中国组织工程研究 ›› 2022, Vol. 26 ›› Issue (1): 76-83.doi: 10.12307/2022.013
• 肿瘤干细胞 cancer stem cells • 上一篇 下一篇
CD44+膀胱癌干细胞中角蛋白6B的表达和调控
刘 璐,杨宇明,商艾晨,刘 翠,孙 策,王 静
- 天津医科大学中新生态城医院病理科,天津市 300456
-
收稿日期:2020-11-28修回日期:2020-12-08接受日期:2021-01-06出版日期:2022-01-08发布日期:2021-10-25 -
通讯作者:杨宇明,博士,副教授,天津医科大学中新生态城医院病理科,天津市 300456 -
作者简介:刘璐,女,1995年生,陕西省渭南市人,汉族,天津医科大学在读硕士。 -
基金资助:滨海新区卫生和计划生育委员会科技项目(2017BWKY033),项目负责人:杨宇明
Expression and regulation of keratin 6B in CD44+ bladder cancer stem cells
Liu Lu, Yang Yuming, Shang Aichen, Liu Cui, Sun Ce, Wang Jing
- Department of Pathology, Sino-Singapore Eco-City Hospital of Tianjin Medical University, Tianjin 300456, China
-
Received:2020-11-28Revised:2020-12-08Accepted:2021-01-06Online:2022-01-08Published:2021-10-25 -
Contact:Yang Yuming, MD, Associate professor, Department of Pathology, Sino-Singapore Eco-City Hospital of Tianjin Medical University, Tianjin 300456, China -
About author:Liu Lu, Master candidate, Department of Pathology, Sino-Singapore Eco-City Hospital of Tianjin Medical University, Tianjin 300456, China -
Supported by:the Science and Technology Project of Tianjin Binhai New District Health and Family Planning Commission, No. 2017BWKY033 (to YYM)
摘要:
文题释义:
角蛋白6B:角蛋白6B基因位于染色体12q13,由9个外显子组成,其编码的蛋白是Ⅱ型角蛋白家族中的一员,属于高分子量角蛋白。CD44:是一种跨膜糖蛋白,其基因位于11号染色体上,转录后经过选择性剪切形成多个异构体,其中包括CD44v6。CD44在多种恶性肿瘤中异常表达,尤其在肿瘤干细胞中表达升高。
背景:膀胱癌干细胞参与促进膀胱癌的术后复发及耐药性,而多项研究表明角蛋白6B参与多个不同类型肿瘤的发生和进展,并与肿瘤的预后密切相关。
目的:观察角蛋白6B在CD44+膀胱癌干细胞中的表达差异及其对膀胱癌干细胞增殖、迁移及自我更新的影响,并进一步探讨角蛋白6B表达对膀胱癌患者预后的影响。
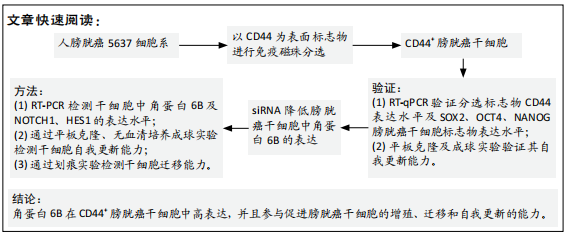
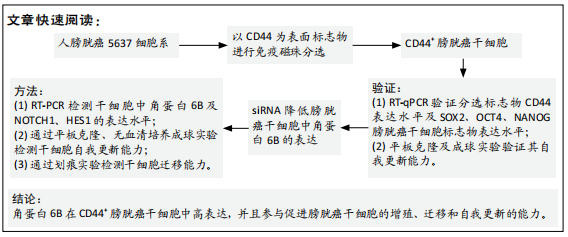
方法:①使用免疫磁珠分选法分选出人CD44+ 5637膀胱癌干细胞,使用RT-PCR检测分选后的相关干性基因SOX2,OCT4和NANOG的表达,采用体外肿瘤球形成实验进一步验证CD44+细胞具有肿瘤干细胞的自我更新能力,RT-PCR检测CD44+膀胱癌干细胞中的角蛋白6B的表达情况;②将CD44+ 5637膀胱癌干细胞分为2组,其中角蛋白6B沉默组将角蛋白6B小干扰RNA转染至分选后的CD44+膀胱癌干细胞,未转染的CD44+膀胱癌干细胞作为空白对照组,转染2 d后收集细胞,分别用平板克隆实验、划痕实验和体外肿瘤球形成实验检测角蛋白6B沉默后膀胱癌干细胞的增殖,迁移和自我更新能力;③采用免疫组织化学染色检测膀胱癌患者24例膀胱肿瘤组织切片的角蛋白6B和CD44v6的表达;④利用ONCOMINE数据库比较膀胱肿瘤组织中角蛋白6B的表达情况对患者总生存率的影响。
结果与结论:①分选后的CD44+细胞中相关干性基因(SOX2,OCT4,NANOG)和角蛋白6B表达高于CD44-细胞(P < 0.05),其细胞增殖,迁移和体外成球能力显著升高(P < 0.05),角蛋白6B小干扰RNA可下调角蛋白6B在CD44+膀胱癌干细胞中的表达(P < 0.05);②与空白对照组相比,角蛋白6B小干扰RNA转染后的CD44+膀胱癌干细胞增殖和迁移能力显著降低(P < 0.05),肿瘤球的数量明显降低(P < 0.05),而Notch1和Hes1 mRNA表达升高(P < 0.05);③角蛋白6B与CD44v6在患者膀胱癌组织中的表达差异有显著性意义(P=0.006),其中角蛋白6B高表达的膀胱癌患者总生存率低于角蛋白6B低表达的患者;④上述数据证实,角蛋白6B可在CD44+膀胱癌干细胞中可高度表达,并且促进膀胱癌干细胞的增殖、迁移和自我更新,角蛋白6B高表达有助于膀胱癌患者生存情况的改善。
https://orcid.org/0000-0002-1365-3073 (Yang Yuming)
中国组织工程研究杂志出版内容重点:干细胞;骨髓干细胞;造血干细胞;脂肪干细胞;肿瘤干细胞;胚胎干细胞;脐带脐血干细胞;干细胞诱导;干细胞分化;组织工程
中图分类号:
引用本文
刘 璐, 杨宇明, 商艾晨, 刘 翠, 孙 策, 王 静. CD44+膀胱癌干细胞中角蛋白6B的表达和调控[J]. 中国组织工程研究, 2022, 26(1): 76-83.
