中国组织工程研究 ›› 2021, Vol. 25 ›› Issue (1): 108-115.doi: 10.3969/j.issn.2095-4344.2135
• 干细胞综述 stem cell review • 上一篇 下一篇
转录因子NKX2-5分子结构与调控心血管前体细胞的生物学功能
王宏书1,韩 燊2,刘 卒2,李晓芳3,钟崇斌4,李立峰4,李亚雄2,蒋立虹5
- 昆明医科大学附属延安医院,1中心实验室,2心脏大血管外科,云南省昆明市 650000;楚雄州人民医院,3全科医学科,4胸心外科,云南省楚雄市 675000;5云南省第一人民医院,云南省昆明市 650100
-
收稿日期:2019-12-05修回日期:2019-12-10接受日期:2020-02-19出版日期:2021-01-08发布日期:2020-11-19 -
通讯作者:蒋立虹,博士,教授,主任医师,云南省第一人民医院,云南省昆明市 650100 李亚雄,教授,主任医师,昆明医科大学附属延安医院心脏大血管外科,云南省昆明市 650000 -
作者简介:王宏书,男,1986年生,云南省大姚县人,彝族,在读博士,医师,主要从事胸心外科疾病的基础与临床研究。 -
基金资助:国家自然科学基金(81560060)
Transcription factor NKX2-5: molecular structure and biological function in regulating cardiovascular precursor cells
Wang Hongshu1, Han Shen2, Liu Zu2, Li Xiaofang3, Zhong Chongbin4, Li Lifeng4, Li Yaxiong2, Jiang Lihong5
- 1Central Laboratory, 2Department of Cardiovascular Surgery, Yan’an Hospital Affiliated to Kunming Medical University, Kunming 650000, Yunnan Province, China; 3Department of General Medicine, 4Department of Thoracic Surgery, the People’s Hospital of Chuxiong Prefecture, Chuxiong 675000, Yunnan Province, China; 5The First People’s Hospital of Yunnan Province, Kunming 650100, Yunnan Province, China
-
Received:2019-12-05Revised:2019-12-10Accepted:2020-02-19Online:2021-01-08Published:2020-11-19 -
Contact:Jiang Lihong, MD, Professor, Chief physician, The First People’s Hospital of Yunnan Province, Kunming 650100, Yunnan Province, China Li Yaxiong, Professor, Chief physician, Department of Cardiovascular Surgery, Yan’an Hospital Affiliated to Kunming Medical University, Kunming 650000, Yunnan Province, China -
About author:Wang Hongshu, MD candidate, Physician, Central Laboratory, Yan’an Hospital Affiliated to Kunming Medical University, Kunming 650000, Yunnan Province, China -
Supported by:the National Natural Science Foundation of China, No. 81560060
摘要:
文题释义:
转录因子:本质为蛋白质,其DNA序列经转录、翻译后形成蛋白质,可进入细胞核中,结合于下游基因启动子区的特定DNA序列,激活或抑制其调控的下游基因的转录,从而调控该基因功能。
先天性心脏病:是一类因胚胎期心脏发育异常而导致的出生缺陷性疾病,根据病变位置,可分为房间隔缺损、室间隔缺损、动脉导管未闭、法洛氏四联症等临床常见亚型。因心脏结构异常,导致血流动力学紊乱,是引起新生儿死亡的常见疾病。
背景:NKX2-5是哺乳动物心脏发育过程中极其重要的一个转录调节因子。来自于模式动物如大鼠、小鼠及斑马鱼的研究表明,NKX2-5的缺失或功能异常,影响心脏发育,出现类似于人类先天性心脏病的病理表现。然而,对于NKX2-5在心脏发育过程中的调控作用及具体机制,目前依然不明确。
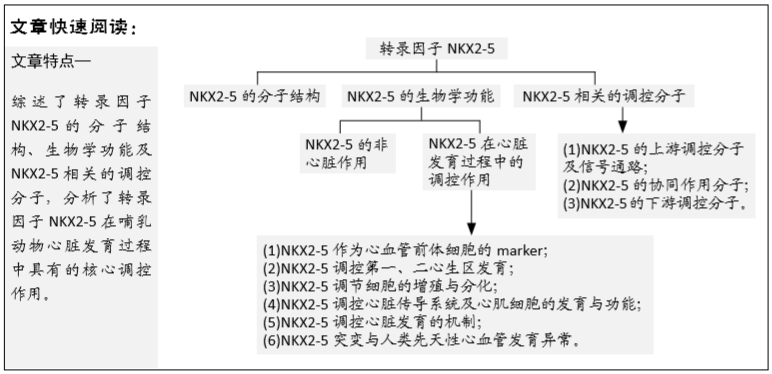
目的:对NKX2-5的分子结构、功能作用、上下游调控分子进行综述。
方法:以检索词“NKX2-5、congenital heart disease、CHD、heart development”在PubMed数据库(网址https://www.ncbi.nlm.nih.gov/pubmed/)中进行检索,文献发表截止时间为2019-04-01。通过对检索到的文献进行下载、阅读,排除与NKX2-5无关的文献、观点重复、结论一致的文献,对剩余的97篇文献进行详细阅读,提取文献中的观点、结论进行综述。
结果与结论:①NKX2-5在心脏发育调控中起关键作用,其功能异常或基因突变导致心脏发育异常与功能异常,与人类先天性心脏病的发生密切相关;②NKX2-5调控作用的发挥依赖于其自身的多个功能性结构域。NKX2-5经转录翻译后进入细胞核内,通过与CO-factors结合或单独结合于特定DNA序列,对下游调控分子起激活或抑制作用,从而对心肌前体细胞的增殖、迁移、分化、功能产生影响,维持正确的心脏发育过程及心脏传导系统的正常发育与功能;③NKX2-5的转录、翻译、入核过程、转录激活或抑制作用的发挥也受到多种方式的调控,主要从染色质构象、启动子与增强子功能、RNA解链、转录后修饰、细胞核定位层面对NKX2-5进行调控。
中图分类号:
引用本文
王宏书, 韩 燊, 刘 卒, 李晓芳, 钟崇斌, 李立峰, 李亚雄, 蒋立虹. 转录因子NKX2-5分子结构与调控心血管前体细胞的生物学功能[J]. 中国组织工程研究, 2021, 25(1): 108-115.
Wang Hongshu, Han Shen, Liu Zu, Li Xiaofang, Zhong Chongbin, Li Lifeng, Li Yaxiong, Jiang Lihong. Transcription factor NKX2-5: molecular structure and biological function in regulating cardiovascular precursor cells[J]. Chinese Journal of Tissue Engineering Research, 2021, 25(1): 108-115.
NK2结构域富含脯氨酸,可能为NKX2-5蛋白之间或与其他互作蛋白之间的反应接触面,在体外实验中发现其功能是调节NKX2-5蛋白的转录激活活性[11]。
YRD结构域富含酪氨酸,对NKX2-5的DNA结合特性和转录激活活性无太大影响,但在胚胎干细胞分化为心肌细胞过程中,为NKX2-5发挥功能所必需[12],同时在NKX2-5蛋白HD结构域缺失的情况下,YDR结构域可作为与野生型NKX2-5蛋白或互作蛋白相互作用的接触面以形成异源二聚体[13]。
而TN结构域的功能到目前为止依然不清。
2.2 NKX2-5的生物学功能
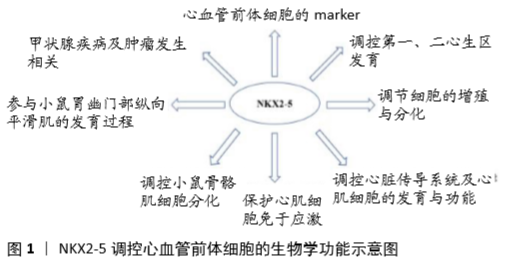
2.2.1 NKX2-5在心脏发育过程中的调控作用 NKX2-5在哺乳动物中具有高度同源性,其中人类NKX2-5与小鼠Nkx2-5具有87%的整体同源性,而HD结构域同源性则为100%[14]。同时小鼠具有与人类相似的心脏结构及心脏发育过程,因此多作为研究心脏发育过程的模式动物[15]。研究发现,NKX2-5主要具有以下作用,见图1。
(1)NKX2-5作为心血管前体细胞的marker:NKX2-5最早表达于胚胎头折早期时的心肌前体细胞,其表达早于心肌细胞的分化,并持续表达于胚胎期、胎儿期和成人期心脏[16]。NKX2-5阳性的心脏前体细胞发育成熟为心肌细胞[17]。当从发育中的小鼠胚胎中将NKX2-5阳性细胞分离出来继续培养,可发现大多数细胞分化为心肌细胞和传导系统细胞,部分细胞发育为平滑肌细胞[18]。而在生心区外,NKX2-5一过性表达,这些瞬时NKX2-5阳性的细胞朝心脏和背动脉迁移,最终形成血液干细胞和血管细胞[19]。PAFFETT-LUGASSY等[20]研究发现NKX2-5阳性细胞也是咽弓动脉内皮细胞的前体细胞,同时咽弓动脉内皮细胞的分化也依赖于NKX2-5。上述研究表明NKX2-5为心血管系统前体细胞的标志,同时也提示其与心血管系统发育密切相关。
(2)NKX2-5调控第一、二心生区发育:ZHANG等[3]通过免疫组化染色对小鼠胚胎进行研究,发现NKX2-5表达于胚胎期心脏的全区域,包括原始的左心室结构、第二生心区发育形成的流出道干、右心室、心房及静脉窦。当分别对内胚层和中胚层的Nkx2-5进行敲除,发现内胚层中Nkx2-5的敲除并不对心脏发育产生明显影响,而中胚层的Nkx2-5敲除则导致心脏发育缺陷,此外当对Nkx2-5-/-小鼠进行中胚层Nkx2-5再表达时可挽救其第二生心区发育缺陷[3]。
在第二生心区内,Nkx2-5的表达存在自我调节机制,对于维持Nkx2-5的表达具有重要作用,而这种自我调控机制具有进化保守型,但在哺乳动物和非哺乳动物之间存在一定差异。在鸡这类非哺乳动物中,NKX2-5直接结合于DNA增强子区域以维持Nkx2-5的表达,而在小鼠为代表的哺乳动物中,NKX2-5间接通过Mef2c实现自我调节[21]。
PRALL等[22]研究发现,在小鼠第二生心区内,Nkx2-5通过负反馈抑制Bmp2/Smad1信号通路,调控第二生心区细胞增殖及流出道的形成过程。具体机制:Nkx2-5突变引起调控心肌细胞特异性和维持心脏前体细胞处于前体状态的基因失调,起初引起心脏前体细胞过度心肌特异化,随后这种特异化的心肌细胞不能增殖以形成第二生心区和流出道干,导致心脏发育异常[22]。COLOMBO等[23]研究发现斑马鱼模型中Nkx基因通过不同的机制调控流出道、静脉极和窦房结处心脏前体细胞的增殖和分化,以调控斑马鱼心脏流出道结构的发育。
GEORGE等[24]研究发现,NKX2-5的心脏发育调控作用在不同的细胞群体中具有时间性。在Nkx2-5-/-斑马鱼胚胎中,心脏发育早期的腔室结构可以正常的分化和特异化,但在心脏发育后期出现心房和心室的特异性转换,而在早期过表达Nkx2-5则可以挽救这种晚期出现的病理改变,而且这种早期过表达效果还可以维持心室特性及影响成年期后的心脏功能。这些调控作用可能是通过NKX2-5诱导下游靶基因激活实现的。而这些下游靶基因在心脏发育分化的早期、晚期阶段,为维持心脏腔室的特异性所必需[24]。
上述研究结果表明,NKX2-5的心脏调控作用主要作用于心脏发育的中胚层区域,尤其是第二生心区,并为第二生心区的发育所必需。在第二生心区中NKX2-5的表达具有自我调控机制,这种自我调控机制使得NKX2-5的表达得以维持。NKX2-5通过对第二生心区内的心脏前体细胞的增殖、分化进行调控,以形成完整而正常的心脏流出道结构及静脉极和窦房结结构。同时NKX2-5的调控作用具有时间位点性,在早期和晚期可产生不同的调控效应。此外,NKX2-5的调控作用还具有细胞位置特异性[25],RYAN等[26]研究发现肌球蛋白磷酸化酶可影响NKX2-5的细胞定位,使NKX2-5从细胞核中排出,从而抑制其转录激活活性并对心肌发生过程产生抑制。
(3)NKX2-5调节细胞的增殖与分化:NAKASHIMA等[27]研究发现,在心脏腔室形成后,NKX2-5抑制小鼠心房细胞和传导系统细胞的增殖。在小鼠心房特异性敲除Nkx2-5后,小鼠心脏原发孔变大,心房肌变薄,同时出现房室传导阻滞,伴随Nkx2-5下游调控基因的表达异常[27]。
DORN等[28]研究中发现NKX2-5直接结合于下游调控基因Isi1的增强子区域,抑制Isi1的转录活性,这种抑制作用决定了Isi1阳性的心脏前体细胞发育为心室肌细胞系,同时也提示NKX2-5与其他基因相互调节,以决定心脏前体细胞朝不同的心肌细胞系分化,并使其获得相应的心肌细胞特性[28]。
在间充质干细胞这类具有多向分化潜能的干细胞中,过表达NKX2-5可以促进脐带间充质干细胞向心肌样细胞分化,一些心肌基因如CTNI、DESMIN、GATA4等表达上调[29]。在大鼠骨髓间充质干细胞中过表达NKX2-5可以促进移植入大鼠体内的间充质干细胞分化为心肌样细胞,促进间充质干细胞对受损心肌的修复作用,改善心衰大鼠模型的心功能和降低心肌纤维化,同样过表达NKX2-5也会引起心肌基因的表达上调[30]。这些结果提示,即使在非心脏前体细胞的多能性干细胞中过表达NKX2-5,依然会引起具有多能分化特性的干细胞向心肌样细胞分化。这种过表达心肌调控因子而实现的诱导分化,有可能使得未来利用非心脏前体细胞诱导分化为心肌细胞成为可能。
(4)NKX2-5调控心脏传导系统及心肌细胞的发育与功能:JAY等[31]研究发现敲除胚胎期小鼠Nkx2-5导致小鼠心脏的房室结、His束和浦肯野系统的发育不全,说明Nkx2-5对心脏传导系统发育具有调控作用。BRIGGS等[32]研究发现,在围产期敲除小鼠Nkx2-5会导致小鼠出现快速进展的心脏传导功能障碍和心衰。通过基因表达谱分析发现,多种离子通道基因的表达出现改变,包括T型钙离子通道、Nav1.5-a亚基和RYR2等基因表达降低。比较特殊的是作为心脏内Na离子通道的主要亚型Nav1.5-a亚基的基因表达只在心室内降低,而在心房内则无明显改变[32]。此外,Nkx2-5敲除导致Ca离子处理缺陷,由此出现心肌细胞收缩功能障碍[32]。TAKEDA 等[33]在2周龄时对已经完成终末分化的小鼠心肌细胞进行Nkx2-5敲除,同样发现了相同的心脏收缩功能障碍和传导障碍。PASHMFOROUSH等[34]于心室特异性敲除小鼠Nkx2-5,发现并不会导致房间隔缺损等心脏结构改变,但却会出现进展性的心脏传导阻滞和大量的心室肌小梁增生。小鼠于出生时表现为房室结发育不良,而随后发生选择性的传导细胞脱落和纤维化。基因表达谱分析发现,Bmp10、Hop等与心肌细胞增殖和心室肌小梁形成的基因及传导系统的基因表达均出现异常[34]。HARRINGTON等[35]研究表明Nkx2-5为维持斑马鱼心律变异性所必需。上述结果说明,Nkx2-5对心脏传导系统的发育及分化成熟后的功能维持都具有关键性作用。通过对心律失常患者的基因测序研究,也发现NKX2-5的变异与心律失常,如心脏骤停猝死、室性心律失常,尤其是房颤及心脏收缩功能异常存在相关性[36-38]。
(5)NKX2-5调控心脏发育的机制:①NKX2-5通过HEY2依赖性的转录网络调控心脏发育:ANDERSON等[39]利用人胚胎干细胞研究发现,当在人胚胎干细胞中敲除NKX2-5将会导致心肌形成障碍,同时心肌细胞收缩和电生理异常。通过RNA-seq及chip-seq技术发现,IRX4、HAND1和MYL2等心室特异性标记基因表达下调,而心肌前体细胞的标记基因如ISL1、PDGFRA、BMP2和FGF10上调,同时NKX2-5依赖性基因如HEY2、IRX4、NPPA、MYL2和VCAM1等下调。通过挽救实验验证了依赖于HEY2的转录调控网络的异常,导致了上述的细胞表型及下游调控基因改变。②NKX2-5通过影响mRNA的polyA尾长度调控心脏发育:转录因子在被翻译为蛋白质前需由DNA转录为mRNA,而mRNA多聚A尾的长度决定了mRNA被翻译为蛋白质的效率。NIMURA等[40]研究发现NKX2-5除了经典的转录因子调控作用外,NKX2-5同5’-3’核酸外切酶XRN2可通过调控与心脏发育相关基因的mRNA多聚A尾的长度而调控小鼠心脏发育。Nkx2-5敲除导致长的3’ UTR的mRNA出现,而这些基因与心脏发育相关,这种调节方式可能是NKX2-5通过其HD结构域识别特定DNA序列,并招募XRN2聚集于此而发挥作用[40]。③NKX2-5与MEIS1通过次序性结合下游基因调控心脏发育:转录因子通过功能结构域识别并结合特定DNA序列,发挥转录激活或抑制作用。在DUPAYS等[41]研究中发现NKX2-5与MEIS1含有相同的DNA结合域,可相互排斥性的结合下游调控基因。通过次序性的结合下游基因,NKX2-5与MEIS1调控心脏分化过程。
(6)NKX2-5突变与人类先天性心血管发育异常:NKX2-5作为心脏发育过程中关键的调控分子,其功能异常往往导致心脏结构及功能异常[42]。在人类心血管疾病患者中,通过基因测序研究发现近50个NKX2-5突变[43],而这些突变与多种心血管发育异常相关[44-48],其中NKX2-5突变与先天性心脏,尤其是房间隔缺损密切相关[49-50]。
ZAKARIYAH等[51]研究发现,通过在小鼠Nkx2-5中敲入与人类先心病发生高度相关的R142C突变,导致小鼠胚胎于E10.5出现心脏发育延迟及胚胎停滞并伴有下游基因的表达下调。组织学检查发现,小鼠心脏出现房间隔缺损、间隔缺损,离子通道基因下调,心电图PR间期延长。R142C突变位于NKX2-5的DNA结合域和核定位序列内,来源于人类先心病病例的研究提示,R142C突变与房间隔缺损和房室传导阻滞相关[52]。PABST等[53]在家族性先天性心脏病患者中经测序研究发现位于NKX2-5第一外显子处的c.325G>T突变,这种突变被预测会在109位氨基酸位置处产生一个终止密码子从而导致断裂蛋白形成,断裂蛋白中缺乏NKX2-5的功能性结构域。在这些家族患者中,心脏发育异常表现有房间隔缺损、房室传导阻滞、室间隔缺损、肺动脉狭窄、卵圆孔未闭。在另外的研究中也有突变导致NKX2-5断裂蛋白形成的报道[50]。HUANG等[54]在室间隔缺损患者的NKX2-5上游增强子区域中发现g.17483576C>G和g.17483564C>T突变,通过功能验证发现这2个突变能显著降低增强子的活性。OUYANG等[46]研究发现位于NKX2-5的第2外显子2 bp的插入突变c.512insGC,这种插入型突变损害了NKX2-5的核定位结构,导致转录激活活性的丧失。除了上述常见的畸形外,还有NKX2-5突变导致其他心血管畸形的报道。QU等[47]发现NKX2-5的p.K192X突变被认为与二叶式主动脉瓣畸形相关,而BIBEN等[48]也报道了NKX2-5突变与瓣膜狭窄的相关性。WATANABE等[55]发现NKX2-5的移码突变导致内脏异位及静脉窦性房间隔缺损。
到目前为止,在先天性心脏病患者中检测出的NKX2-5突变中,单核苷酸多态性位点rs2277923和rs3729753被报道的较多[43,56-57]。rs2277923和rs3729753为无义突变,致病机制依然不明确,其是否在mRNA层面上致病依然值得继续深入研究。
2.2.2 NKX2-5的非心脏作用 随着对NKX2-5研究的深入,发现NKX2-5除了具有心脏发育调控作用外,还有其他功能。例如Nkx2-5除了调控心脏特异性基因表达外,还能保护心肌细胞免于应激[58-59]。除了调控心脏发育外,Nkx2-5还能调控小鼠骨骼肌细胞分化,在体外促进骨骼肌细胞向神经元样细胞分化[60]。在小鼠胃幽门部纵向平滑肌的发育过程中也需要NKX2-5和GATA3的参与[61]。HWANG等[62]研究发现NKX2-5可能在皮肤细胞从增殖向分化转变的过程中起作用,同时参与异常的皮肤角化性疾病的病理过程。DRITSOULA等[63]发现在系统性硬化的血管重构过程中出现NKX2-5异常表达。
除了上述作用外,NKX2-5还与甲状腺疾病和肿瘤相关。在甲状腺发育不全患者中,发现有NKX2-5的突变出现[64]。PENHA等[65]发现NKX2-5表达于乳头状甲状腺癌中,调控甲状腺细胞分化。NAGEL等[66]关于儿童急性淋巴细胞白血病的研究发现,与BCL11B处于并立位置的NKX2-5出现异常代替了TLX3表达。而SHIBATA等[67]的研究也提示NKX2-5参与肿瘤发生的作用。
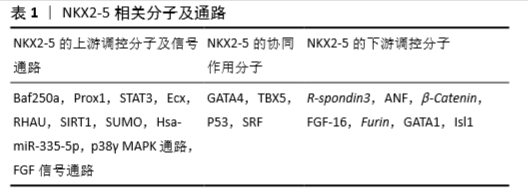
2.3 NKX2-5相关的调控分子 作为一个转录调节因子,NKX2-5也接受上游调控因子的调节,并单独或与共调节分子作用,调控下游靶分子,从而完成转录调节过程。这些NKX2-5相关的调控分子可分为上游调控分子、共同作用分子及下游调控分子,见表1。
2.3.1 NKX2-5的上游调控分子及信号通路
(1)Baf250a:Baf250a是依赖ATP的染色质构象调节复合物SWI/SNF的关键调节成分。在小鼠窦房结中,Baf250a通过激活Tbx3的表达,然后同TBX3、HDAC3一起抑制窦房结中Nkx2-5的表达[68]。
(2)Prox1:Prox1属于同源盒因子,其在心脏发育中起维持肌肉结构的作用[69]。在小鼠心脏传导系统的已分化心肌细胞中,Prox1与Nkx2-5共同表达并负向调控Nkx2-5的表达,以确保成熟的心房和心室传导系统成分的解剖学完整,尤其是房室结和浦肯野纤维系统[70]。Prox1的敲低可以挽救Nkx2-5杂合子突变小鼠中浦肯野纤维系统发育不良、房室结发育不良的病理改变[70]。
(3)STAT3:SNYDER等[71]利用小鼠畸胎瘤细胞P19研究发现,STAT3直接结合于Nkx2-5的启动子区,调控其表达。
(4)心脏结合素Ecx:在斑马鱼中,突变ftk导致类似于人类先天性心脏病的进展性的心肌成熟异常和心脏形成异常,伴随有Nkx2-5表达严重下调。利用挽救实验证明了Nkx2-5是Ecx衍生信号通路的下游调节成分,并且与ftk异常所致病理表型相关[72]。
(5)RHAU:RNA G-4链体在RNA的生物学过程中起重要作用[73]。RHAU(DHX36, G4R1)是一种RNA解旋酶,可以解旋G-4链体。在Nkx2-5 mRNA的5’非编码区含有4链体结构,这个4链体结构需要RHAU的解链作用,同时在3’非编码区含有AU-富集原件,此原件有利于RHAU介导的mRNA降解。RHAU通过对Nkx2-5的mRNA调控,影响其稳定性和翻译效益,从而实现Nkx2-5的转录后调控[74]。
(6)SIRT1:转录因子的乙酰化与去乙酰化作用,会对其细胞器定位、稳定性和转录激活活性产生影响[75]。TANG等[76]研究发现SIRT1能对NKX2-5去乙酰化,抑制其转录激活活性。SIRT1与NKX2-5的COOH端相互作用,并在HD结构域的第182位赖氨酸处去乙酰化,此处的突变也会导致NKX2-5的转录活性降低。此外,SIRT1还可通过降低NKX2-5与其共作用分子的结合而影响NKX2-5的下游靶基因的表达。
(7)SUMO:小泛素样修饰物(mall ubiquitin-like modifier,
SUMO)是一类可以共价可逆性结合到赖氨酸残基上的小分子物质。SUMO通过对NKX2-5的第51位赖氨酸进行修饰而增强其转录激活活性[77],而K51R的突变显著降低了NKX2-5的DNA结合活性与转录激活活性[75]。
(8)ErbB4和p38γ MAPK:RAMACHRISHAN等[78]通过敲低ErbB4 和p38γ,然后分离核内和胞内NKX2-5进行检测,发现p38γ可能通过影响NKX2-5的核内定位而影响其作用。
(9)FGF信号通路:心脏的心房腔和心室腔具有不同的组织形成过程、电生理特性和收缩特性,这取决于不同的基因表达。FGF通路通过维持Nkx2-5表达而调控心房、心室特性。FGF信号缺失导致心室内异位的心房性基因表达,心房性基因高表达,心室性基因低表达,心房细胞增多和同期心室细胞减少,这些结果提示FGF通路作为Nkx2-5的上游调控其功能[79]。
(10)Hsa-miR-335-5p:除了常见的转录因子与信号通路外,miRNA也有报道参与对NKX2-5的调控。KAY等[80]在人胚胎干细胞中通过过表达及抑制hsa-miR-335-3p和hsa-miR-335-5p,发现与心肌分化相关的基因如NKX2-5的表达改变,提示hsa-miR-335-5p对其表达有调控作用。
2.3.2 NKX2-5的协同作用分子 研究表明,NKX2-5在发挥调控作用时并非单独发生作用,也存在与其相互合作的调控分子。
(1)GATA4:GATA4和NKX2-5均为心脏前体细胞的标记,为心肌细胞分化所必需。以ANF启动子序列为基础构建载体,通过共同表达GATA4和NKX2-5,发现协同促进作用[81]。这种协同作用涉及GATA4的C端锌指结构和NKX2-5 HD结构域的第3螺旋结构[82-83]。
(2)TBX5:通过共转染Nkx2-5和Tbx5,发现二者之间相互作用,协同促进下游基因的表达。pull-down实验提示,TBX5氨基端的结构域和N端的部分序列以及NKX2-5的HD结构域为二者发生相互作用所必需[84]。
(3)P53:KOJIC等[85]发现NKX2-5与P53可协同激活Ankrd2的启动子。P53可以同野生型及突变型的NKX2-5发生相互作用,这种作用发生于NKX2-5的C端区域,而P53则位于C端外。
(4)SRF:依赖于SRF完整的血清反应元件结构,NKX2-5与SRF共同对小鼠心肌α-Actin的启动子产生强烈的激活作用。这一过程的实现依赖于SRF通过DNA结合作用将NKX2-5募集于α-Actin的启动子区[86]。
2.3.3 NKX2-5的下游调控分子
(1)R-spondin3 (Rspo3):R-spondin3 (RSPO3)是分泌型因子,可强力激活WnT通路而调控干细胞增殖,当突变Nkx2-5时R-spondin3表达显著下调。通过BLAST和VISTA分析,提示在R-spondin3的5’存在有NKX2-5的结合位点,而进一步的CHIP和luciferase实验发现了6个结合位点并验证了NKX2-5对R-spondin3的调控作用[87]。
(2)ANF:ANF大量表达于心房肌细胞和心室肌小梁层,其编码的28个氨基酸的多肽作用于心血管发育中,调控液体和电解质平衡。而NKX2-5是ANF最主要的转录激活分子之一。研究发现,NKX2-5可通过结合于ANF的3个调控元件(kb-34,-31和-21)和ANF的启动子远端而调控其表达[82,88]。
(3)β-Catenin:RIAZI等[89]发现NKX2-5通过结合到
β-Catenin启动子区的特殊元件而调控β-Catenin的表达。通过对β-Catenin表达的调控,NKX2-5实现对WNT通路的调节。
(4)FGF-16:FGF-16在出生后由新生心肌细胞表达,并逐步增加,直至成年,其在维持心肌存活方面发挥作用,包括可增加心肌细胞对阿霉素诱导引起的损伤。敲低Nkx2-5会降低内源性的Fgf-16的RNA和蛋白水平,而过表达Nkx2-5则增加FGF-16的表达水平,同时增强其对快速型阿霉素诱导损伤的抵抗作用,这些结果提示FGF-16的表达受Nkx2-5直接调控[90]。
(5)Furin:Furin是一种分泌型的前蛋白转化酶基因,可通过有限的水解位于蛋白内部的一个或多个位点从而激活蛋白前体为具有生物学活性的激活体[91-92]。因此,多种转录因子的成熟受其调控,包括与心脏发育相关的一些转录因子都被证实是其作用底物[91-93]。DUPAYS等[94]研究发现Furin为心脏前体细胞的成熟和房室连接的正确形成所必需,同时发现Furin被NKX2-5直接抑制。
(6)GATA1:GATA因子可被分类为GATA-1/2/3和GATA-4/5/6,其中GATA-1/2/3参与造血分化、红细胞分化、心脏前体细胞增殖和T细胞分化,而GATA-4/5/6则在心脏形成中起作用[95]。NKX2-5可通过结合于Gata1远端的NKE元件而抑制GATA1表达[96]。
(7)Isl1:Isl1属于LIM同源性转录因子,在心脏第二生心区前体细胞的存活、增殖、迁移中起关键性的调控作用,可调控前体细胞迁移到第一生心区形成的原始心管结构上促进其延长和形成心脏结构[97]。DORN等[28]研究发现NKX2-5直接结合于Isi1的增强子区域而抑制其转录激活。
| [1] SCHLEICH JM, ABDULLA T, SUMMERS R, et al. An overview of cardiac morphogenesis. Arch Cardiovasc Dis. 2013;106(11): 612-623. [2] SRIVASTAVA D. Making or breaking the heart: from lineage determination to morphogenesis. Cell. 2006;126(6):1037-1048. [3] ZHANG L, NOMURA-KITABAYASHI A, SULTANA N, et al. Mesodermal Nkx2.5 is necessary and sufficient for early second heart field development. Dev Biol. 2014;390(1):68-79. [4] MEGANATHAN K, SOTIRIADOU I, NATARAJAN K, et al. Signaling molecules, transcription growth factors and other regulators revealed from in-vivo and in-vitro models for the regulation of cardiac development. Int J Cardiol. 2015;183:117-128. [5] OLSON EN. Gene regulatory networks in the evolution and development of the heart. Science. 2006;313(5795):1922-1927. [6] SCHOTT JJ, BENSON DW, BASSON CT, et al. Congenital heart disease caused by mutations in the transcription factor NKX2-5. Science. 1998; 281(5373):108-111. [7] HARVEY RP. NK-2 homeobox genes and heart development. Dev Biol. 1996;178(2):203-216. [8] SHIOJIMA I, KOMURO I, MIZUNO T, et al. Molecular cloning and characterization of human cardiac homeobox gene CSX1. Circ Res. 1996;79(5):920-929. [9] KASAHARA H, LEE B, SCHOTT JJ, et al. Loss of function and inhibitory effects of human CSX/NKX2.5 homeoprotein mutations associated with congenital heart disease. J Clin Invest. 2000; 106(2):299-308. [10] AKAZAWA H, KOMURO I. Cardiac transcription factor Csx/Nkx2-5: Its role in cardiac development and diseases. Pharmacol Ther. 2005; 107(2):252-268. [11] WATADA H, MIRMIRA RG, KALAMARAS J, et al. Intramolecular control of transcriptional activity by the NK2-specific domain in NK-2 homeodomain proteins. Proc Natl Acad Sci U S A. 2000; 97(17): 9443-9448. [12] ELLIOTT DA, SOLLOWAY MJ, WISE N, et al. A tyrosine-rich domain within homeodomain transcription factor Nkx2-5 is an essential element in the early cardiac transcriptional regulatory machinery. Development. 2006;133(7):1311-1322. [13] BOUVERET R, WAARDENBERG AJ, SCHONROCK N, et al. NKX2-5 mutations causative for congenital heart disease retain functionality and are directed to hundreds of targets. Elife. 2015;4. [14] TURBAY D, WECHSLER SB, BLANCHARD KM, et al. Molecular cloning, chromosomal mapping, and characterization of the human cardiac-specific homeobox gene hCsx. Mol Med. 1996; 2(1):86-96. [15] BRUNEAU BG. The developmental genetics of congenital heart disease. Nature. 2008;451(7181):943-948. [16] LINTS TJ, PARSONS LM, HARTLEY L, et al. Nkx-2.5: a novel murine homeobox gene expressed in early heart progenitor cells and their myogenic descendants. Development. 1993;119(2): 419-431. [17] SERPOOSHAN V, LIU YH, BUIKEMA JW, et al. Nkx2.5+ Cardiomyoblasts Contribute to Cardiomyogenesis in the Neonatal Heart. Sci Rep. 2017; 7(1):12590. [18] WU SM, FUJIWARA Y, CIBULSKY SM, et al. Developmental origin of a bipotential myocardial and smooth muscle cell precursor in the mammalian heart. Cell. 2006;127(6):1137-1150. [19] ZAMIR L, SINGH R, NATHAN E, et al. Nkx2.5 marks angioblasts that contribute to hemogenic endothelium of the endocardium and dorsal aorta. Elife. 2017;6: e20994. [20] PAFFETT-LUGASSY N, SINGH R, NEVIS KR, et al. Heart field origin of great vessel precursors relies on nkx2.5-mediated vasculogenesis. Nat Cell Biol. 2013;15(11):1362-1369. [21] CLARK CD, ZHANG B, LEE B, et al. Evolutionary conservation of Nkx2.5 autoregulation in the second heart field. Dev Biol. 2013; 374(1): 198-209. [22] PRALL OW, MENON MK, SOLLOWAY MJ, et al. An Nkx2-5/ Bmp2/Smad1 negative feedback loop controls heart progenitor specification and proliferation. Cell. 2007;128(5): 947-959. [23] COLOMBO S, DE SENA-TOMÁS C, GEORGE V, et al. Nkx genes establish second heart field cardiomyocyte progenitors at the arterial pole and pattern the venous pole through Isl1 repression. Development. 2018;145(3): dev161497. [24] GEORGE V, COLOMBO S, TARGOFF KL. An early requirement for nkx2.5 ensures the first and second heart field ventricular identity and cardiac function into adulthood. Dev Biol. 2015;400(1):10-22. [25] ARMIÑÁN A, GANDÍA C, BARTUAL M, et al. Cardiac differentiation is driven by NKX2.5 and GATA4 nuclear translocation in tissue-specific mesenchymal stem cells. Stem Cells Dev. 2009;18(6):907-918. [26] RYAN T, SHELTON M, LAMBERT JP, et al. Myosin phosphatase modulates the cardiac cell fate by regulating the subcellular localization of Nkx2.5 in a Wnt/Rho-associated protein kinase-dependent pathway. Circ Res. 2013;112(2):257-266. [27] NAKASHIMA Y, YANEZ DA, TOUMA M, et al. Nkx2-5 suppresses the proliferation of atrial myocytes and conduction system. Circ Res. 2014; 114(7):1103-1113. [28] DORN T, GOEDEL A, LAM JT, et al. Direct nkx2-5 transcriptional repression of isl1 controls cardiomyocyte subtype identity. Stem Cells. 2015;33(4):1113-1129. [29] RUAN Z, ZHU L, YIN Y, et al. Overexpressing NKx2.5 increases the differentiation of human umbilical cord drived mesenchymal stem cells into cardiomyocyte-like cells. Biomed Pharmacother. 2016;78:110-115. [30] DENG B, WANG JX, HU XX, et al. Nkx2.5 enhances the efficacy of mesenchymal stem cells transplantation in treatment heart failure in rats. Life Sci. 2017;182:65-72. [31] JAY PY, HARRIS BS, BUERGER A, et al. Function follows form: cardiac conduction system defects in Nkx2-5 mutation. Anat Rec A Discov Mol Cell Evol Biol. 2004;280(2):966-972. [32] BRIGGS LE, TAKEDA M, CUADRA AE, et al. Perinatal loss of Nkx2-5 results in rapid conduction and contraction defects. Circ Res. 2008; 103(6):580-590. [33] TAKEDA M, BRIGGS LE, WAKIMOTO H, et al. Slow progressive conduction and contraction defects in loss of Nkx2-5 mice after cardiomyocyte terminal differentiation. Lab Invest. 2009;89(9): 983-993. [34] PASHMFOROUSH M, LU JT, CHEN H, et al. Nkx2-5 pathways and congenital heart disease; loss of ventricular myocyte lineage specification leads to progressive cardiomyopathy and complete heart block. Cell. 2004;117(3):373-386. [35] HARRINGTON JK, SORABELLA R, TERCEK A, et al. Nkx2.5 is essential to establish normal heart rate variability in the zebrafish embryo. Am J Physiol Regul Integr Comp Physiol. 2017;313(3): R265-R271. [36] XIE WH, CHANG C, XU YJ, et al. Prevalence and spectrum of Nkx2.5 mutations associated with idiopathic atrial fibrillation. Clinics (Sao Paulo). 2013;68(6):777-784. [37] PERERA JL, JOHNSON NM, JUDGE DP, et al. Novel and highly lethal NKX2.5 missense mutation in a family with sudden death and ventricular arrhythmia. Pediatr Cardiol. 2014;35(7):1206-1212. [38] MONACO I, SANTACROCE R, CASAVECCHIA G, et al. Double de novo mutations in dilated cardiomyopathy with cardiac arrest. J Electrocardiol. 2019;53:40-43. [39] ANDERSON DJ, KAPLAN DI, BELL KM, et al. NKX2-5 regulates human cardiomyogenesis via a HEY2 dependent transcriptional network. Nat Commun. 2018;9(1):1373. [40] NIMURA K, YAMAMOTO M, TAKEICHI M, et al. Regulation of alternative polyadenylation by Nkx2-5 and Xrn2 during mouse heart development. Elife. 2016;5: e16030. [41] DUPAYS L, SHANG C, WILSON R, et al. Sequential Binding of MEIS1 and NKX2-5 on the Popdc2 Gene: A Mechanism for Spatiotemporal Regulation of Enhancers during Cardiogenesis. Cell Rep. 2015;13(1): 183-195. [42] BENSON DW, SILBERBACH GM, KAVANAUGH-MCHUGH A, et al. Mutations in the cardiac transcription factor NKX2.5 affect diverse cardiac developmental pathways. J Clin Invest. 1999; 104(11):1567-1573. [43] CHUNG IM, RAJAKUMAR G. Genetics of Congenital Heart Defects: The NKX2-5 Gene, a Key Player. Genes (Basel). 2016;7(2): E6. [44] GOLDMUNTZ E, GEIGER E, BENSON DW. NKX2.5 mutations in patients with tetralogy of fallot. Circulation. 2001;104(21): 2565-2568. [45] KÖNIG K, WILL JC, BERGER F, et al. Familial congenital heart disease, progressive atrioventricular block and the cardiac homeobox transcription factor gene NKX2.5: identification of a novel mutation. Clin Res Cardiol. 2006;95(9):499-503. [46] OUYANG P, SAAREL E, BAI Y, et al. A de novo mutation in NKX2.5 associated with atrial septal defects, ventricular noncompaction, syncope and sudden death. Clin Chim Acta. 2011;412(1-2):170-175. [47] QU XK, QIU XB, YUAN F, et al. A novel NKX2.5 loss-of-function mutation associated with congenital bicuspid aortic valve. Am J Cardiol. 2014; 114(12):1891-1895. [48] BIBEN C, WEBER R, KESTEVEN S, et al. Cardiac septal and valvular dysmorphogenesis in mice heterozygous for mutations in the homeobox gene Nkx2-5. Circ Res. 2000;87(10):888-895. [49] LIU XY, WANG J, YANG YQ, et al. Novel NKX2-5 mutations in patients with familial atrial septal defects. Pediatr Cardiol. 2011; 32(2):193-201. [50] IKEDA Y, HIROI Y, HOSODA T, et al. Novel point mutation in the cardiac transcription factor CSX/NKX2.5 associated with congenital heart disease. Circ J. 2002;66(6):561-563. [51] ZAKARIYAH AF, RAJGARA RF, VEINOT JP, et al. Congenital heart defect causing mutation in Nkx2.5 displays in vivo functional deficit. J Mol Cell Cardiol. 2017;105:89-98. [52] ELLESØE SG, JOHANSEN MM, BJERRE JV, et al. Familial Atrial Septal Defect and Sudden Cardiac Death: Identification of a Novel NKX2-5 Mutation and a Review of the Literature. Congenit Heart Dis. 2016; 11(3):283-290. [53] PABST S, WOLLNIK B, ROHMANN E, et al. A novel stop mutation truncating critical regions of the cardiac transcription factor NKX2-5 in a large family with autosomal-dominant inherited congenital heart disease. Clin Res Cardiol. 2008;97(1):39-42. [54] HUANG W, MENG H, QIAO Y, et al. Two novel and functional DNA sequence variants within an upstream enhancer of the human NKX2-5 gene in ventricular septal defects. Gene. 2013; 524(2):152-155. [55] WATANABE Y, BENSON DW, YANO S, et al. Two novel frameshift mutations in NKX2.5 result in novel features including visceral inversus and sinus venosus type ASD. J Med Genet. 2002;39(11):807-811. [56] DINESH SM, KUSUMA L, SMITHA R, et al. Single-nucleotide polymorphisms of NKX2.5 found in congenital heart disease patients of Mysore, South India. Genet Test Mol Biomarkers. 2010;14(6):873-879. [57] CAO Y, LAN W, LI Y, et al. Single nucleotide polymorphism of NKX2-5 gene with sporadic congenital heart disease in Chinese Bai population. Int J Clin Exp Pathol. 2015;8(11):14917-14924. [58] MONZEN K, ZHU W, KASAI H, et al. Dual effects of the homeobox transcription factor Csx/Nkx2-5 on cardiomyocytes. Biochem Biophys Res Commun. 2002;298(4):493-500. [59] TOKO H, ZHU W, TAKIMOTO E, et al. Csx/Nkx2-5 is required for homeostasis and survival of cardiac myocytes in the adult heart. J Biol Chem. 2002;277(27):24735-24743. [60] RIAZI AM, LEE H, HSU C, et al. CSX/Nkx2.5 modulates differentiation of skeletal myoblasts and promotes differentiation into neuronal cells in vitro. J Biol Chem. 2005;280(11): 10716-10720. [61] UDAGER AM, PRAKASH A, SAENZ DA, et al. Proper development of the outer longitudinal smooth muscle of the mouse pylorus requires Nkx2-5 and Gata3. Gastroenterology. 2014;146(1):157-165.e10. [62] HWANG C, JANG S, CHOI DK, et al. The role of nkx2.5 in keratinocyte differentiation. Ann Dermatol. 2009;21(4):376-381. [63] DRITSOULA A, PAPAIOANNOU I, GUERRA SG, et al. Molecular Basis for Dysregulated Activation of NKX2-5 in the Vascular Remodeling of Systemic Sclerosis. Arthritis Rheumatol. 2018;70(6):920-931. [64] VAN ENGELEN K, MOMMERSTEEG MT, BAARS MJ, et al. The ambiguous role of NKX2-5 mutations in thyroid dysgenesis. PLoS One. 2012;7(12):e52685. [65] PENHA RCC, BUEXM LA, RODRIGUES FR, et al. NKX2.5 is expressed in papillary thyroid carcinomas and regulates differentiation in thyroid cells. BMC Cancer. 2018;18(1):498. [66] NAGEL S, KAUFMANN M, DREXLER HG, et al. The cardiac homeobox gene NKX2-5 is deregulated by juxtaposition with BCL11B in pediatric T-ALL cell lines via a novel t(5;14)(q35.1;q32.2). Cancer Res. 2003;63(17):5329-5334. [67] SHIBATA K, KAJIYAMA H, YAMAMOTO E, et al. Establishment and characterization of an ovarian yolk sac tumor cell line reveals possible involvement of Nkx2.5 in tumor development. Oncology. 2008;74 (1-2):104-111. [68] WU M, PENG S, YANG J, et al. Baf250a orchestrates an epigenetic pathway to repress the Nkx2.5-directed contractile cardiomyocyte program in the sinoatrial node. Cell Res. 2014;24(10):1201-1213. [69] RISEBRO CA, SEARLES RG, MELVILLE AA, et al. Prox1 maintains muscle structure and growth in the developing heart. Development. 2009;136(3):495-505. [70] RISEBRO CA, PETCHEY LK, SMART N, et al. Epistatic rescue of Nkx2.5 adult cardiac conduction disease phenotypes by prospero-related homeobox protein 1 and HDAC3. Circ Res. 2012;111(2):e19-31. [71] SNYDER M, HUANG XY, ZHANG JJ. Stat3 directly controls the expression of Tbx5, Nkx2.5, and GATA4 and is essential for cardiomyocyte differentiation of P19CL6 cells. J Biol Chem. 2010;285(31): 23639-23646. [72] SULTANA N, NAG K, HOSHIJIMA K, et al. Zebrafish early cardiac connexin, Cx36.7/Ecx, regulates myofibril orientation and heart morphogenesis by establishing Nkx2.5 expression. Proc Natl Acad Sci U S A. 2008;105(12):4763-4768. [73] JODOIN R, CARRIER JC, RIVARD N, et al. G-quadruplex located in the 5’UTR of the BAG-1 mRNA affects both its cap-dependent and cap-independent translation through global secondary structure maintenance. Nucleic Acids Res. 2019;47(19):10247-10266. [74] NIE J, JIANG M, ZHANG X, et al. Post-transcriptional Regulation of Nkx2-5 by RHAU in Heart Development. Cell Rep. 2015;13(4): 723-732. [75] WANG J, ZHANG H, IYER D, et al. Regulation of cardiac specific nkx2.5 gene activity by small ubiquitin-like modifier. J Biol Chem. 2008; 283(34):23235-23243. [76] TANG X, MA H, HAN L, et al. SIRT1 deacetylates the cardiac transcription factor Nkx2.5 and inhibits its transcriptional activity. Sci Rep. 2016;6:36576. [77] COSTA MW, LEE S, FURTADO MB, et al. Complex SUMO-1 regulation of cardiac transcription factor Nkx2-5. PLoS One. 2011;6(9):e24812. [78] RAMACHANDRA CJ, MEHTA A, WONG P, et al. ErbB4 Activated p38γ MAPK Isoform Mediates Early Cardiogenesis Through NKx2.5 in Human Pluripotent Stem Cells. Stem Cells. 2016;34(2):288-298. [79] PRADHAN A, ZENG XI, SIDHWANI P, et al. FGF signaling enforces cardiac chamber identity in the developing ventricle. Development. 2017;144(7):1328-1338. [80] KAY M, SOLTANI BM, AGHDAEI FH, et al. Hsa-miR-335 regulates cardiac mesoderm and progenitor cell differentiation. Stem Cell Res Ther. 2019; 10(1):191. [81] DUROCHER D, CHARRON F, WARREN R, et al. The cardiac transcription factors Nkx2-5 and GATA-4 are mutual cofactors. EMBO J. 1997;16(18): 5687-5696. [82] LEE Y, SHIOI T, KASAHARA H, et al. The cardiac tissue-restricted homeobox protein Csx/Nkx2.5 physically associates with the zinc finger protein GATA4 and cooperatively activates atrial natriuretic factor gene expression. Mol Cell Biol. 1998;18(6):3120-3129. [83] KINNUNEN S, VÄLIMÄKI M, TÖLLI M, et al. Nuclear Receptor-Like Structure and Interaction of Congenital Heart Disease-Associated Factors GATA4 and NKX2-5. PLoS One. 2015;10(12):e0144145. [84] HIROI Y, KUDOH S, MONZEN K, et al. Tbx5 associates with Nkx2-5 and synergistically promotes cardiomyocyte differentiation. Nat Genet. 2001;28(3):276-280. [85] KOJIC S, NESTOROVIC A, RAKICEVIC L, et al. Cardiac transcription factor Nkx2.5 interacts with p53 and modulates its activity. Arch Biochem Biophys. 2015;569:45-53. [86] CHEN CY, SCHWARTZ RJ. Recruitment of the tinman homolog Nkx-2.5 by serum response factor activates cardiac alpha-actin gene transcription. Mol Cell Biol. 1996;16(11):6372-6384. [87] CAMBIER L, PLATE M, SUCOV HM, et al. Nkx2-5 regulates cardiac growth through modulation of Wnt signaling by R-spondin3. Development. 2014;141(15):2959-2971. [88] WARREN SA, TERADA R, BRIGGS LE, et al. Differential role of Nkx2-5 in activation of the atrial natriuretic factor gene in the developing versus failing heart. Mol Cell Biol. 2011;31(22): 4633-4645. [89] RIAZI AM, TAKEUCHI JK, HORNBERGER LK, et al. NKX2-5 regulates the expression of beta-catenin and GATA4 in ventricular myocytes. PLoS One. 2009;4(5):e5698. [90] WANG J, JIN Y, CATTINI PA. Expression of the Cardiac Maintenance and Survival Factor FGF-16 Gene Is Regulated by Csx/Nkx2.5 and Is an Early Target of Doxorubicin Cardiotoxicity. DNA Cell Biol. 2017;36(2): 117-126. [91] THOMAS G. Furin at the cutting edge: from protein traffic to embryogenesis and disease. Nat Rev Mol Cell Biol. 2002;3(10):753-766. [92] CUI Y, HACKENMILLER R, BERG L, et al. The activity and signaling range of mature BMP-4 is regulated by sequential cleavage at two sites within the prodomain of the precursor. Genes Dev. 2001;15(21):2797-2802. [93] CUI Y, JEAN F, THOMAS G, et al. BMP-4 is proteolytically activated by furin and/or PC6 during vertebrate embryonic development. EMBO J. 1998;17(16):4735-4743. [94] DUPAYS L, TOWERS N, WOOD S, et al. Furin, a transcriptional target of NKX2-5, has an essential role in heart development and function. PLoS One. 2019;14(3):e0212992. [95] PATIENT RK, MCGHEE JD. The GATA family (vertebrates and invertebrates). Curr Opin Genet Dev. 2002;12(4):416-422. [96] CAPRIOLI A, KOYANO-NAKAGAWA N, IACOVINO M, et al. Nkx2-5 represses Gata1 gene expression and modulates the cellular fate of cardiac progenitors during embryogenesis. Circulation. 2011; 123(15): 1633-1641. [97] CAI CL, LIANG X, SHI Y, et al. Isl1 identifies a cardiac progenitor population that proliferates prior to differentiation and contributes a majority of cells to the heart. Dev Cell. 2003;5(6):877-889. |
| [1] | 蒲 锐, 陈子扬, 袁凌燕. 不同细胞来源外泌体保护心脏的特点与效应[J]. 中国组织工程研究, 2021, 25(在线): 1-. |
| [2] | 林清凡, 解一新, 陈婉清, 叶振忠, 陈幼芳. 人胎盘源间充质干细胞条件培养液可上调缺氧状态下BeWo细胞活力和紧密连接因子的表达[J]. 中国组织工程研究, 2021, 25(在线): 4970-4975. |
| [3] | 张 宇, 田少奇, 曾国波, 胡 川. 初次下肢全关节置换后发生心肌梗死的危险因素[J]. 中国组织工程研究, 2021, 25(9): 1340-1345. |
| [4] | 顾 霞, 赵 敏, 王平义, 李一梅, 李文华. 低氧诱导因子1α与低氧相关疾病信号通路的关系[J]. 中国组织工程研究, 2021, 25(8): 1284-1289. |
| [5] | 姬志祥, 蓝常贡. 尿酸盐转运蛋白在痛风中的多态性和治疗相关性[J]. 中国组织工程研究, 2021, 25(8): 1290-1298. |
| [6] | 吴 训, 孟娟红, 张建运, 王 亮. 浓缩生长因子修复兔髁突全层软骨损伤[J]. 中国组织工程研究, 2021, 25(8): 1166-1171. |
| [7] | 刘志超, 张 帆, 孙 旗, 康晓乐, 袁巧妹, 柳根哲, 陈 江. 不同静水压下人椎间盘髓核细胞的形态和活性[J]. 中国组织工程研究, 2021, 25(8): 1172-1176. |
| [8] | 申晋波, 张 林. 急性力竭运动导致大鼠跟腱微损伤的超微结构变化及机制[J]. 中国组织工程研究, 2021, 25(8): 1190-1195. |
| [9] | 李 静, 谢建山, 崔慧林, 曹锡梅, 杨艳萍, 李海荣. 不同毛色小鼠背部皮肤中甘油二酯激酶ζ和蛋白激酶CβⅡ的表达和定位[J]. 中国组织工程研究, 2021, 25(8): 1196-1200. |
| [10] | 李嘉程, 梁学振, 刘金豹, 许 波, 李 刚. 骨性关节炎mRNA差异表达谱及竞争性内源RNA调控的网络分析[J]. 中国组织工程研究, 2021, 25(8): 1212-1217. |
| [11] | 柴 乐, 吕建兰, 胡劲涛, 胡华辉, 许庆军, 余进伟, 全仁夫. 诱导急性脊髓损伤模型大鼠炎症反应信号通路的变化[J]. 中国组织工程研究, 2021, 25(8): 1218-1223. |
| [12] | 耿秋东, 葛海雅, 王和鸣, 李 楠. 基于网络药理学探讨龟鹿二仙胶治疗骨关节炎的作用及机制[J]. 中国组织工程研究, 2021, 25(8): 1229-1236. |
| [13] | 刘翔翔, 黄云梅, 陈文列, 林如辉, 卢小冬, 李钻芳, 许亚晔, 黄美雅, 李西海. 早期骨关节炎模型大鼠半月板白区细胞的超微结构变化[J]. 中国组织工程研究, 2021, 25(8): 1237-1242. |
| [14] | 谭婧玉, 刘海文. 成骨细胞特异因子2基因家族的全基因组鉴定、分类及进化分析[J]. 中国组织工程研究, 2021, 25(8): 1243-1248. |
| [15] | 张秀梅, 翟运开, 赵 杰, 赵 萌. 类器官模型国内外数据库近10年文献研究热点分析[J]. 中国组织工程研究, 2021, 25(8): 1249-1255. |
胞[1]。心脏前体细胞的增殖、分化、迁移,形成了原始的心管结构、心管的环化折叠和心脏间隔的形成,最终形成完整的功能性四腔室的心脏结构。
胚胎期心脏发育是一个复杂的生物学过程。在这个过程中,转录因子、信号通路、基因表观遗传学因素及小分子RNA都参与心脏发育过程的调控[4-5]。在这些因素中,转录因子NKX2-5是调控心脏发育的关键分子之一,其功能异常被认为与先天性心脏病尤其是房间隔缺损密切相关[6]。但到目前为止,NKX2-5功能异常导致先天性心脏病发生的机制依然不清。因此,作者在查阅相关文献后对NKX2-5的分子结构、功能作用、上下游调控分子进行综述。
中国组织工程研究杂志出版内容重点:干细胞;骨髓干细胞;造血干细胞;脂肪干细胞;肿瘤干细胞;胚胎干细胞;脐带脐血干细胞;干细胞诱导;干细胞分化;组织工程
1.2 文献筛选标准 排除与NKX2-5无关的文献,观点重复、结论一致的文献。
1.3 质量评估及数据的提取 计算机初检得到803篇文献,经资料收集者互相评估纳入文献的有效性和适用性,通过阅读文题和摘要进行初步筛选;排除重复性研究以及内容不相关的文献,最后纳入97篇文献进行综述。
中国组织工程研究杂志出版内容重点:干细胞;骨髓干细胞;造血干细胞;脂肪干细胞;肿瘤干细胞;胚胎干细胞;脐带脐血干细胞;干细胞诱导;干细胞分化;组织工程

文题释义:#br# 转录因子:本质为蛋白质,其DNA序列经转录、翻译后形成蛋白质,可进入细胞核中,结合于下游基因启动子区的特定DNA序列,激活或抑制其调控的下游基因的转录,从而调控该基因功能。#br#
先天性心脏病:是一类因胚胎期心脏发育异常而导致的出生缺陷性疾病,根据病变位置,可分为房间隔缺损、室间隔缺损、动脉导管未闭、法洛氏四联症等临床常见亚型。因心脏结构异常,导致血流动力学紊乱,是引起新生儿死亡的常见疾病。
中国组织工程研究杂志出版内容重点:干细胞;骨髓干细胞;造血干细胞;脂肪干细胞;肿瘤干细胞;胚胎干细胞;脐带脐血干细胞;干细胞诱导;干细胞分化;组织工程
#br#
转录因子NKX2-5是哺乳动物心脏发育过程中极其重要的一个调节因子。来自于模式动物如大鼠、小鼠及斑马鱼的研究表明,NKX2-5的缺失或功能异常,影响心脏发育,导致心脏发育异常,出现类似于人类先天性心脏病的病理表现。然而,对于NKX2-5在心脏发育过程中的调控作用及具体机制,目前依然不明确。近年来的诸多研究表明,NKX2-5的位点变异如rs2277923、rs3729753与先天性心脏病如房间隔缺损、心律失常等的发生密切相关,因此深入了解转录因子NKX2-5的研究与进展有助于相关领域的研究人员把握和了解房间隔缺损发生机制的研究动态和方向。
中国组织工程研究杂志出版内容重点:干细胞;骨髓干细胞;造血干细胞;脂肪干细胞;肿瘤干细胞;胚胎干细胞;脐带脐血干细胞;干细胞诱导;干细胞分化;组织工程
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||