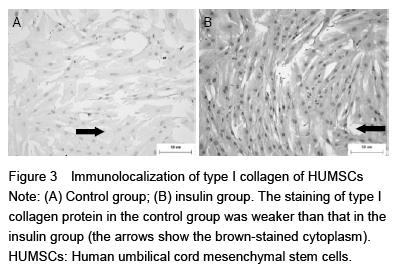
0.01 mol/L PBS (pH 7.2), cells were permeabilized when needed with 0.05% Triton X-100 in PBS for 30 minutes at 4 ℃ prior to labeling and then washed again with PBS. Nonspecific binding sites were subsequently blocked with 3% normal goat serum (Dako Cytomation) in PBS for 20 minutes. After washing, cells were incubated with rabbit polyclonal antibody against type I collagen (diluted concentration of 1:100) overnight at 4 ℃. The cells were then washed again and incubated for 30 minutes with goat anti-rabbit horseradish peroxidase-conjugated secondary antibodies. After washing in PBS, a high-sensitivity diaminobenzidine chromogenic substrate system (DAB) and Mayer’s hematoxylin were respectively used to visualize the peroxidase and for counter staining. After mounting in an aqueous mounting medium (Aquatex; Merck, Darmstadt, Germany), cell-seeded coverslips were examined using a photomicroscope equipped with an automated camera.
Real-time PCR
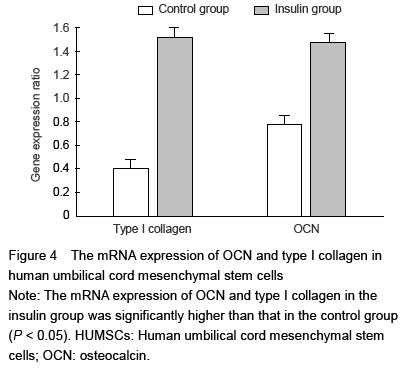
To assess the osteogenesis of HUMSCs, type I collagen protein and osteocalcin mRNA expressions were detected by real-time PCR. Total RNA was extracted from cultured HUMSCs with Trizol. The separation and precipitation of RNA were accomplished with chloroform and isopropyl alcohol. Total RNA (1 μg) was converted to cDNA and amplification was performed using One-Step qPCR Kit following the manufacturer’s protocol. For determination of type I collagen and osteocalcin mRNA expression, SYBR Green detection was used and the values were normalized using glyceraldehyde-3-phosphate dehydrogenase (GAPDH). Real-time quantitative PCR was performed in a 7300 Sequence Detection System (Applied Biosystems). Primers pairs used in this study are presented in Table 1.
.jpg)
The thermal cycling conditions for the real-time PCR were 95 ℃ for 2 minutes followed by 45 cycles of
94 ℃ for 15 seconds, 58 ℃ for 20 seconds and
72 ℃ for 30 seconds. Data were collected at 68 ℃. Relative expression levels for each primer set were normalized to the expression of GAPDH by the 2-??Ct methods[17].
Main outcome measures
(1) The proliferative capacity of HUMSCs; (2) ALP activity; (3) type I collagen protein expression; (4) the gene expression of type I collagen and osteocalcin.
Statistical analysis
All data were analyzed statistically using SPSS 21.0 statistical software. Each experiment was repeated at least three times, and representative data were reported. Continuous variables are expressed as mean±SD. Independent sample t-test was used to compare the continuous variables if these data normally distributed, which was confirmed with Kolmogorov-Smirnov test. The Wilcoxon signed-rank test was used to assess non-parametric data. A P-value of less than 0.05 was considered statistically significant.
.jpg)
.jpg)