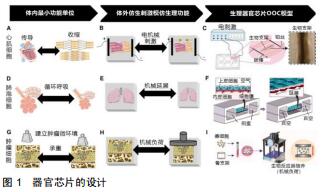
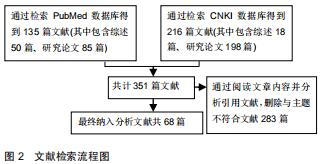
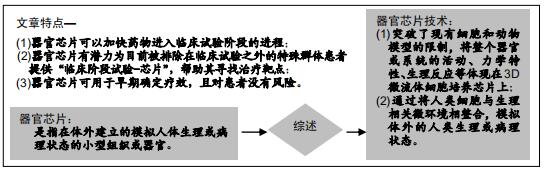
|
[1] HUH D, MATTHEWS BD, MAMMOTO A, et al. Reconstituting organ-level lung functions on a chip. Science.2010;328(5986): 1662-1668.
[2] BOOTH R, KIM H. Characterization of a microfluidic in vitro model of the blood-brain barrier (muBBB). Lab Chip.2012;12(10): 1784-1792.
[3] 王帅超,杨仕平.用于肿瘤微环境研究的微流控细胞芯片构建[D]. 上海:上海师范大学,2018.
[4] WIKSWO JP, CURTIS EL, EAGLETON ZE, et al. Scaling and systems biology for integrating multiple organs-on-a-chip. Lab Chip.2013;13(18):3496-3511.
[5] PARK TH, SHULER ML. Integration of cell culture and microfabrication technology. Biotechnol Prog.2003;19(2):243-253.
[6] BHATIA SN, INGBER DE. Microfluidic organs-on-chips. Nat Biotechnol.2014;32(8):760-772.
[7] GINTANT G, SAGER PT, STOCKBRIDGE N. Evolution of strategies to improve preclinical cardiac safety testing. Nat Rev Drug Discov.2016;15(7):457-471.
[8] GOBAA S, HOEHNEL S, ROCCIO M, et al. Artificial niche microarrays for probing single stem cell fate in high throughput. Nat Methods.2011;8(11):949-955.
[9] PASSIER R, ORLOVA V, MUMMERY C. Complex Tissue and Disease Modeling using hiPSCs. Cell Stem Cell.2016;18(3): 309-321.
[10] POLINI A, PRODANOV L, BHISE NS, et al. Organs-on-a-chip: a new tool for drug discovery. Expert Opin Drug Discov.2014; 9(4): 335-352.
[11] ZHANG B, RADISIC M. Organ-on-a-chip devices advance to market. Lab Chip.2017;17(14):2395-2420.
[12] RUSSELL WMS, BURCH RL. The Principle of Humane Experimental Technique (Wheathampstead, UK: Universities Federation of Animal Welfare). 1959.
[13] HOLMES AM, CRETON S, CHAPMAN K. Working in partnership to advance the 3Rs in toxicity testing. Toxicology.2010;267(1-3): 14-19.
[14] AHADIAN S, CIVITARESE R, BANNERMAN D, et al. Organ-On-A-Chip Platforms: A Convergence of Advanced Materials, Cells, and Microscale Technologies. Adv Healthc Mater.2018;7(14).
[15] LELIEVRE SA, KWOK T, CHITTIBOYINA S. Architecture in 3D cell culture: An essential feature for in vitro toxicology. Toxicol In Vitro.2017;45(Pt 3):287-295.
[16] ROTHBAUER M, ZIRATH H, ERTL P. Recent advances in microfluidic technologies for cell-to-cell interaction studies. Lab Chip.2018;18(2):249-270.
[17] SMITH AS, LONG CJ, BERRY BJ, et al. Microphysiological systems and low-cost microfluidic platform with analytics. Stem Cell Res Ther.2013;4 Suppl 1: S9.
[18] XU Z, LI E, GUO Z, et al. Design and Construction of a Multi-Organ Microfluidic Chip Mimicking the in vivo Microenvironment of Lung Cancer Metastasis. ACS Appl Mater Interfaces.2016;8(39): 25840-25847.
[19] ABACI H E, GLEDHILL K, GUO Z, et al. Pumpless microfluidic platform for drug testing on human skin equivalents. Lab Chip. 2015;15(3):882-888.
[20] ATAC B, WAGNER I, HORLAND R, et al. Skin and hair on-a-chip: in vitro skin models versus ex vivo tissue maintenance with dynamic perfusion. Lab Chip.2013;13(18):3555-3561.
[21] ESCH MB, UENO H, APPLEGATE DR, et al. Modular, pumpless body-on-a-chip platform for the co-culture of GI tract epithelium and 3D primary liver tissue. Lab Chip.2016;16(14):2719-2729.
[22] SHAH P, FRITZ JV, GLAAB E, et al. A microfluidics-based in vitro model of the gastrointestinal human-microbe interface. Nat. Commun. 2016; 7:11535.
[23] TAKAHASHI K, TANABE K, OHNUKI M, et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell. 2007;131(5):861-872.
[24] LIANG P, SALLAM K, WU H, et al. Patient-Specific and Genome-Edited Induced Pluripotent Stem Cell-Derived Cardiomyocytes Elucidate Single-Cell Phenotype of Brugada Syndrome. J Am Coll Cardiol.2016;68(19):2086-2096.
[25] KIM K, DOI A, WEN B, et al. Epigenetic memory in induced pluripotent stem cells. Nature.2010; 467(7313):285-290.
[26] RAJAMOHAN D, MATSA E, KALRA S, et al. Current status of drug screening and disease modelling in human pluripotent stem cells. BioEssays. 2013; 35(3):281-298.
[27] NUNES SS, MIKLAS JW, LIU J, et al. Biowire: a platform for maturation of human pluripotent stem cell-derived cardiomyocytes. Nat. Methods 2013; 10(8): 781–787.
[28] VILLASANTE A, MARTURANO-KRUIK A, VUNJAK-NOVAKOVIC G. Bioengineered human tumor within a bone niche. Biomaterials. 2014;35(22):5785-5794.
[29] VILLASANTE A, MARTURANO-KRUIK A, ROBINSON ST, et al. Tissue-Engineered Model of Human Osteolytic Bone Tumor. Tissue Eng Part C Methods.2017;23(2):98-107.
[30] ZHANG YS, ALEMAN J, SHIN SR, et al. Multisensor-integrated organs-on-chips platform for automated and continual in situ monitoring of organoid behaviors. Proc Natl Acad Sci U S A.2017; 114(12): E2293-E2302.
[31] SHIRURE VS, GEORGE SC. Design considerations to minimize the impact of drug absorption in polymer-based organ-on-a-chip platforms. Lab Chip.2017;17(4):681-690.
[32] VAN MEER BJ, DE VRIES H, FIRTH K, et al. Small molecule absorption by PDMS in the context of drug response bioassays. Biochem Biophys Res Commun.2017;482(2):323-328.
[33] TSAMANDOURAS N, CHEN W, EDINGTON CD, et al. Integrated Gut and Liver Microphysiological Systems for Quantitative In Vitro Pharmacokinetic Studies. AAPS J.2017;19(5):1499-1512.
[34] ONAKPOYA IJ, HENEGHAN CJ, ARONSON JK. Post-marketing withdrawal of 462 medicinal products because of adverse drug reactions: a systematic review of the world literature. BMC Med. 2016,14:10.
[35] OLSSON S, EDWARDS IR. Tachycardia during cisapride treatment. BMJ.1992;305(6856):748-749.
[36] GIACOMELLI E, BELLIN M, SALA L, et al. Three-dimensional cardiac microtissues composed of cardiomyocytes and endothelial cells co-differentiated from human pluripotent stem cells. Development.2017;144(6):1008-1017.
[37] BURRIDGE PW, HOLMSTROM A, and WU JC. Chemically defined culture and cardiomyocyte differentiation of human pluripotent stem cells. Curr Protoc Hum Genet. 2015;87: 1-15.
[38] MANNHARDT I, BRECKWOLDT K, LETUFFE-BRENIERE D, et al. Human Engineered Heart Tissue: Analysis of Contractile Force. Stem Cell Reports.2016;7(1):29-42.
[39] KOSTADINOVA R, BOESS F, APPLEGATE D, et al. A long-term three dimensional liver co-culture system for improved prediction of clinically relevant drug-induced hepatotoxicity. Toxicol Appl Pharmacol.2013;268(1):1-16.
[40] SCHEPERS A, LI C, CHHABRA A, et al. Engineering a perfusable 3D human liver platform from iPS cells. Lab Chip.2016;16(14): 2644-2653.
[41] DOMANSKY K, INMAN W, SERDY J, et al. Perfused multiwell plate for 3D liver tissue engineering. Lab Chip.2010;10(1):51-58.
[42] BHISE NS, MANOHARAN V, MASSA S, et al. A liver-on-a-chip platform with bioprinted hepatic spheroids. Biofabrication.2016; 8(1):14101.
[43] KHETANI SR, BHATIA SN. Microscale culture of human liver cells for drug development. Nat Biotechnol.2008;26(1):120-126.
[44] CHO CH, PARK J, TILLES AW, et al. Layered patterning of hepatocytes in co-culture systems using microfabricated stencils. Biotechniques.2010;48(1):47-52.
[45] RAMAIAHGARI SC, DEN BRAVER MW, HERPERS B, et al. A 3D in vitro model of differentiated HepG2 cell spheroids with improved liver-like properties for repeated dose high-throughput toxicity studies. Arch Toxicol.2014;88(5):1083-1095.
[46] GORAL VN, HSIEH YC, PETZOLD ON, et al. Perfusion-based microfluidic device for three-dimensional dynamic primary human hepatocyte cell culture in the absence of biological or synthetic matrices or coagulants. Lab Chip.2010;10(24):3380-3386.
[47] LEE PJ, HUNG PJ, LEE LP. An artificial liver sinusoid with a microfluidic endothelial-like barrier for primary hepatocyte culture. Biotechnol Bioeng.2007;97(5):1340-1346.
[48] RENNERT K, STEINBORN S, GROGER M, et al. A microfluidically perfused three dimensional human liver model. Biomaterials.2015;71:119-131.
[49] GODOY P, HEWITT NJ, ALBRECHT U, et al. Recent advances in 2D and 3D in vitro systems using primary hepatocytes, alternative hepatocyte sources and non-parenchymal liver cells and their use in investigating mechanisms of hepatotoxicity, cell signaling and ADME. Arch Toxicol.2013;87(8):1315-1530.
[50] LUBBERSTEDT M, MULLER-VIEIRA U, MAYER M, et al. HepaRG human hepatic cell line utility as a surrogate for primary human hepatocytes in drug metabolism assessment in vitro. J Pharmacol Toxicol Methods.2011;63(1):59-68.
[51] WARE BR, BERGER DR, KHETANI SR. Prediction of Drug-Induced Liver Injury in Micropatterned Co-cultures Containing iPSC-Derived Human Hepatocytes. Toxicol Sci.2015; 145(2):252-262.
[52] LI Y, YANG Y, XIONG A, et al. Comparative Gene Expression Analysis of Lymphocytes Treated with Exosomes Derived from Ovarian Cancer and Ovarian Cysts. Front Immunol.2017;8:607.
[53] YIN P, LV H, LI Y, et al. Exosome-Mediated Genetic Information Transfer, a Missing Piece of Osteoblast-Osteoclast Communication Puzzle. Front Endocrinol (Lausanne).2017;8:336.
[54] LI AP, UZGARE A, LAFORGE YS. Definition of metabolism- dependent xenobiotic toxicity with co-cultures of human hepatocytes and mouse 3T3 fibroblasts in the novel integrated discrete multiple organ co-culture (IdMOC) experimental system: results with model toxicants aflatoxin B1, cyclophosphamide and tamoxifen. Chem Biol Interact.2012;199(1):1-8.
[55] LI AP, BODE C, SAKAI Y. A novel in vitro system, the integrated discrete multiple organ cell culture (IdMOC) system, for the evaluation of human drug toxicity: comparative cytotoxicity of tamoxifen towards normal human cells from five major organs and MCF-7 adenocarcinoma breast cancer cells. Chem Biol Interact. 2004;150(1):129-136.
[56] LAU YY, CHEN YH, LIU TT, et al. Evaluation of a novel in vitro Caco-2 hepatocyte hybrid system for predicting in vivo oral bioavailability. Drug Metab Dispos.2004;32(9):937-942.
[57] PHAN D, WANG X, CRAVER BM, et al. A vascularized and perfused organ-on-a-chip platform for large-scale drug screening applications. Lab Chip.2017;17(3):511-520.
[58] OLEAGA C, BERNABINI C, SMITH AS, et al. Multi-Organ toxicity demonstration in a functional human in vitro system composed of four organs. Sci Rep.2016; 6:20030.
[59] MA L, BARKER J, ZHOU C, et al. Towards personalized medicine with a three-dimensional micro-scale perfusion-based two-chamber tissue model system. Biomaterials.2012; 33(17): 4353-4361.
[60] MILLER PG, SHULER ML. Design and demonstration of a pumpless 14 compartment microphysiological system. Biotechnol Bioeng.2016;113(10):2213-2227.
[61] UPADHYAY RK. Drug delivery systems, CNS protection, and the blood brain barrier. Biomed Res Int. 2014; 2014: 869269.
[62] KIMURA H, IKEDA T, NAKAYAMA H, et al. An on-chip small intestine-liver model for pharmacokinetic studies. J Lab Autom. 2015;20(3):265-273.
[63] HABERT R, MUCZYNSKI V, GRISIN T, et al. Concerns about the widespread use of rodent models for human risk assessments of endocrine disruptors. Reproduction.2014;147(4): R119-R129.
[64] CHO I, BLASER MJ. The human microbiome: at the interface of health and disease. Nat Rev Genet.2012;13(4):260-270.
[65] DIMASI JA, HANSEN RW, GRABOWSKI HG. The price of innovation: new estimates of drug development costs. J Health Econ.2003;22(2):151-185.
[66] JEKUNEN A. Decision-making in product portfolios of pharmaceutical research and development--managing streams of innovation in highly regulated markets. Drug Des Devel Ther. 2014;8:2009-2016.
[67] NG KS, THAM LS, TAN HH, et al. Cisapride and torsades de pointes in a pacemaker patient. Pacing Clin Electrophysiol.2000; 23(1):130-132.
[68] INGBER DE. Reverse Engineering Human Pathophysiology with Organs-on-Chips. Cell.2016;164(6):1105-1109.
|