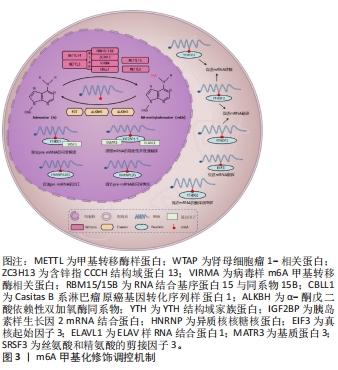
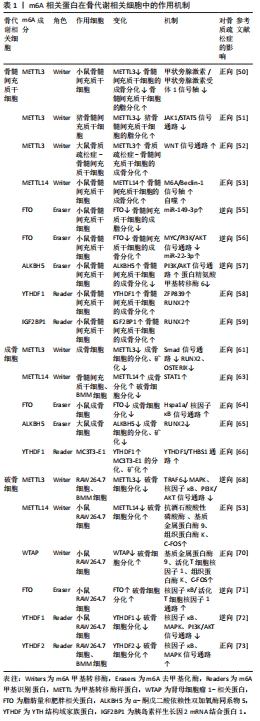
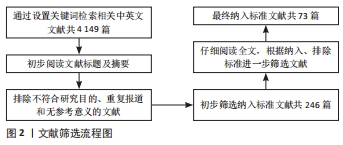
[1] ENSRUD KE, CRANDALL CJ. Osteoporosis. Ann Intern Med. 2017;167(3):ITC17-ITC32.
[2] BROWN C. Osteoporosis: Staying strong. Nature. 2017;550(7674):S15-S17.
[3] RACHNER TD, KHOSLA S, HOFBAUER LC. Osteoporosis: now and the future. Lancet. 2011;377(9773):1276-1287.
[4] TYSOE O. Liver-bone crosstalk implicated in osteoporosis progression. Nat Rev Endocrinol. 2023;19(8):440.
[5] YANG C, DONG Z, LING Z, et al. The crucial mechanism and therapeutic implication of RNA methylation in bone pathophysiology. Ageing Res Rev. 2022;79:101641.
[6] DAI X, REN T, ZHANG Y, et al. Methylation multiplicity and its clinical values in cancer. Expert Rev Mol Med. 2021;23:e2.
[7] JONKHOUT N, TRAN J, SMITH MA, et al. The RNA modification landscape in human disease. RNA. 2017;23(12):1754-1769.
[8] DESROSIERS R, FRIDERICI K, ROTTMAN F. Identification of Methylated Nucleosides in Messenger RNA from Novikoff Hepatoma Cells. Proc National Acad Sci. 1974;71(10): 3971-3975.
[9] BATISTA PJ, MOLINIE B, WANG J, et al. m(6)A RNA modification controls cell fate transition in mammalian embryonic stem cells. Cell Stem Cell. 2014;15(6):707-719.
[10] YANG Y, HSU PJ, CHEN YS, et al. Dynamic transcriptomic m(6)A decoration: writers, erasers, readers and functions in RNA metabolism. Cell Res. 2018;28(6):616-624.
[11] SUN T, WU R, MING L. The role of m6A RNA methylation in cancer. Biomed Pharmacother. 2019;112:108613.
[12] HAN M, LIU Z, XU Y, et al. Abnormality of m6A mRNA Methylation Is Involved in Alzheimer’s Disease. Front Neurosci. 2020;14:98.
[13] FU J, CUI X, ZHANG X, et al. The Role of m6A Ribonucleic Acid Modification in the Occurrence of Atherosclerosis. Front Genet. 2021;12:733871.
[14] NIE K, YIJ, YANG Y, et al. A Broad m6A Modification Landscape in Inflammatory Bowel Disease. Front Cell Dev Biol. 2021;9:782636.
[15] HUANG H, WENG H, CHEN J. The Biogenesis and Precise Control of RNA m6A Methylation. Trends Genet. 2020;36(1):44-52.
[16] ZHANG Y, GENG X, LI Q, et al. m6A modification in RNA: biogenesis, functions and roles in gliomas. J Exp Clin Cancer Res. 2020;39(1):192.
[17] AN Y, DUAN H. The role of m6A RNA methylation in cancer metabolism. Mol Cancer. 2022;21(1):14.
[18] SHI H, WEI J, HE C. Where, When, and How: Context-Dependent Functions of RNA Methylation Writers, Readers, and Erasers. Mol Cell. 2019;74(4):640-650.
[19] OERUM S, MEYNIER V, CATALA M, et al. A comprehensive review of m6A/m6Am RNA methyltransferase structures. Nucleic Acids Res. 2021;49(13):7239-7255.
[20] YANG B, WANG JQ, TAN Y, et al. RNA methylation and cancer treatment. Pharmacol Res. 2021;174:105937.
[21] SCHOLLER E, WEICHMANN F, TREIBER T, et al. Interactions, localization, and phosphorylation of the m(6)A generating METTL3-METTL14-WTAP complex. RNA. 2018; 24(4):499-512.
[22] CHEN J, FANG Y, XU Y, et al. Role of m6A modification in female infertility and reproductive system diseases. Int J Biol Sci. 2022;18(9):3592-3604.
[23] BUJNICKI JM, FEDER M, RADLINSKA M, et al. Structure prediction and phylogenetic analysis of a functionally diverse family of proteins homologous to the MT-A70 subunit of the human mRNA:m(6)A methyltransferase. J Mol Evol. 2002;55(4):431-444.
[24] PING XL, SUN BF, WANG L, et al. Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Res. 2014;24(2):177-189.
[25] LIU J, YUE Y, HAN D, et al. A METTL3-METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation. Nat Chem Biol. 2014;10(2):93-95.
[26] WEN J, LV R, MA H, et al. Zc3h13 Regulates Nuclear RNA m(6)A Methylation and Mouse Embryonic Stem Cell Self-Renewal. Mol Cell. 2018;69(6):1028-1038.
[27] ZHANG X, LI MJ, XIA L, et al. The biological function of m6A methyltransferase KIAA1429 and its role in human disease. Peer J. 2022;10:e14334.
[28] PATIL DP, CHEN CK, PICKERING BF, et al. m(6)A RNA methylation promotes XIST-mediated transcriptional repression. Nature. 2016;537(7620):369-373.
[29] WANG Y, ZHANG L, REN H, et al. Role of Hakai in m(6)A modification pathway in Drosophila. Nat Commun. 2021;12(1):2159.
[30] FANG Z, MEI W, QU C, et al. Role of m6A writers, erasers and readers in cancer. Exp Hematol Oncol. 2022;11(1):45.
[31] GAO R, YE M, LIU B, et al. m6A Modification: A Double-Edged Sword in Tumor Development. Front Oncol. 2021;11:679367.
[32] FU Y, JIA G, PANG X, et al. FTO-mediated formation of N6-hydroxymethyladenosine and N6-formyladenosine in mammalian RNA. Nat Commun. 2013;4:1798.
[33] JIA G, FU Y, ZHAO X, et al. N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat Chem Biol. 2011;7(12):885-887.
[34] ZHENG G, DAHL JA, NIU Y, et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol Cell. 2013;49(1):18-29.
[35] UEDA Y, OOSHIO I, FUSAMAE Y, et al. AlkB homolog 3-mediated tRNA demethylation promotes protein synthesis in cancer cells. Sci Rep. 2017;7:42271.
[36] LIU C, GU L, DENG W, et al. N6-Methyladenosine RNA Methylation in Cardiovascular Diseases. Front Cardiovasc Med. 2022;9:887838.
[37] LIAO S, SUN H, XU C. YTH Domain: A Family of N(6)-methyladenosine (m(6)A) Readers. Genomics Proteomics Bioinformatics. 2018;16(2):99-107.
[38] ZACCARA S, JAFFREY SR. A Unified Model for the Function of YTHDF Proteins in Regulating m(6)A-Modified mRNA. Cell. 2020;181(7):1582-1595.
