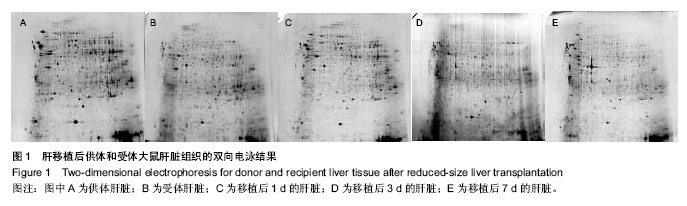
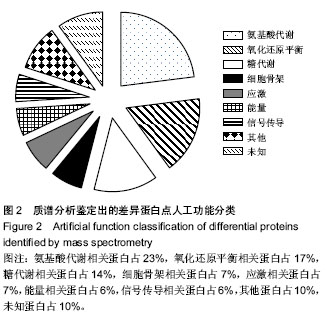
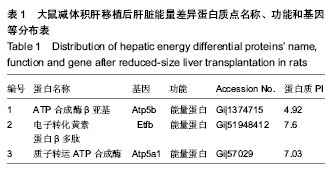
| [1] Brewis IA, Brennan P. Proteomics technologies for the global identification and quantification of proteins. Adv Protein Chem Struct Biol. 2010;80:1-44.
[2] Woods AG, Ngounou Wetie AG, Sokolowska I, et al. Mass spectrometry as a tool for studying autism spectrum disorder. J Mol Psychiatry. 2013;1(1):6.
[3] Woods AG, Wormwood KL, Wetie AG, et al. Autism spectrum disorder: An omics perspective. Proteomics Clin Appl. 2014. in press.
[4] Mayer G, Jones AR, Binz PA, et al. Controlled vocabularies and ontologies in proteomics: overview, principles and practice. Biochim Biophys Acta. 2014;1844(1 Pt A):98-107.
[5] Dowling P, Hayes C, Ting KR, et al. Identification of proteins found to be significantly altered when comparing the serum proteome from Multiple Myeloma patients with varying degrees of bone disease. BMC Genomics. 2014; 15:904.
[6] Pooladi M, Abad SK, Hashemi M. Proteomics analysis of human brain glial cell proteome by 2D gel. Indian J Cancer. 2014;51(2):159-162.
[7] Hashemi M, Pooladi M, Razi Abad SK. The investigation of changes in proteins expression (Apolipoprotein A1 and albumin) in malignant astrocytoma brain tumor. J Cancer Res Ther. 2014;10(1):107-111.
[8] Kienzl-Wagner K, Pratschke J, Brandacher G. Proteomics--a blessing or a curse? Application of proteomics technology to transplant medicine. Transplantation. 2011;92(5):499-509.
[9] Kienzl-Wagner K, Pratschke J, Brandacher G. Biomarker discovery in transplantation--proteomic adventure or mission impossible? Clin Biochem. 2013;46(6):497-505.
[10] Cohen Freue GV, Meredith A, Smith D, et al. Computational biomarker pipeline from discovery to clinical implementation: plasma proteomic biomarkers for cardiac transplantation. PLoS Comput Biol. 2013;9(4):e1002963.
[11] Metzger J, Chatzikyrkou C, Broecker V, et al. Diagnosis of subclinical and clinical acute T-cell-mediated rejection in renal transplant patients by urinary proteome analysis. Proteomics Clin Appl. 2011;5(5-6):322-333.
[12] Kim SC, Page EK, Knechtle SJ. Urine proteomics in kidney transplantation. Transplant Rev (Orlando). 2014;28(1): 15-20.
[13] Halawa A. The early diagnosis of acute renal graft dysfunction: a challenge we face. The role of novel biomarkers. Ann Transplant. 2011;16(1):90-98.
[14] Sigdel TK, Salomonis N, Nicora CD, et al. The identification of novel potential injury mechanisms and candidate biomarkers in renal allograft rejection by quantitative proteomics. Mol Cell Proteomics. 2014;13(2):621-631.
[15] Pisitkun T, Gandolfo MT, Das S, et al. Application of systems biology principles to protein biomarker discovery: urinary exosomal proteome in renal transplantation. Proteomics Clin Appl. 2012;6(5-6):268-278.
[16] Hossain MA, De Souza AI, Bagul A, et al. HSP70, Peroxiredoxin-3 and -6 are upregulated during renal warm ischaemia in a donation after circulatory death model. J Proteomics. 2014;108:133-145.
[17] Kornasiewicz O, Bojarczuk K, Bugajski M, et al. Application of a proteomic approach to identify proteins associated with primary graft non-function after liver transplantation. Int J Mol Med. 2012;30(4):755-764.
[18] Baker ES, Burnum-Johnson KE, Jacobs JM, et al. Advancing the high throughput identification of liver fibrosis protein signatures using multiplexed ion mobility spectrometry. Mol Cell Proteomics. 2014;13(4):1119-1127.
[19] 刘静,李江,张升宁,等.改良法构建大鼠减体积肝移植模型[J].中国组织工程研究与临床康复杂志,2010,14(18):3252-3257.
[20] Jing L, Li L, Jiang-Hua R, et al. A causal analysis of intra- abdominal hemorrhage after reduced-size liver transplantation in rat. Cell Biochem Biophys. 2011;61(3):685-690.
[21] Okayama A, Miyagi Y, Oshita F, et al. Proteomic analysis of proteins related to prognosis of lung adenocarcinoma. J Proteome Res. 2014;13(11):4686-4694.
[22] Kim MS, Pinto SM, Getnet D, et al. A draft map of the human proteome. Nature. 2014;509(7502):575-581.
[23] Wilhelm M, Schlegl J, Hahne H, et al. Mass-spectrometry- based draft of the human proteome. Nature. 2014;509(7502): 582-587.
[24] Rigbolt KT, Prokhorova TA, Akimov V, et al. System-wide temporal characterization of the proteome and phosphoproteome of human embryonic stem cell differentiation. Sci Signal. 2011;4(164):rs3.
[25] Mardinoglu A, Kampf C, Asplund A, et al. Defining the human adipose tissue proteome to reveal metabolic alterations in obesity. J Proteome Res. 2014;13(11):5106-5119.
[26] Martins-de-Souza D. Proteomics, metabolomics, and protein interactomics in the characterization of the molecular features of major depressive disorder. Dialogues Clin Neurosci. 2014; 16(1):63-73.
[27] Sleddering MA, Markvoort AJ, Dharuri HK, et al. Proteomic Analysis in Type 2 Diabetes Patients before and after a Very Low Calorie Diet Reveals Potential Disease State and Intervention Specific Biomarkers. PLoS One. 2014;9(11): e112835.
[28] Jov?evska I, Zupanec N, Ko?evar N, et al. TRIM28 and β-Actin Identified via Nanobody-Based Reverse Proteomics Approach as Possible Human Glioblastoma Biomarkers. PLoS One. 2014;9(11):e113688.
[29] Webb-Robertson BJ, Matzke MM, Datta S, et al. Bayesian proteoform modeling improves protein quantification of global proteomic measurements. Mol Cell Proteomics. 2014;13(12): 3639-3646.
[30] Dallas DC, Guerrero A, Parker EA, et al. Current peptidomics: Applications, purification, identification, quantification and functional analysis. Proteomics. 2014. in press.
[31] Dong Y, Cai J, Chen S. Proteomics of drug-resistant cancer biomarkers. Bioanalysis. 2014;6(19):2519-2521.
[32] Tang B, Li Y, Zhao L, et al. Stable isotope dimethyl labeling combined with LTQ mass spectrometric detection, a quantitative proteomics technology used in liver cancer research. Biomed Rep. 2013;1(4):549-554.
[33] Sharma S, Ray S, Mukherjee S, et al. Multipronged quantitative proteomic analyses indicate modulation of various signal transduction pathways in human meningiomas. Proteomics. 2014. in press.
[34] Hirokawa S, Shimanuki T, Kitajima H, et al. Knockdown of electron transfer flavoprotein β subunit reduced TGF-β-induced α-SMA mRNA expression but not COL1A1 in fibroblast-populated three-dimensional collagen gel cultures. J Dermatol Sci. 2012;68(3):179-186.
[35] Malecki J, Ho AY, Moen A, et al. Human METTL20 is a Mitochondrial Lysine Methyltransferase that Targets the Beta Subunit of Electron Transfer Flavoprotein (ETFβ) and Modulates Its Activity. J Biol Chem. 2014. in press.
[36] Rhein VF, Carroll J, He J, et al. Human METTL20 methylates lysine residues adjacent to the recognition loop of the electron transfer flavoprotein in mitochondria. J Biol Chem. 2014;289 (35): 24640-24651.
[37] Kasumov EA, Kasumov RE, Kasumova IV. A mechano- chemiosmotic model for the coupling of electron and proton transfer to ATP synthesis in energy-transforming membranes: a personal perspective. Photosynth Res. 2014. in press.
[38] Schertl P, Braun HP. Respiratory electron transfer pathways in plant mitochondria. Front Plant Sci. 2014;5:163.
[39] Morelli AM, Ravera S, Calzia D, et al. Hypothesis of lipid-phase-continuity proton transfer for aerobic ATP synthesis. J Cereb Blood Flow Metab. 2013;33(12):1838-1842.
[40] Koumandou VL, Kossida S. Evolution of the F0F1 ATP synthase complex in light of the patchy distribution of different bioenergetic pathways across prokaryotes. PLoS Comput Biol. 2014;10(9):e1003821.
[41] Falk G, Walker JE. DNA sequence of a gene cluster coding for subunits of the F0 membrane sector of ATP synthase in Rhodospirillum rubrum. Support for modular evolution of the F1 and F0 sectors. Biochem J. 1988;254(1):109-122.
[42] von Ballmoos C, Wiedenmann A, Dimroth P. Essentials for ATP synthesis by F1F0 ATP synthases. Annu Rev Biochem. 2009;78:649-672.
[43] Dimroth P, Kaim G, Matthey U. Crucial role of the membrane potential for ATP synthesis by F(1)F(o) ATP synthases. J Exp Biol. 2000;203(Pt 1):51-59.
[44] Arakaki N, Nagao T, Niki R, et al. Possible role of cell surface H+ -ATP synthase in the extracellular ATP synthesis and proliferation of human umbilical vein endothelial cells. Mol Cancer Res. 2003;1(13):931-99.
[45] Fillingame RH. Coupling H+ transport and ATP synthesis in F1F0-ATP synthases: glimpses of interacting parts in a dynamic molecular machine. J Exp Biol. 1997;200(Pt 2):217-224.
[46] Deshpande M, Notari L, Subramanian P, et al. Inhibition of tumor cell surface ATP synthesis by pigment epithelium -derived factor: implications for antitumor activity. Int J Oncol. 2012;41(1):219-227.
[47] Wang J, Li R, Zhang G. Research on relevance between mitochondrial ATP synthase and malignant tumor. Sheng Wu Yi Xue Gong Cheng Xue Za Zhi. 2014;31(3):714-717.
[48] Sánchez-Aragó M, Formentini L, Cuezva JM. Mitochondria-mediated energy adaption in cancer: the H(+)-ATP synthase-geared switch of metabolism in human tumors. Antioxid Redox Signal. 2013;19(3):285-298.
[49] Bockenhauer SD, Duncan TM, Moerner WE, et al. The regulatory switch of F1-ATPase studied by single-molecule FRET in the ABEL Trap. Proc Soc Photo Opt Instrum Eng. 2014;8950:89500H.
[50] Duncan TM, Düser MG, Heitkamp T, et al. Regulatory conformational changes of the ε subunit in single FRET-labeled FoF1-ATP synthase. Proc Soc Photo Opt Instrum Eng. 2014;8948:89481J.
[51] Börsch M, