[1] KOTINI M, BARRIGA EH, LESLIE J, et al. Gap junction protein connexin-43 is a direct transcriptional regulator of N-cadherin in vivo. Nat Commun. 2018;9(1):3846.
[2] WAXSE BJ, SENGUPTA P, HESKETH GG, et al. Myosin VI facilitates connexin 43 gap junction accretion. J Cell Sci. 2017;130(5):827-840.
[3] GHAMESI TAHRIR F, GUPTA M, MYERS V, et al. Role of bcl2-associated athanogene 3 in turnover of gap junction protein,connexin 43, in neonatal cardiomyocytes. Sci Rep. 2019;9(1):7658.
[4] PERVOLARAKI E, DACHTLER J, ANDERSON RA, et al. Ventricular myocardium development and the role of connexins in the human fetal heart. Sci Rep. 2017;7(1):12272.
[5] KABA RA, COPPEN SR, DUPONT E, et al. Comparison of connexin 43, 40 and 45 expression patterns in the developing human and mouse hearts. Cell Commun Adhes. 2001;8(4-6):339-343.
[6] REAUME AG, SOUSA PAD, KULKARNI S, et al. Cardiac malformation in neonatal mice lacking connexin 43. Science. 1995;5205(267): 1831-1834.
[7] MATSUYAMA D, KAWAHARA K. Proliferation of neonatal cardiomyocytes by connexin 43 knockdown via synergistic inactivation of p38 MAPK and increased expression of FGF1. Basic Res Cardiol. 2009;104(6): 631-642.
[8] KRETZ M, ECKARDT D, KRUGER O, et al. Normal embryonic development and cardiac morphogenesis in mice with Wnt1-Cre-mediated deletion of connexin43. Genesis. 2006;44(6):269-276.
[9] GREENBERG JM, LUMBRERAS V, PELAEZ D, et al. Neural crest stem cells can differentiate to a cardiomyogenic lineage with an ability to contract in response to pulsed infrared stimulation. Tissue Eng PartC Methods. 2016;22(10):982-990.
[10] AGHAJANIAN H, CHO YK, RIZER NW, et al. Pdgfralpha functions in endothelial-derived cells to regulate neural crest cells and the development of the great arteries. Dis Models Mech. 2017;10(9): 1101-1108.
[11] HUANG GY, WESSELS A, SMITH BR, et al. Alteration in connexin 43 gap junction gene dosage impairs conotruncal heart development. Dev Biol. 1998;32-44.
[12] ANDERSEN P, TAMPAKAKIS E, JIMENEZ DV, et al. Precardiac organoids form two heart fields via Bmp/Wnt signaling. Nat Commun. 2018;9(1): 3140.
[13] CORTES C, FRANCOU A, DE BONO C, et al. Epithelial properties of the second heart field. Circ Res. 2018;122(1):142-154.
[14] ANDERSON RH, CHAUDHRY B, MOHUN TJ, et al. Normal and abnormal development of the intrapericardial arterial trunks in humans and mice. Cardiovasc Res. 2012;95(1):108-115.
[15] KEYTE A, HUTSON MR. The neural crest in cardiac congenital anomalies. Differentiation. 2012;84(1):25-40.
[16] GIBBS BC, DAMERLA RR, VLADAR EK, et al. Prickle1 mutation causes planar cell polarity and directional cell migration defects associated with cardiac outflow tract anomalies and other structural birth defects. Biol Open. 2016;5(3):323-335.
[17] OKAMOTO N, AKIMOTO N, HIDAKA N, et al. Formal genesis of the outflow tracts of the heart revisited: previous works in the light of recent observations. Congenit Anom. 2010;50(3):141-158.
[18] AGHAJANIAN H, CHO YK, RIZER NW, et al. Pdgfralpha functions in endothelial-derived cells to regulate neural crest cells and the development of the great arteries. Dis Models Mech. 2017;10(9): 1101-1108.
