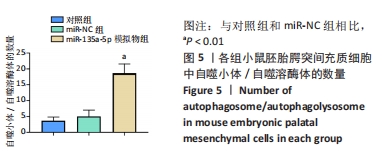
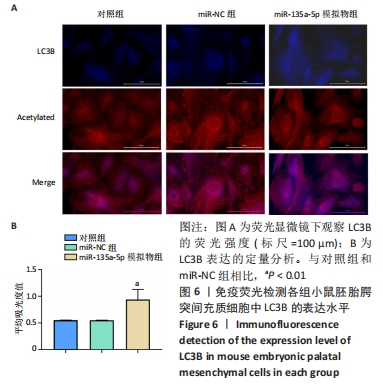
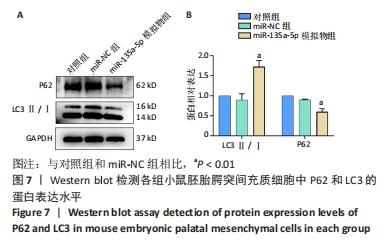
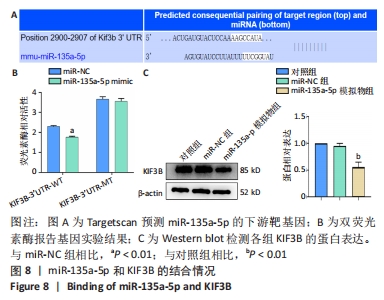
[1] MARTINELLI M, PALMIERI A, CARINCI F, et al. Non-syndromic Cleft Palate: An Overview on Human Genetic and Environmental Risk Factors. Front Cell Dev Biol. 2020;8:592271.
[2] MOSSEY PA, LITTLE J, MUNGER RG, et al. Cleft lip and palate. Lancet. 2009; 374(9703):1773-1785.
[3] DIXON MJ, MARAZITA ML, BEATY TH, et al. Cleft lip and palate: understanding genetic and environmental influences. Nat Rev Genet. 2011; 12(3):167-178.
[4] GRITLI-LINDE A. The etiopathogenesis of cleft lip and cleft palate: usefulness and caveats of mouse models. Curr Top Dev Biol. 2008;84:37-138.
[5] JURILOFF DM, HARRIS MJ, MAGER DL, et al. Epigenetic mechanism causes Wnt9b deficiency and nonsyndromic cleft lip and palate in the A/WySn mouse strain. Birth Defects Res A Clin Mol Teratol. 2014;100(10):772-788.
[6] ZHANG Y, DONG S, WANG J, et al. Involvement of Notch2 in all‑trans retinoic acid‑induced inhibition of mouse embryonic palate mesenchymal cell proliferation. Mol Med Rep. 2017;16(3):2538-2546.
[7] ZHANG H, LIU X, GAO Z, et al. Excessive retinoic acid inhibit mouse embryonic palate mesenchymal cell growth through involvement of Smad signaling. Anim Cells Syst (Seoul). 2016;21(1):31-36.
[8] HAMMOND NL, BROOKES KJ, DIXON MJ. Ectopic Hedgehog Signaling Causes Cleft Palate and Defective Osteogenesis. J Dent Res. 2018;97(13):1485-1493.
[9] HE W, MENG T, WU M, et al. Perturbation of Fgf10 signal pathway in mouse embryonic palate by dexamethasone and vitamin B12 in vivo. J Pediatr Surg. 2010;45(10):2030-2035.
[10] MIRVIS M, STEARNS T, JAMES NELSON W. Cilium structure, assembly, and disassembly regulated by the cytoskeleton. Biochem J. 2018;475(14):2329-2353.
[11] PREVO B, SCHOLEY JM, PETERMAN EJG. Intraflagellar transport: mechanisms of motor action, cooperation, and cargo delivery. FEBS J. 2017;284(18): 2905-2931.
[12] CHANDRA B, TUNG ML, HSU Y, et al. Retinal ciliopathies through the lens of Bardet-Biedl Syndrome: Past, present and future. Prog Retin Eye Res. 2022;89:101035.
[13] KONG JH, SIEBOLD C, ROHATGI R. Biochemical mechanisms of vertebrate hedgehog signaling. Development. 2019;146(10): dev166892.
[14] 殷杰,郭阁,王明雪,等.小鼠胚胎腭间充质细胞的初级纤毛观察[J].武汉大学学报(医学版),2018,39(4):587-590,609.
[15] 郭佳男. IFT122通过初级纤毛介导的Shh信号通路调控mEPMCs增殖的机制研究[D].遵义:遵义医科大学,2020.
[16] SERMERSHEIM MA, PARK KH, GUMPPER K, et al. MicroRNA regulation of autophagy in cardiovascular disease. Front Biosci (Landmark Ed). 2017; 22(1):48-65.
[17] PAMPLIEGA O, ORHON I, PATEL B, et al. Functional interaction between autophagy and ciliogenesis. Nature. 2013;502(7470):194-200.
[18] ORHON I, DUPONT N, PAMPLIEGA O, et al. Autophagy and regulation of cilia function and assembly. Cell Death Differ. 2015;22(3):389-397.
[19] LOU S, MA L, KAN S, et al. Association Study of Genetic Variants in Autophagy Pathway and Risk of Non-syndromic Cleft Lip With or Without Cleft Palate. Front Cell Dev Biol. 2020;8:576.
[20] MENS MMJ, GHANBARI M. Cell Cycle Regulation of Stem Cells by MicroRNAs. Stem Cell Rev Rep. 2018;14(3):309-322.
[21] TANG J, LIAN SB, BAI Y, et al. Comprehensive analysis of plasma miRNA and related ceRNA network in non-syndromic cleft lip and/or palate. Int J Pediatr Otorhinolaryngol. 2022;162:111306.
[22] DING HL, HOOPER JE, BATZEL P, et al. MicroRNA Profiling during Craniofacial Development: Potential Roles for Mir23b and Mir133b. Front Physiol. 2016;7:281.
[23] YOSHIOKA H, SUZUKI A, IWAYA C, et al. Suppression of microRNA 124-3p and microRNA 340-5p ameliorates retinoic acid-induced cleft palate in mice. Development. 2022;149(9):dev200476.
[24] LIU H, PU L, TSAUO C, et al. A new congenital cleft palate New Zealand rabbit model for surgical research. Sci Rep. 2021;11(1):3865.
[25] FUNABASHI T, KATOH Y, OKAZAKI M, et al. Interaction of heterotrimeric kinesin-II with IFT-B-connecting tetramer is crucial for ciliogenesis. J Cell Biol. 2018;217(8):2867-2876.
[26] LANG Z, FAN X, LIN H, et al. Silencing of SNHG6 alleviates hypoxia/reoxygenation-induced cardiomyocyte apoptosis by modulating miR-135a-5p/HIF1AN to activate Shh/Gli1 signalling pathway. J Pharm Pharmacol. 2021;73(1):22-31.
[27] MOSSEY PA, MODELL B. Epidemiology of oral clefts 2012: an international perspective. Front Oral Biol. 2012;16:1-18.
[28] KHANDELWAL KD, VAN BOKHOVEN H, ROSCIOLI T, et al. Genomic approaches for studying craniofacial disorders. Am J Med Genet C Semin Med Genet. 2013;163C(4):218-231.
[29] GAO LY, ZHANG FQ, ZHAO WH, et al. LncRNA H19 and Target Gene-mediated Cleft Palate Induced by TCDD. Biomed Environ Sci. 2017;30(9):676-680.
[30] HE Z, LIU X, LIU X, et al. The role of MEG3 in the proliferation of palatal mesenchymal cells is related to the TGFβ/Smad pathway in TCDD inducing cleft palate. Toxicol Appl Pharmacol. 2021;419:115517.
[31] RADHAKRISHNA U. Small players with a big role: MicroRNAs in pathophysiology of cleft lip and palate. Indian J Hum Genet. 2012;18(3): 272-273.
[32] SHI L, LI B, ZHANG B, et al. Mouse embryonic palatal mesenchymal cells maintain stemness through the PTEN-Akt-mTOR autophagic pathway. Stem Cell Res Ther. 2019;10(1):217.
[33] 瞿功玲. Shh信号通路变化对小鼠胚胎腭突间充质细胞自噬的影响[D].遵义:遵义医科大学,2021.
[34] KO JY, LEE EJ, PARK JH. Interplay Between Primary Cilia and Autophagy and Its Controversial Roles in Cancer. Biomol Ther (Seoul). 2019;27(4):337-341.
[35] CHAKRABORTY J, CAICCI F, ROY M, et al. Investigating mitochondrial autophagy by routine transmission electron microscopy: Seeing is believing? Pharmacol Res. 2020;160:105097.
[36] DIKIC I, ELAZAR Z. Mechanism and medical implications of mammalian autophagy. Nat Rev Mol Cell Biol. 2018;19(6):349-364.
[37] QIN Y, HUANG J, ZHAO X, et al. MiR-135a-5p and Mst1 regulate MPP + -1 induced apoptosis and autophagy in Parkinson’s disease model in vitro. Cell Signal. 2022;94:110328.
[38] CHEN JY, XU LF, HU HL, et al. MiRNA-215-5p alleviates the metastasis of prostate cancer by targeting PGK1. Eur Rev Med Pharmacol Sci. 2020; 24(2):639-646.
[39] COGNÉ B, LATYPOVA X, SENARATNE LDS, et al. Mutations in the Kinesin-2 Motor KIF3B Cause an Autosomal-Dominant Ciliopathy. Am J Hum Genet. 2020;106(6):893-904.
[40] YUAN G, SINGH G, CHEN S, et al. Cleft Palate and Aglossia Result From Perturbations in Wnt and Hedgehog Signaling. Cleft Palate Craniofac J. 2017;54(3):269-280.
[41] SHIN JO, SONG J, CHOI HS, et al. Activation of sonic hedgehog signaling by a Smoothened agonist restores congenital defects in mouse models of endocrine-cerebro-osteodysplasia syndrome. EBioMedicine. 2019;49: 305-317.
[42] ABRAMYAN J. Hedgehog Signaling and Embryonic Craniofacial Disorders. J Dev Biol. 2019;7(2):9.
[43] WANG Q, KUROSAKA H, KIKUCHI M, et al. Perturbed development of cranial neural crest cells in association with reduced sonic hedgehog signaling underlies the pathogenesis of retinoic-acid-induced cleft palate. Dis Model Mech. 2019;12(10):dmm040279.
[44] HUANGFU D, LIU A, RAKEMAN AS, et al. Hedgehog signalling in the mouse requires intraflagellar transport proteins. Nature. 2003;426(6962):83-87.
[45] JIMENEZ-SANCHEZ M, MENZIES FM, CHANG YY, et al. The Hedgehog signalling pathway regulates autophagy. Nat Commun. 2012;3:1200.
[46] WEI C, PAN Y, ZHANG Y, et al. Overactivated sonic hedgehog signaling aggravates intrauterine adhesion via inhibiting autophagy in endometrial stromal cells. Cell Death Dis. 2020;11(9):755.
[47] XIAO Q, YANG Y, QIN Y, et al. AMP-activated protein kinase-dependent autophagy mediated the protective effect of sonic hedgehog pathway on oxygen glucose deprivation-induced injury of cardiomyocytes. Biochem Biophys Res Commun. 2015;457(3):419-425.
[48] PIETROBONO S, GAGLIARDI S, STECCA B. Non-canonical Hedgehog Signaling Pathway in Cancer: Activation of GLI Transcription Factors Beyond Smoothened. Front Genet. 2019;10:556.
[49] YU JS, CUI W. Proliferation, survival and metabolism: the role of PI3K/AKT/mTOR signalling in pluripotency and cell fate determination. Development. 2016;143(17):3050-3060.
[50] PAQUETTE M, EL-HOUJEIRI L, PAUSE A. mTOR Pathways in Cancer and Autophagy. Cancers (Basel). 2018;10(1):18.
[51] MANNING BD, TEE AR, LOGSDON MN, et al. Identification of the tuberous sclerosis complex-2 tumor suppressor gene product tuberin as a target of the phosphoinositide 3-kinase/akt pathway. Mol Cell. 2002;10(1):151-162.
[52] LONG X, LIN Y, ORTIZ-VEGA S, et al. Rheb binds and regulates the mTOR kinase. Curr Biol. 2005;15(8):702-713.
[53] DOBRENEL T, CALDANA C, HANSON J, et al. TOR Signaling and Nutrient Sensing. Annu Rev Plant Biol. 2016;67:261-285.
[54] BOND P. Regulation of mTORC1 by growth factors, energy status, amino acids and mechanical stimuli at a glance. J Int Soc Sports Nutr. 2016;13:8.
[55] HE D, WU H, XIANG J, et al. Gut stem cell aging is driven by mTORC1 via a p38 MAPK-p53 pathway. Nat Commun. 2020;11(1):37.
[56] HE C, KLIONSKY DJ. Regulation mechanisms and signaling pathways of autophagy. Annu Rev Genet. 2009;43:67-93.
|