Liu Lu, Yang Yuming, Shang Aichen, Liu Cui, Sun Ce, Wang Jing. Expression and regulation of keratin 6B in CD44+ bladder cancer stem cells[J]. Chinese Journal of Tissue Engineering Research, 2022, 26(1): 76-83.
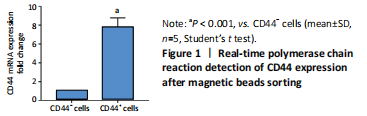
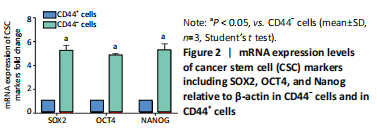
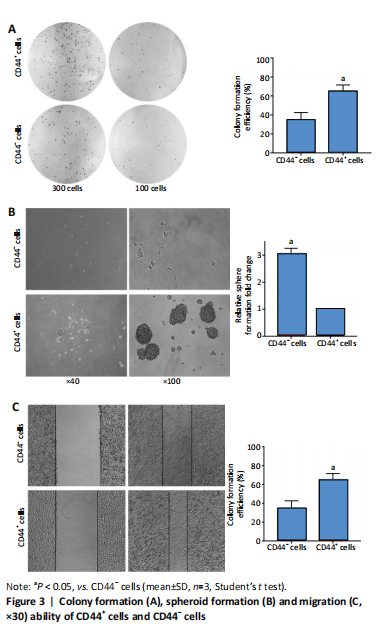
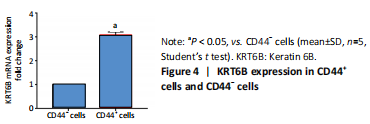
Initially, the cell suspension was prepared by digesting the 10 bladder cancer samples. The CD44v6+EMA- bladder cancer stem cells were isolated by magnetic bead cell sorting. The results of transcriptome sequencing compared the expression profiles of the bladder cancer genes in the CD44v6+EMA- BCSCs, control subjects, and the KRT6B expression levels in BCSCs were significantly higher. To explore the expression of KRT6B in bladder cancer cell lines, the CD44+ cells were isolated by magnetic bead cell sorting from 5637 cell line. RT-PCR assay was used for detection of CD44 expression after magnetic sorting (P < 0.05; Figure 1). RT-PCR results showed that CD44+ cells expressed higher level of CSCs-related genes (Sox2, Oct4, and Nanog; P < 0.05; Figure 2). In the next step, colony formation assays indicated a higher proliferation and colony formation ability of CD44+ cells (P < 0.05; Figure 3A). Sphere formation assays showed that CD44+ BCSCs could form larger and more spheres than CD44- bladder cancer cells (P < 0.05; Figure 3B). It is demonstrated that CD44+ cells possessed enhanced self-renewal ability. Wound healing assays showed that the migration capacity of CD44+ cells was higher compared with the control group (P < 0.05; Figure 3C). A series of experiments showed that the CD44+ cells possessed the characteristic of BCSC populations, including self-renewal and spheroid formation capacity. Subsequently, the expression of KRT6B in CD44+ BCSCs and CD44- bladder cancer cells was analyzed. RT-PCR indicated that KRT6B was enriched in CD44+ cells of 5637 human bladder cancer cell line (P < 0.05; Figure 4). These results indicated that the KRT6B expression trend remained consistent between RNA sequencing and RT-PCR.

Silencing KRT6B in CD44+ BCSCs inhibited spheroid formation ability (in vitro)
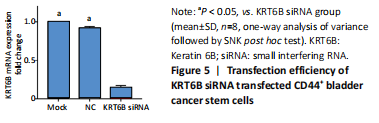
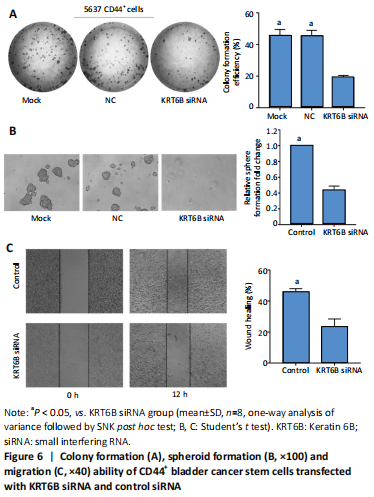
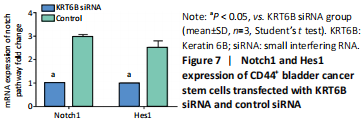
We explored the role of KRT6B in the CD44+ bladder cancer stem-like properties by silencing KRT6B in CD44+ BCSCs. KRT6B was knockdown by siRNA in CD44+ BCSCs and the downregulation of KRT6B was confirmed by RT-PCR (P < 0.05; Figure 5). Colony formation assays showed that KRT6B knockdown in CD44+ BCSCs inhibited proliferation and colony formation of CD44+ BCSCs (P < 0.05; Figure 6A). Sphere formation assay indicated that silence of KRT6B expression decreased the spheroid formation and self-renewal ability in CD44+ BCSCs (P < 0.05; Figure 6B). Wound healing assay displayed that ability of migration in KRT6B siRNA cells was decreased compared with the control group (P < 0.05; Figure 6C). This suggested that the downregulated KRT6B inhibited cell self-renewal and spheroid formation ability in CD44+ bladder cancer stem cells. To further explore the effect on a cellular level, we assessed the effect of KRT6B down-regulation in CD44- cells. Next, we aimed to initially explore the potential mechanism by which KRT6B maintains stemness in BCSCs. The RT-PCR analysis demonstrated that the silence of KRT6B in CD44+ cells increased the expression of NOTCH1 and HES1 in mRNA level (P < 0.05; Figure 7). Collectively, we speculated that KRT6B maintained the CD44+ BCSCs stemness which might be related to the Notch signal pathway.

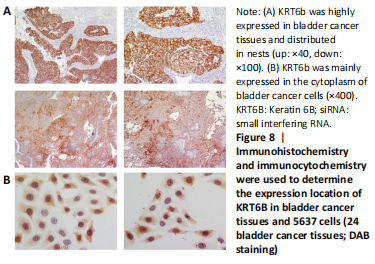
Expression of KRT6B in immunohistochemistry and immunocytochemistry (in vitro)
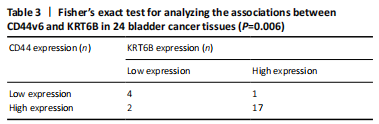
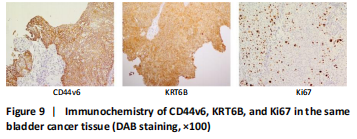
Immunohistochemistry was performed in 24 bladder cancer tissues to analyze the expression of CD44v6 and KRT6B. Immunohistochemical analysis showed that the cells with KRT6B expression located in the basal and intermediate layer of normal bladder uroepithelium. During uroepithelium carcinoma differentiation, immunohistochemical analysis indicated that basal cells expressed KRT6B and KRT6B+ cells distributed in nest-pattern in bladder cancer (Figure 8A). Immunocytochemistry analysis showed that KRT6B was mainly located in the cytoplasm and expressed in different subsets of the bladder cancer cells (Figure 9B). It is proposed that CD44v6+ cells located in the basal layer of bladder cancer. Immunohistochemical analysis confirmed co-expression of CD44v6 and KRT6B defined basal layer cells in bladder cancer and expanded from basal towards differentiated layer. Furthermore, it was observed that KRT6B expression was correlated with CD44v6 expression in bladder cancer specimens (n=24, P=0.006, Bilateral, Fisher’s exact test; Table 3). In summary, these results showed that the expression of KRT6B was correlated with CD44v6 and KRT6B+ cells, which might be related to CD44+ BCSCs.

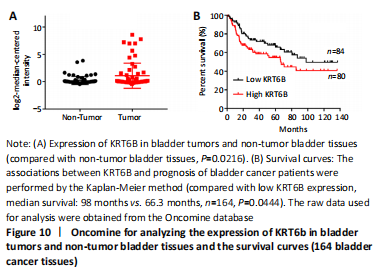
Expression and prognosis of KRT6B in normal bladder tissue and bladder cancer (in vitro)
To further explore the effect of KRT6B expression on the clinical outcome of bladder cancer patients, we used the Lee database from ONCOMINE aimed at mining bladder gene information. We analyzed the expression of KRT6B at the protein level by GraphPad Prism 7.0 statistical software. The result displayed that the expression of KRT6B in bladder cancer tissues was significantly increased compared with normal bladder tissues (P=0.022; Figure 10A). Subsequently, we evaluated the possible association between KRT6B expression and the prognostic role of KRT6B in human bladder cancer tissues. The result showed that bladder cancer patients with high KRT6B expression had a shorter overall survival rate than that of patients with low KRT6B expression (median survival=66.3 months vs. 98 months, n=164, P=0.044; Figure 10B). Therefore, these results indicate that the high expression of KRT6B is related to the poor prognosis of bladder cancer patients.
| [1] BAJAJ J, DIAZ E, REYA T. Stem cells in cancer initiation and progression. J Cell Biol. 2020;219(1):e201911053. [2] AZZARELLI R. Organoid models of glioblastoma to study brain tumor stem cells. Front Cell Dev Biol. 2020;8:220. [3] JAHANAFROOZ Z, MOSAFER J, AKBARI M, et al. Colon cancer therapy by focusing on colon cancer stem cells and their tumor microenvironment. J Cell Physiol. 2020;235(5):4153-4166. [4] XIAOBIN Z, CHENGXIAO L, JIE Y, et al. Breast cancer stem cells, heterogeneity, targeting therapies and therapeutic implications. Pharmacol Res. 2020;163: 105320. [5] COLES GL, CRISTEA S, WEBBER JT, et al. Unbiased proteomic profiling uncovers a targetable GNAS/PKA/PP2A axis in small cell lung cancer stem cells. Cancer Cell. 2020;38(1):129-143. [6] BARBATO L, BOCCHETTI M, DI BIASE A, et al. Cancer stem cells and targeting strategies. Cells. 2019;8(8):926. [7] AFIFY SM, CALLE AS, HASSAN G, et al. A novel model of liver cancer stem cells developed from induced pluripotent stem cells. Br J Cancer. 2020; 122(9):1378-1390. [8] O’CONOR CJ, CHEN T, GONZÁLEZ I, et al. Cancer stem cells in triple-negative breast cancer: a potential target and prognostic marker. Biomark Med. 2018;12(7):813-820. [9] GZIL A, ZARĘBSKA I, BURSIEWICZ W, et al. Markers of pancreatic cancer stem cells and their clinical and therapeutic implications. Mol Biol Rep. 2019;46(6):6629-6645. [10] NAGARE RP, SNEHA S, KRISHNAPRIYA S, et al. ALDH1A1+ ovarian cancer stem cells co-expressing surface markers CD24, EPHA1 and CD9 form tumours in vivo. Exp Cell Res. 2020;392(1):112009. [11] ALMANAA TN, GEUSZ ME, JAMASBI RJ. A new method for identifying stem-like cells in esophageal cancer cell lines. J Cancer. 2013;4(7):536. [12] HUNTLY BJP, GILLILAND DG. Leukaemia stem cells and the evolution of cancer-stem-cell research. Nat Rev Cancer. 2005;5(4):311-321. [13] ZHU J, HUANG G, HUA X, et al. CD44s is a crucial ATG7 downstream regulator for stem-like property, invasion, and lung metastasis of human bladder cancer (BC) cells. Oncogene. 2019;38(17):3301-3315. [14] CHAN KS, ESPINOSA I, CHAO M, et al. Identification, molecular characterization, clinical prognosis, and therapeutic targeting of human bladder tumor-initiating cells. Proc Natl Acad Sci U S A. 2009;106(33): 14016-14021. [15] TYNER AL, EICHMAN MJ, FUCHS E. The sequence of a type II keratin gene expressed in human skin: conservation of structure among all intermediate filament genes. Proc Natl Acad Sci U S A. 1985;82(14):4683-4687. [16] FUCHS EV, COPPOCK SM, GREEN H, et al. Two distinct classes of keratin genes and their evolutionary significance. Cell. 1981;27(1):75-84. [17] LIU YR, YIN PN, SILVERS CR, et al. Enhanced metastatic potential in the MB49 urothelial carcinoma model. Sci Rep. 2019;9(1):1-9. [18] DYRSKJØT L, KRUHØFFER M, THYKJAER T, et al. Gene expression in the urinary bladder: a common carcinoma in situ gene expression signature exists disregarding histopathological classification. Cancer Res. 2004;64(11):4040-4048. [19] HOGGARTH ZE, OSOWSKI DB, SLUSSER-NORE A, et al. Enrichment of genes associated with squamous differentiation in cancer initiating cells isolated from urothelial cells transformed by the environmental toxicant arsenite. Toxicol Appl Pharmacol. 2019;374:41-52. [20] HOGGARTH ZE, OSOWSKI DB, FREEBERG BA, et al. The urothelial cell line UROtsa transformed by arsenite and cadmium display basal characteristics associated with muscle invasive urothelial cancers. PLoS One. 2018;13(12):e0207877. [21] SJÖDAHL G, LAUSS M, LÖVGREN K, et al. A molecular taxonomy for urothelial carcinoma. Clin Cancer Res. 2012;18(12):3377-3386. [22] GU C, WANG Z, ZHOU N, et al. Mettl14 inhibits bladder TIC self-renewal and bladder tumorigenesis through N 6-methyladenosine of Notch1. Mol Cancer. 2019;18(1):1-12. [23] XU Z, HUANG B, ZHANG Q, et al. NOTCH1 regulates the proliferation and migration of bladder cancer cells by cooperating with long non‐coding RNA HCG18 and microRNA‐34c‐5p. J Cellular Biochem. 2019;120(4):6596-6604. [24] ARSHI A, RAEISI F, MAHMOUDI E, et al. A comparative study of HOTAIR expression in breast cancer patient tissues and cell lines. Cell J. 2020;22(2): 31721532. [25] LEE JS, LEEM SH, LEE SY, et al. Expression signature of E2F1 and its associated genes predict superficial to invasive progression of bladder tumors. J Clin Oncol. 2010;28(16):2660-2667. [26] SIEGEL RL, MILLER KD, JEMAL A. Cancer statistics, 2020. CA: a cancer journal for clinicians. 2020;70(1):7-30. [27] OHISHI T, KOGA F, MIGITA T. Bladder cancer stem-like cells: their origin and therapeutic perspectives. Int J Mol Sci. 2016;17(1):43. [28] BRANDT WD, MATSUI W, ROSENBERG JE, et al. Urothelial carcinoma: stem cells on the edge. Cancer Metastasis Rev. 2009;28(3):291-304. [29] JUNG M, KIM B, MOON KC. Immunohistochemistry of cytokeratin (CK) 5/6, CD44 and CK20 as prognostic biomarkers of non-muscle-invasive papillary upper tract urothelial carcinoma. Histopathology. 2019;74(3):483-493. [30] WU CT, LIN WY, CHEN WC, et al. Predictive Value of CD44 in Muscle-Invasive Bladder Cancer and Its Relationship with IL-6 Signaling. Ann Surg Oncol. 2018;25(12):3518-3526. [31] YANG YM, CHANG JW. Bladder cancer initiating cells (BCICs) are among EMA-CD44v6+ subset: novel methods for isolating undetermined cancer stem (initiating) cells . Cancer Invest. 2008;26(7):725-733. [32] WANG F, CHEN S, LIU HB, et al. Keratin 6 regulates collective keratinocyte migration by altering cell–cell and cell–matrix adhesion. J Cell Biol. 2018; 217(12):4314-4330. [33] WONG P, COULOMBE PA. Loss of keratin 6 (K6) proteins reveals a function for intermediate filaments during wound repair. J Cell Biol. 2003;163(2): 327-337. [34] VOLKMER JP, SAHOO D, CHIN RK, et al. Three differentiation states risk-stratify bladder cancer into distinct subtypes. Proc Natl Acad Sci U S A. 2012;109(6):2078-2083. [35] MAZZUCCHELLI R, SCARPELLI M, GALOSI AB, et al. Pathology of upper tract urothelial carcinoma with emphasis on staging. Int J Immunopathol Pharmacol. 2014;27(4):509-516. [36] SESTERHENN AM, MANDIC R, DüNNE AA, et al. Cytokeratins 6 and 16 are frequently expressed in head and neck squamous cell carcinoma cell lines and fresh biopsies . Anticancer Res. 2005;25(4):2675-2680. [37] LIM SC, PARAJULI KR, HAN SI. Keratin 6, induced by chronic cisplatin exposure, confers chemoresistance in human gastric carcinoma cells. Oncol Rep. 2019;42(2):797-804. [38] KUDO I, ESUMI M, KUSUMI Y, et al. Particular gene upregulation and p53 heterogeneous expression in TP53-mutated maxillary carcinoma. Oncol Lett. 2017;14(4):4633-4640. [39] TSAI FJ, LAI MT, CHENG J, et al. Novel K6-K14 keratin fusion enhances cancer stemness and aggressiveness in oral squamous cell carcinoma. Oncogene. 2019;38(26):5113-5126. [40] FLESKEN-NIKITIN A, HWANG CI, CHENG CY, et al. Ovarian surface epithelium at the junction area contains a cancer-prone stem cell niche. Nature. 2013; 495(7440):241-245. [41] HU J, ZHANG LC, SONG X, et al. KRT6 interacting with notch1 contributes to progression of renal cell carcinoma, and aliskiren inhibits renal carcinoma cell lines proliferation in vitro. Int J Clin Exp Pathol. 2015;8(8):9182-9188. [42] ARUMUGAM A, WENG Z, CHAUDHARY SC, et al. Keratin-6 driven ODC expression to hair follicle keratinocytes enhances stemness and tumorigenesis by negatively regulating Notch. Biochem Biophys Research Commun. 2014;451(3):394-401. [43] Abdou AG, El-Wahed MMA, Kandil MAE, et al. Immunohistochemical analysis of the role and relationship between Notch‐1 and Oct‐4 expression in urinary bladder carcinoma. Apmis. 2013;121(10):982-996. [44] LU N, PIAO MH, FENG CS, et al. Isoflurane promotes epithelial-to-mesenchymal transition and metastasis of bladder cancer cells through HIF-1α-β-catenin/Notch1 pathways. Life Sci. 2020;258:118154. [45] MARAVER A, FERNANDEZ-MARCOS PJ, CASH TP, et al. NOTCH pathway inactivation promotes bladder cancer progression. J Clin Invest. 2015;125(2): 824-830. |
| [1] | 王 景, 熊 山, 曹 金, 冯林伟, 王 信. 白细胞介素3在骨代谢中的作用及机制[J]. 中国组织工程研究, 2022, 26(8): 1316-1322. |
| [2] | 肖 豪, 刘 静 , 周 君. 脉冲电磁场治疗绝经后骨质疏松症的研究进展[J]. 中国组织工程研究, 2022, 26(8): 1323-1329. |
| [3] | 安维政, 何 萧, 任 帅, 刘建宇. 肌源干细胞在周围神经再生中的潜力[J]. 中国组织工程研究, 2022, 26(7): 1183-1190. |
| [4] | 范一鸣, 刘方煜, 张洪宇, 李 帅, 王岩松. 脊髓损伤后室管膜区内源性神经干细胞反应的系列问题[J]. 中国组织工程研究, 2022, 26(7): 1191-1197. |
| [5] | 梁学振, 杨 曦, 李嘉程, 骆 帝, 许 波, 李 刚. 补肾活血胶囊介导Hedgehog信号通路调控大鼠骨髓间充质干细胞成骨成脂分化[J]. 中国组织工程研究, 2022, 26(7): 1068-1074. |
| [6] | 王继芳, 鲍 祯, 乔亚红. miR-206调控小细胞肺癌干细胞EVI1基因表达及细胞生物学行为[J]. 中国组织工程研究, 2022, 26(7): 1075-1079. |
| [7] | 刘 峰, 彭宇环, 罗良平, 吴本清. 植物源性碱性成纤维细胞生长因子维持人胚胎干细胞的生长与分化[J]. 中国组织工程研究, 2022, 26(7): 1080-1085. |
| [8] | 闻丹丹, 李 强, 沈才齐, 纪 哲, 金培生. 外用红色诺卡氏菌细胞壁骨架提高脂肪间充质干细胞活性修复糖尿病创面[J]. 中国组织工程研究, 2022, 26(7): 1086-1092. |
| [9] | 朱兵兵, 邓江华, 陈晶晶, 慕晓玲. 白细胞介素8受体可提高脐带间充质干细胞迁移和向损伤内皮的黏附[J]. 中国组织工程研究, 2022, 26(7): 1093-1098. |
| [10] | 罗小玲, 张 丽, 杨茂桦, 徐 洁, 徐晓梅. 柚皮素干预人牙周膜干细胞的成骨分化能力[J]. 中国组织工程研究, 2022, 26(7): 1099-1104. |
| [11] | 熊挺淋, 应梦慧, 张丽莎, 张晓刚, 杨 燕. 诱导多能干细胞分化的心肌细胞电生理特点[J]. 中国组织工程研究, 2022, 26(7): 1111-1115. |
| [12] | 王新民, 刘 飞, 许 杰, 白玉玺, 吕 剑. 髓芯减压联合牙髓干细胞治疗兔早期激素性股骨头坏死[J]. 中国组织工程研究, 2022, 26(7): 1122-1127. |
| [13] | 房晓磊, 冷 军, 张 晨, 刘会敏, 郭 文. 间充质干细胞不同移植途径治疗缺血性脑卒中疗效差异的系统评价[J]. 中国组织工程研究, 2022, 26(7): 1133-1140. |
| [14] | 郭 嘉, 丁琼桦, 刘 泽, 吕思懿, 周泉程, 高玉花, 白春雨. 间充质干细胞来源外泌体的生物学特性及免疫调控作用[J]. 中国组织工程研究, 2022, 26(7): 1141-1150. |
| [15] | 张璟琳, 冷 敏, 朱博恒, 汪 虹. 干细胞源外泌体促进糖尿病创面愈合的机制及应用[J]. 中国组织工程研究, 2022, 26(7): 1163-1169. |
Keratin 6B (KRT6B) is required for the formation of intermediate filament, which is involved in cytoskeleton construction[16]. A previous study indicated that enhanced KRT6B expression in metastatic tumors of MB-49 sub-clones urothelial carcinoma and enhanced expression of KRT6B in infiltrating bladder cancer patient tissues rather than normal bladder tissues[17-18]. It was demonstrated that KRT6b was significantly elevated in spheres cells (cancer-initiating cells) compared to transformed cell lines and associated with squamous differentiation[19-20]. Moreover, KRT6B has demonstrated high-level expression in the squamous cell cancer-like subtype of urothelial carcinoma, and the elevated expression of KRT6B was associated with poorer prognosis in squamous cell cancer[21], suggesting that KRT6B is an important role in cancer stem cells. The inactivation of the notch pathway is related to the production and progression of bladder cancer. A study revealed that knockout of notch1 suppresses the capacity of tumorsphere formation and invasion of bladder cancer initiating cells[22]. XU et al.[23] found that notch1 inhibited the proliferation and migration of bladder cancer cells with lncRNA HCG18 and microRNA-34c-5p. Numerous studies discussed the functional role of notch pathway in bladder cancer, which may be related to the inactivation of notch pathway.
Therefore, to design targeted therapy for bladder cancer stem cells (BCSCs), we need to fully study the regulatory mechanism of maintaining stemness property including self-renewal in BCSCs. In this article, we attempt to find the regulation of KRT6B on the self-renewal ability and spheroid ability of CD44+ bladder cancer cells and try to discuss the relationship between KRT6B and bladder stem cells, which may be a potential prognostic factor for bladder cancer patients.
中国组织工程研究杂志出版内容重点:干细胞;骨髓干细胞;造血干细胞;脂肪干细胞;肿瘤干细胞;胚胎干细胞;脐带脐血干细胞;干细胞诱导;干细胞分化;组织工程
Cytological in vitro test combined with clinical sample validation.
Time and setting
The experiment was conducted in the Molecular Biology Laboratory of Sino-Singapore Eco-City Hospital of Tianjin Medical University from June 2019 to October 2020.
Materials
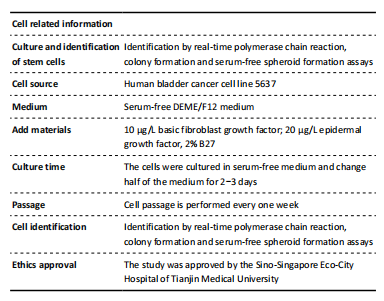
There were human bladder cancer cell line 5637 (Cell Bank of the Chinese Academy of Sciences, Shanghai, China); RPMI 1640 Medium (HyClone, Logan, UT, USA); 10% fetal bovine serum (Biological Industries, Israel); 1% penicillin/streptomycin (Invitrogen, Carlsbad, CA, USA); 10 μg/L human recombinant basic fibroblast growth factor (100-18B; PeproTech, NJ, USA); 20 μg/L human recombinant epidermal growth factor (AF-100-15; PeproTech, NJ, USA); 2% B27 supplement (17504044; GIBCO, Grand Island, NY, USA), and DEME/F12 medium (HyClone, Logan, UT, USA). Two pathologists independently assessed immunohistochemistry staining scores. Totally 24 bladder cancer samples (Immunocytochemistry) were collected for immunohistochemistry undergoing curative resection in Sino-Singapore Eco-City Hospital Affiliated Tianjin Medical University (Tianjin, China).
Methods
Magnetic bead cell sorting (in vitro)
Cells in the logarithmic growth phase were used for magnetic beads sorting. CD44+ and CD44- populations from 5637 cell lines were enriched by MACS. MACS buffer containing MACS BSA Stock Solution (130-091-376; Miltenyi Biotec, Gladbach, Germany) and autoMACS Ringsing Solution (130-091-222; Miltenyi Biotec) were used for preparing the single-cell suspension. Cells were washed and resuspended in 200 μL MASC buffer. CD44 Microbeads (Miltenyi Biotec) were used to incubate cells for 15 minutes at 4 °C. After incubation, CD44+ and CD44- cells were isolated using MACS separation columns (Miltenyi Biotec). All steps were strictly performed according to the protocols of the Miltenyi Biotec manufacturer (Miltenyi Biotec).
Real-time polymerase chain reaction (RT-PCR) (in vitro)
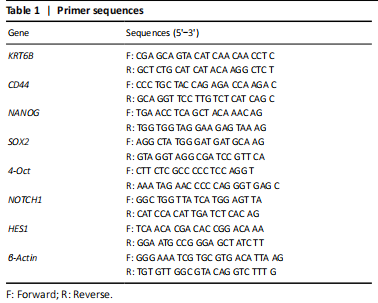
Total RNA was extracted using GeneBetter Cell Total RNA Mini Kit (R017-50, Beijing, China). The total RNA containing 2 μg was reversely transcribed into cDNA using 5× All-In-One RT Master Mix (Applied Biological Materials Inc., Richmond, BC, Canada). The components were mixed well and put into a PCR machine (Bioer Technology, Hangzhou, China) with a setting of 25 °C for 10 minutes, 32 °C for 50 minutes, and 85 °C for 5 minutes. BrightGreen 2× qPCR Master Mix-No Dye (Applied Biological Materials Inc.) and LightCycler 480 qPCR machine (Roche, USA) were used to perform RT-PCR. The cycling conditions were to activate the enzyme for 10 minutes at 95 °C and ran for 40 cycles of 15 seconds at 95 °C and 60 seconds at 60 °C. β-Actin was used as an internal control for Relative RNA expression normalized. The relative quantification was determined using the ΔΔCt method[24]. Specific primer sequences are listed in Table 1.
RNA interference (in vitro)
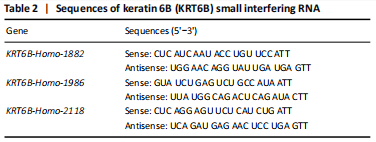
KRT6B small interfering RNA (siRNA; KRT6B-Home-1882, KRT6B-Home-2118, KRT6B-Home-1986) and control siRNA were obtained from Genepharma (Shanghai, China). The 5637 bladder cancer cells were digested for magnetic beads sorting and counted. 5×104 numbers of CD44+ cells were plated in six-well plates adding medium without antibiotics and cultured at 37 °C in an incubator containing 5% CO2 for 15-24 hours. Cells were transfected by KRT6B siRNA with siRNA-mate (Genepharma) according to the manufacturing’ protocol. Cells were collected on 72 hours post-transfection. The sequences of KRT6B siRNA are listed in Table 2. KRT6B was knockdown by siRNAs in BCSCs and the KRT6B siRNA (#KRT6B-Home-1882) was shown as representative data. RT-PCR was used to detect whether the expression of KRT6B was decreased and the relative quantification was determined using the ΔΔCt method.

Colony formation assay (in vitro)
300 cells per well were seeded in the 12-well plates and cultured in an incubator containing 5% CO2 at 37 °C. After 2 weeks, the cells were fixed by 4% methanol for 30 minutes and stained with 0.5% crystal violet solution for 3 minutes. Cell populations of more than 50 cells were counted by an invert microscope (Nikon) with a camera system (NIS-Elements Software, Nikon). Colony formation efficiency was equal to (number of clones/number of seeded cells)×100%.
Wound healing assay (in vitro)
After 6 and 12 hours, for wound healing analysis, 5×105 cells per well were seeded in the 6-well plates and were cultured to 90% confluence at 37 °C and 5% CO2. The bladder cancer cells monolayers were scratched with a 200 μL pipette tip. The floating cells were gently washed with phosphate buffer saline to avoid the cells seeding on scratch. Images of the wound were retained by the photograph using an invert microscope with a camera system. The cells were cultured in serum-free RPMI 1640 medium. After 6 and 12 hours, the cells were observed and photographed by an invert microscope (Nikon). The cell migration areas were calculated by ImageJ software.
Spheroid formation assay (in vitro)
The serum-free DEME/F12 medium (HyClone, Logan, UT, USA) was supplemented with 1% penicillin/streptomycin (Invitrogen), 10 μg/L
human recombinant basic fibroblast growth factor (100-18B; PeproTech, NJ, USA), 20 μg/L human recombinant epidermal growth factor (AF-100-15; PeproTech) and 2% B27 supplement (17504044; Gibco, Grand Island, NY, USA). Then, 2×104 cells per well were seeded on ultra-low attachment 6-well plates (Corning, NY, USA). Cells were cultured in serum-free DEME/F12 medium and half of the medium was changed for 2-3 days. Cell passage was performed every one week. The diameters of tumorspheres were measured and spheres with diameters more than 50 μm were counted by invert microscope (Nikon, Tokyo, Japan) after culture for 2 weeks.
Immunohistochemistry (24 bladder cancer samples)
Immunohistochemistry for protein expression in bladder cancer FEEP sections was performed using specific antibodies including CD44v6 (Zhongshan Inc., Beijing, China) and KRT6B (PeproTech). H2O2 solution was added to the slides for 10 minutes to block endogenous peroxidase activity at room temperature. The slides were put into a staining container full of citrate buffer (Neocell Biotech Co.), and put in a microwave oven set to medium heat for 8 minutes. The container was removed and allowed to cool at room temperature. The diluted KRT6B antibody was added to the sections on the slides and incubated overnight at 4 °C. Biotinylated goat anti-mouse/rabbit IgG (Celplor, Germany) was added to the sections on the slides and incubated at room temperature for 30 minutes. The KRT6B antibody was diluted at a dilution of 1:1000 in phosphate buffer saline. The staining intensity was scored on a scale as achromat (0), stramineous (1), buffy (2), or brown (3). The percentage of positive areas of cells was scored on a scale of 1 (< 25%), 2 (26%-50%), 3 (51%-75%), and 4 (75%-100%). Final protein expression levels were derived from the intensity score adding positive cells staining score. 1-3 was identified as low expression and 4-7 was identified as high expression.
Immunocytochemistry (in vitro)
The diagnosis of bladder cancer was verified by pathological results. Cultured on the coverslips cells were fixed with cold methanol for 30 minutes at 37 °C when the confluency reached 80%. The primary antibodies used for immunocytochemistry was KRT6B (PeproTech) diluted 1:1000 in phosphate buffer saline. The cells were incubated overnight with primary antibodies at 4 °C. Biotinylated goat anti-mouse/rabbit IgG (Celplor, Germany) was added to the slides for 20 minutes. Cells on the slides were incubated with horseradish peroxidase-labeled streptomycin (Zhongshan Inc.) for 15 minutes. The staining process was performed using DAB (Zhongshan Inc.) according to the manufacturer’s protocol.
ONCOMINE database (bioinformatics analysis of 164 bladder cancer tissues)
ONCOMINE is an online microarray database for collecting tumor genetic information. In this study, ONCOMINE was used to compare KRT6B mRNA levels between tumor and normal bladder tissues. Lee database was downloaded to identify the differential expression of KRT6B in bladder cancer[25].
Main outcome measures
There were sorting and identification of CD44+ bladder cancer stem cells, KRT6B mRNA and protein expression level of cells sorted by magnetic beads, cell proliferation, migration, and self-renewal ability after silencing KRT6B.
Statistical analysis
For statistical analysis, GraphPad Prism 7.0 statistical software (GraphPad Software Inc., San Diego, CA, USA) was used to analyze results. Comparisons between groups were performed by Student’s t-test and one-way analysis of variance followed by SNK post hoc test with respect to data in measurement data. The measurement data that obeyed the normal distribution were expressed as the mean ± SD. Two-sided Fisher’s exact test was used for smaller samples. The Kaplan-Meier method was used to analyze the survival curve, and the Gehan-Breslow-Wilcoxon test was used to evaluate the significance. In all results, P ≤ 0.05 was considered statistically significant.
The expression of CD44 was confirmed in basal layer cells both in normal urothelial and bladder cancer[28]. CD44 has been reported associated with production and progression and be involved in worse survival in non-muscle-invasive bladder cancer[29]. In muscle-invasive bladder cancer, the study displayed a high expression of EMT and invasion-related factors in CD44+ cells[30]. In our previous study, the EMA-CD44v6+ subset was considered to possess the features of CSCs as a series of experiments[31]. Another study showed that the ability of tumorigenic of CD44+ cells in mice higher than CD44- cells[14]. CD44 as bladder cancer stem cells biomarker was important for therapeutic target in bladder cancer. The mechanism of maintaining the stemness properties in BCSCs remains to be elucidated. In this study, we found KRT6B was highly related to CD44 in bladder cancer and elevated in CD44+ cells compared to CD44- cells. It is indicated that KRT6B might have an important role in BCSCs. KRT6B is an isoform of human type II keratin 6, which encodes intermediate filament protein in epithelial cells. The wound-inducible CK6 could promote keratinocyte migration and enhance wound healing[32]. The high expression of CK6 regulated cell growth in basal cells during wound repair[33]. In the present study, wound healing assays displayed that the migration ability of KRT6B silenced cells was lower than that of the control group. KRT6B was identified to be down-regulated during cell differentiation using mining developmentally regulated genes[34]. Therefore, KRT6B was predicted by locating basal cells in the bladder and associating them with cell proliferation and migration in bladder cancer. Increasing evidence indicates that KRT6B is highly expressed in a variety of tumor tissues. Bladder cancer with squamous differentiation is the most common variant morphologies in invasive urothelial carcinoma[35]. The main histologic features of invasive bladder cancer are urinary epithelium with distinctive squamous differentiation markers, including keratinocytes and intercellular bridges. The expression of keratins 6 (CK6, K6) and 16 were high and stable in head and neck squamous cell carcinoma[36]. A study revealed that the expression of CK6 was up-regulated in cisplatin-resistant gastric carcinoma cells and it was suggested that CK6 enhances chemoresistance[37]. KUDO et al.[38] also found that the cell adhesion gene KRT6B was upregulated in the TP53 mutant subtype of maxillary squamous cell carcinoma. TSAI et al.[39] have shown that ability of K6-K14 variant 7 contributed to stemness, drug tolerance, and up-regulated TGF-β and G-CSF signaling in oral squamous cell cancer. In an epithelial ovarian cancer study, ovarian surface epithelium expressing KRT6B showed long-term stem cells properties that can easily become malignant[40]. Accumulating evidence has indicated that KRT6B expression is highly correlated to the production and progression of a variety of malignant tumors. In our study, EMA-CD44v6+ cells were isolated by magnetic beads sorting from 10 bladder tumor samples. RNA-seq showed that KRT6B expression levels in EMA-CD44v6+ cells were higher than EMA-CD44v6- cells. Consistent with these previous studies, we found KRT6B was increased in CD44+ bladder cancer cells compared to CD44- bladder cancer cells. Gene silencing of KRT6B in the CD44+ BCSCs decreased proliferation, self-renewal and ability of spheroid formation in vitro. Immunohistochemistry confirmed the obviously correlation between CD44 and KRT6B in tumor tissues of bladder cancer patients. To explore the effect of KRT6B on the prognosis of bladder cancer patients, ONCOMINE dataset analyzed that the bladder cancer patients with high KRT6B expression had a worse prognosis than patients with a low expression.
A study showed that overexpression of KRT6B was correlated with worse prognosis in human renal cell carcinoma and regulated the expression of notch1[41]. ARUMUGAM et al.[42] have found K6 could drive ODC overexpression contributing to tumorigenesis in hair follicles via negatively regulating the Notch pathway. Immunohistochemical analysis showed that Notch1 expression in bladder cancer was negatively correlated with OCT4 expression[43]. Interestingly, in our study, Notch1 and Hes1 expression levels were significantly elevated at mRNA level in KRT6B silenced cells. Although a study showed that the Notch pathway could promote the progression in bladder cancer[44], increasing evidence showed that the Notch pathway can inhibit proliferation, invasion, metastasis, and stem cell properties in bladder cancer[45].
According to previous studies, it is demonstrated that the Notch pathway may play a key role in the maintenance of bladder stem cell properties. Here, we have found that KRT6B is highly expressed in bladder cancer stem cells based on a series of in vitro experiments and lack of in vivo experiments. Therefore, further studies are required to determine whether KRT6B is involved in the maintenance of stemness of bladder cancer stem cells. 中国组织工程研究杂志出版内容重点:干细胞;骨髓干细胞;造血干细胞;脂肪干细胞;肿瘤干细胞;胚胎干细胞;脐带脐血干细胞;干细胞诱导;干细胞分化;组织工程
 #br#
#br#
文题释义:
角蛋白6B:角蛋白6B基因位于染色体12q13,由9个外显子组成,其编码的蛋白是Ⅱ型角蛋白家族中的一员,属于高分子量角蛋白。CD44:是一种跨膜糖蛋白,其基因位于11号染色体上,转录后经过选择性剪切形成多个异构体,其中包括CD44v6。CD44在多种恶性肿瘤中异常表达,尤其在肿瘤干细胞中表达升高。 #br#
中国组织工程研究杂志出版内容重点:干细胞;骨髓干细胞;造血干细胞;脂肪干细胞;肿瘤干细胞;胚胎干细胞;脐带脐血干细胞;干细胞诱导;干细胞分化;组织工程
肿瘤干细胞:肿瘤干细胞被认为是肿瘤中具有干细胞特性的细胞亚群,在肿瘤的形成、进展以及对放化疗的抵抗中扮演着重要角色,在通过手术切除,放疗和化学疗法杀死肿瘤细胞后,肿瘤干细胞仍然会导致肿瘤复发并威胁患者的生命,所以识别并分选出肿瘤干细胞很有可能为治疗癌证提供关键线索。
中国组织工程研究杂志出版内容重点:干细胞;骨髓干细胞;造血干细胞;脂肪干细胞;肿瘤干细胞;胚胎干细胞;脐带脐血干细胞;干细胞诱导;干细胞分化;组织工程
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||