[39] XU Y, ZHANG W, SHEN F, et al. YTH Domain Proteins: A Family of m(6)A Readers in Cancer Progression. Front Oncol. 2021;11:629560.
[40] DEGRAUWE N, SUVA ML, JANISZEWSKA M, et al. IMPs: an RNA-binding protein family that provides a link between stem cell maintenance in normal development and cancer. Genes Dev. 2016;30(22):2459-2474.
[41] HUANG H, WENG H, SUN W, et al. Recognition of RNA N(6)-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat Cell Bio. 2018;20(3):285-295.
[42] ALARCON CR, GOODARZI H, LEE H, et al. HNRNPA2B1 Is a Mediator of m(6)A-Dependent Nuclear RNA Processing Events. Cell. 2015;162(6):1299-1308.
[43] LIU N, DAI Q, ZHENG G, et al. N(6)-methyladenosine-dependent RNA structural switches regulate RNA-protein interactions. Nature. 2015;518(7540):560-564.
[44] LIU N, ZHOU KI, PARISIEN M, et al. N6-methyladenosine alters RNA structure to regulate binding of a low-complexity protein. Nucleic Acids Res. 2017;45(10):6051-6063.
[45] MEYER KD, PATIL DP, ZHOU J, et al. 5’ UTR m(6)A Promotes Cap-Independent Translation. Cell. 2015;163(4):999-1010.
[46] ZHANG L, ZHENG YL, WANG R, et al. Exercise for osteoporosis: A literature review of pathology and mechanism. Front Immunol. 2022;13:1005665.
[47] HOCKING LJ, WHITEHOUSE C, HELFRICH MH. Autophagy: a new player in skeletal maintenance? J Bone Miner Res. 2012;27(7):1439-1447.
[48] NOMBELA-ARRIETA C, RITZ J, SILBERSTEIN LE. The elusive nature and function of mesenchymal stem cells. Nat Rev Mol Cell Biol. 2011;12(2):126-131.
[49] RICKARD DJ, WANG FL, RODRIGUEZ-ROJAS AM, et al. Intermittent treatment with parathyroid hormone (PTH) as well as a non-peptide small molecule agonist of the PTH1 receptor inhibits adipocyte differentiation in human bone marrow stromal cells. Bone. 2006;39(6):1361-1372.
[50] WU Y, XIE L, WANG M, et al. Mettl3-mediated m(6)A RNA methylation regulates the fate of bone marrow mesenchymal stem cells and osteoporosis. Nat Commun. 2018;9(1):4772.
[51] YAO Y, BI Z, WU R, et al. METTL3 inhibits BMSC adipogenic differentiation by targeting the JAK1/STAT5/C/EBPbeta pathway via an m(6)A-YTHDF2-dependent manner. FASEB J. 2019;33(6):7529-7544.
[52] WU T, TANG H, YANG J, et al. METTL3-m(6) A methylase regulates the osteogenic potential of bone marrow mesenchymal stem cells in osteoporotic rats via the Wnt signalling pathway. Cell Prolif. 2022;55(5):e13234.
[53] HE M, LEI H, HE X, et al. METTL14 Regulates Osteogenesis of Bone Marrow Mesenchymal Stem Cells via Inducing Autophagy Through m6A/IGF2BPs/Beclin-1 Signal Axis. Stem Cells Transl Med. 2022;11(9):987-1001.
[54] SHEN GS, ZHOU HB, ZHANG H, et al. The GDF11-FTO-PPARgamma axis controls the shift of osteoporotic MSC fate to adipocyte and inhibits bone formation during osteoporosis. Biochim Biophys Acta Mol Basis Dis. 2018;1864(12):3644-3654.
[55] LI Y, YANG F, GAO M, et al. miR-149-3p Regulates the Switch between Adipogenic and Osteogenic Differentiation of BMSCs by Targeting FTO. Mol Ther Nucleic Acids. 2019;17:590-600.
[56] ZHANG X, WANG Y, ZHAO H, et al. Extracellular vesicle-encapsulated miR-22-3p from bone marrow mesenchymal stem cell promotes osteogenic differentiation via FTO inhibition. Stem Cell Res Ther. 2020;11(1):227.
[57] LI Z, WANG P, LI J, et al. The N(6)-methyladenosine demethylase ALKBH5 negatively regulates the osteogenic differentiation of mesenchymal stem cells through PRMT6. Cell Death Dis. 2021;12(6):578.
[58] LIU T, ZHENG X, WANG C, et al. The m(6)A “reader” YTHDF1 promotes osteogenesis of bone marrow mesenchymal stem cells through translational control of ZNF839. Cell Death Dis. 2021;12(11):1078.
[59] ZHOU S, ZHANG G, WANG K, et al. METTL3 potentiates osteogenic differentiation of bone marrow mesenchymal stem cells via IGF2BP1/m6A/RUNX2. Oral Dis. 2023. doi: 10.1111/odi.14526.
[60] KIM JM, LIN C, STAVRE Z, et al. Osteoblast-Osteoclast Communication and Bone Homeostasis. Cells. 2020;9(9):2073.
[61] ZHANG Y, GU X, LI D, et al. METTL3 Regulates Osteoblast Differentiation and Inflammatory Response via Smad Signaling and MAPK Signaling. Int J Mol Sci. 2019; 21(1):199.
[62] SUN Z, WANG H, WANG Y, et al. MiR-103-3p targets the m(6) A methyltransferase METTL14 to inhibit osteoblastic bone formation. Aging Cell. 2021;20(2):e13298.
[63] WANG C, CHEN R, ZHU X, et al. METTL14 alleviates the development of osteoporosis in ovariectomized mice by upregulating m(6)A level of SIRT1 mRNA. Bone. 2023;168: 116652.
[64] ZHANG Q, RIDDLE RC, YANG Q, et al. The RNA demethylase FTO is required for maintenance of bone mass and functions to protect osteoblasts from genotoxic damage. Proc Natl Acad Sci U S A. 2019;116(36):17980-17989.
[65] FENG L, FAN Y, ZHOU J, et al. The RNA demethylase ALKBH5 promotes osteoblast differentiation by modulating Runx2 mRNA stability. FEBS Lett. 2021;595(15):2007-2014.
[66] SHI D, LIU X, LI X, et al. Yth m(6)A RNA-Binding Protein 1 Regulates Osteogenesis of MC3T3-E1 Cells under Hypoxia via Translational Control of Thrombospondin-1. Int J Mol Sci. 2023;24(2):1741.
[67] SUN Y, LI J, XIE X, et al. Recent Advances in Osteoclast Biological Behavior. Front Cell Dev Biol. 2021;9:788680.
[68] LI D, CAI L, MENG R, et al. METTL3 Modulates Osteoclast Differentiation and Function by Controlling RNA Stability and Nuclear Export. Int J Mol Sci. 2020;21(5):1660.
[69] LI D, HE J, FANG C, et al. METTL3 Regulates Osteoclast Biological Behaviors via iNOS/NO-Mediated Mitochondrial Dysfunction in Inflammatory Conditions. Int J Mol Sci. 2023;24(2):1403.
[70] LIU J, YOU Y, SUN Z, et al. WTAP-Mediated m6A RNA Methylation Regulates the Differentiation of Bone Marrow Mesenchymal Stem Cells via the miR-29b-3p/HDAC4 Axis. Stem Cells Transl Med. 2023;12(5):307-321.
[71] ZHUANG J, NING H, WANG M, et al. Downregulated fat mass and obesity-associated protein inhibits bone resorption and osteoclastogenesis by nuclear factor-kappa B inactivation. Cell Signal. 2021;87:110137.
[72] HE M, LI D, FANG C, et al. YTHDF1 regulates endoplasmic reticulum stress, NF-kappaB, MAPK and PI3K-AKT signaling pathways in inflammatory osteoclastogenesis. Arch Biochem Biophys. 2022;732:109464.
[73] FANG C, HE M, LI D, et al. YTHDF2 mediates LPS-induced osteoclastogenesis and inflammatory response via the NF-kappaB and MAPK signaling pathways. Cell Signal. 2021;85:110060.
|