[19] ZHANG J, JULIA YF, CHANGYQ, et al. The FGF-BMP signaling axis regulates outflow tract valve primordium formation by promoting cushion neural crest cell differentiation. Circ Res. 2010;107(10): 1209-1219.
[20] 师亮,李慧超,景雅,等.鼠胚呼吸内胚层相关第二生心区发育中的上皮-间充质转化[J].解剖学报,2018,49(4):480-485.
[21] RAMIREZ A, ASTROF S. Vualization and analysis of pharyngeal arch arteries using whole-mount immunohistochemistry and 3D reconstruction. J Vis Exp. 2020. Doi:10.3791/60797.
[22] SONG YC, DOHN TE, RYDEEN AB, et al. HDAC1-mediated repression of the retinoic acid-responsive gene ripply3 promotes second heart field development. PLoS Genet. 2019;15(5):e1008165.
[23] WITZEL HR, JUNGBLUT B, CHOE CP, et al. The LIM protein ajuba restricts the second heart field progenitor pool by regulating Isl1 activity. Dev Cell. 2012;23(1):58-70.
[24] PIVEN OO, WINATA CL. The canonical way to make a heart: β-catenin and plakoglobin in heart development and remodeling. Exp Biol and Med. 2017;242(18):1735-1745.
[25] ANTHONY J, HORTON AJ, BROOKER J, et al. Nkx2.5 second heart field target gene Ccdc117 regulates DNA metabolism and proliferation. Sci Rep. 2019;9(1):1738.
[26] Colombo S, de Sena-Tomas C, George V, et al. Nkx2.5 genes establish second heart field cardiomyocyte progenitors at the arterial pole and pattern the venous pole through Isl1 repression. Development. 2019. Doi: 10.1242/dev.161497.
[27] ZHOU L, LIU JL, XIANG ML, et al. Gata4 potentiates second heart field proliferation and Hedgehog signaling for cardiac septation. Proc Natl Acad Sci U S A. 2017;114(8):E1422-E1431.
[28] SAWADA H, RATERI DL, MOORLEGHEN JJ, et al. Smooth muscle cells derived from second heart field and cardiac neural crest reside in spatially distinct domains in the media of the ascending aorta-brief report. Arter Thromb Vasc Biol. 2017;37:1722-1726.
[29] MOLIN DG, POELMANN RE, DERUITER MC, et al. Transforming growth factor beta-SMAD2 signaling regulates aortic arch innervation and development. Circ Res. 2004;95(11):1109-1117.
[30] KAREN LW, CECILIA WL, MARGARET L, et al. Connexin 43 expression reflflects neural crest patterns during cardiovascular development. Dev Biol. 1999;208:307-323.
[31] XU X, FRANCIS R, WEI CJ, et al. Connexin 43-mediated modulation of polarized cell movement and the directional migration of cardiac neural crest cells. Development. 2006;133(18):321-324. |
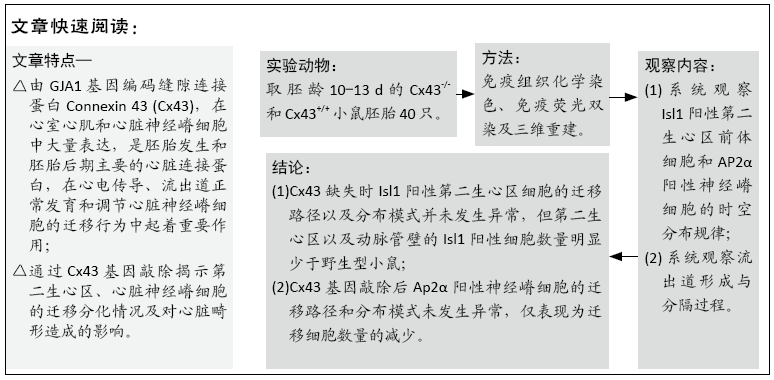 文题释义:
文题释义: