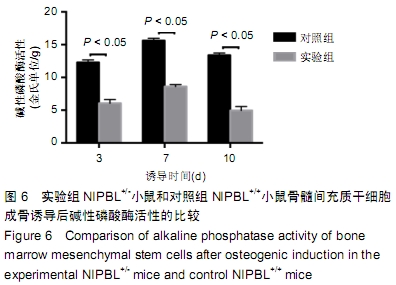
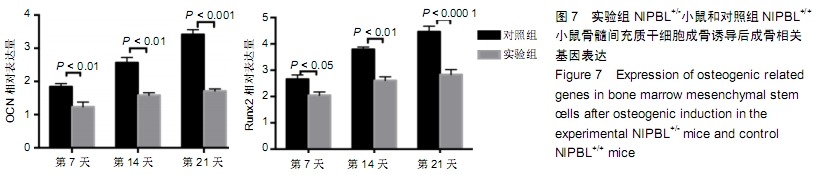
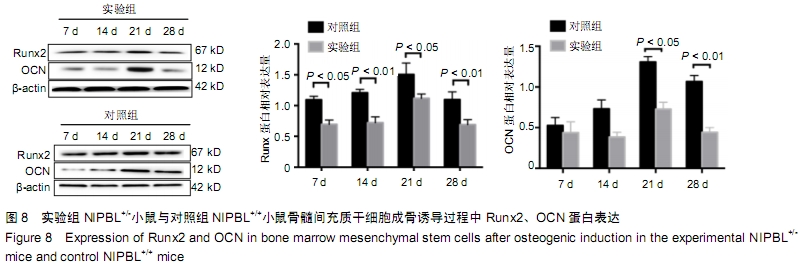
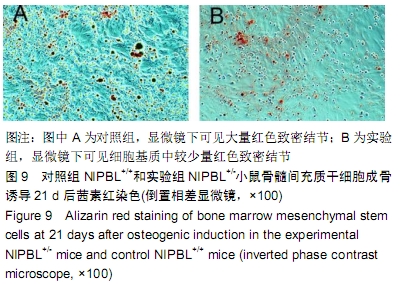
[1] BARISIC I, TOKIC V, LOANE M, et al. Descriptive epidemiology of Cornelia de Lange syndrome in Europe. Am J Med Genet A. 2008; 146A(1):51-59.
[2] MASKOEN AM, LAKSONO B, HAJJAH R, et al. Cornelia de lange syndrome with thyroid agenesis of an indonesian patient. Cell Mol Biol (Noisy-le-grand). 2017;63(8):93-94.
[3] KLINE AD, MOSS JF, SELICORNI A, et al. Diagnosis and management of Cornelia de Lange syndrome: first international consensus statement. Nat Rev Genet. 2018;19(10):649-666.
[4] MUTO A, IKEDA S, LOPEZ-BURKS ME, et al. Nipbl and mediator cooperatively regulate gene expression to control limb development. PLoS Genet. 2014;10(9):e1004671.
[5] ROHATGI S, CLARK D, KLINE AD, et al. Facial diagnosis of mild and variant CdLS: Insights from a dysmorphologist survey. Am J Med Genet A. 2010;152A(7):1641-1653.
[6] BAJAJ S, GAMBHIR P, RANADE S. Short tandem repeats in CdLS-causing genes: distribution and comparison. J Genet. 2014;93(3):e104-107.
[7] KAUR M, MEHTA D, NOON SE, et al. NIPBL expression levels in CdLS probands as a predictor of mutation type and phenotypic severity. Am J Med Genet C Semin Med Genet. 2016;172(2): 163-170.
[8] 卓丽丹,冯顶丽,芦笛,等.大鼠骨髓间充质干细胞和脂肪干细胞体外成骨能力的比较[J].实用口腔医学杂志, 2018,34(6):730-735.
[9] 芮云峰,林禹丞,陈辉,等.晚期骨关节炎患者膝关节滑膜间充质干细胞的体外成骨分化[J].中国组织工程研究, 2013,17(45):7840-7846.
[10] 邢蓬蕊,潘金勇,张惠荣. Shh与Wnt5a基因在Cornelia De Lange综合征中的表达及意义[J].中国当代儿科杂志, 2019,21(5):485-490.
[11] MILLS JA, HERRERA PS, KAUR M, et al. NIPBL+/- haploinsufficiency reveals a constellation of transcriptome disruptions in the pluripotent and cardiac states. Sci Rep. 2018; 8(1):1056.
[12] DEARDORFF MA, BANDO M, NAKATO R, et al. HDAC8 mutations in Cornelia de Lange syndrome affect the cohesin acetylation cycle. Nature. 2012;489(7415):313-317.
[13] BOUDAOUD I, FOURNIER É, BAGUETTE A, et al. Connected Gene Communities Underlie Transcriptional Changes in Cornelia de Lange Syndrome. Genetics. 2017;207(1):139-151.
[14] KRAWCZYNSKA N, WIERZBA J, JASIECKI J, et al. Molecular characterization of two novel intronic variants of NIPBL gene detected in unrelated Cornelia de Lange syndrome patients. BMC Med Genet. 2019;20(1):1.
[15] SIERRA RI, SPECKER BL, JIMÉNEZ F, et al. Biochemical bone markers, bone mineral content, and bone mineral density in rats with experimental nephrotic syndrome. Ren Fail. 1997;19(3): 409-424.
[16] ZHANG Y, XIE RL, CROCE CM, et al. A program of microRNAs controls osteogenic lineage progression by targeting transcription factor Runx2. Proc Natl Acad Sci U S A. 2011;108(24):9863-9868.
[17] TERMINE JD, KLEINMAN HK, WHITSON SW, et al. Osteonectin, a bone-specific protein linking mineral to collagen. Cell. 1981;26(1 Pt 1):99-105.
[18] 陈路,夏先学,蒋科.骨关节炎患者膝关节滑膜组织来源间充质干细胞的增殖与分化[J].中国组织工程研究,2015,19(41):6561-6565.
[19] KRANTZ ID, MCCALLUM J, DESCIPIO C, et al. Cornelia de Lange syndrome is caused by mutations in NIPBL, the human homolog of Drosophila melanogaster Nipped-B. Nat Genet. 2004; 36(6):631-635.
[20] TONKIN ET, WANG TJ, LISGO S, et al. NIPBL, encoding a homolog of fungal Scc2-type sister chromatid cohesion proteins and fly Nipped-B, is mutated in Cornelia de Lange syndrome. Nat Genet. 2004;36(6):636-641.
[21] NEWKIRK DA, CHEN YY, CHIEN R, et al. The effect of Nipped-B-like (Nipbl) haploinsufficiency on genome-wide cohesin binding and target gene expression: modeling Cornelia de Lange syndrome. Clin Epigenetics. 2017;9:89.
[22] NOLEN LD, BOYLE S, ANSARI M, et al. Regional chromatin decompaction in Cornelia de Lange syndrome associated with NIPBL disruption can be uncoupled from cohesin and CTCF. Hum Mol Genet. 2013;22(20):4180-4193.
[23] CARVALHAL S, TAVARES A, SANTOS MB, et al. A quantitative analysis of cohesin decay in mitotic fidelity. J Cell Biol. 2018; 217(10):3343-3353.
[24] ZHOU H, ZHENG L, LU K, et al. Downregulation of Cohesin Loading Factor Nipped-B-Like Protein (NIPBL) Induces Cell Cycle Arrest, Apoptosis, and Autophagy of Breast Cancer Cell Lines. Med Sci Monit. 2017;23:4817-4825.
[25] ALOMER RM, DA SILVA EML, CHEN J, et al. Esco1 and Esco2 regulate distinct cohesin functions during cell cycle progression. Proc Natl Acad Sci U S A. 2017;114(37):9906-9911.
[26] TORTELOTE GG, REIS RR, DE ALMEIDA MENDES F, et al. Complexity of the Wnt/β‑catenin pathway: Searching for an activation model. Cell Signal. 2017;40:30-43.
[27] MAZZOLA M, DEFLORIAN G, PEZZOTTA A, et al. NIPBL: a new player in myeloid cell differentiation. Haematologica. 2019;104(7): 1332-1341.
[28] VEGA OA, LUCERO CMJ, ARAYA HF, et al. Wnt/β-Catenin Signaling Activates Expression of the Bone-Related Transcription Factor RUNX2 in Select Human Osteosarcoma Cell Types. J Cell Biochem. 2017;118(11):3662-3674.
[29] GE C, CAWTHORN WP, LI Y, et al. Reciprocal Control of Osteogenic and Adipogenic Differentiation by ERK/MAP Kinase Phosphorylation of Runx2 and PPARγ Transcription Factors. J Cell Physiol. 2016;231(3):587-596.
[30] CHEN CW, LIN NY, ZHANG Y,et al. SAT0185 Wnt5a Promotes Fibroblast Activation and Tissue Fibrosis by ROR2/ RYK Dependent Activation of PCP-Signaling. Annals of the rheumatic diseases. 2016;75(Suppl 2): 732-735.
[31] 付殷,孙贵才,涂远青,等. 氯化锂促进骨髓间充质干细胞增殖分化的作用途径[J].中国组织工程研究,2019,23(25):3956-3960.
[32] BOT C, PFEIFFER A, GIORDANO F, et al. Independent mechanisms recruit the cohesin loader protein NIPBL to sites of DNA damage. J Cell Sci. 2017;130(6):1134-1146.
|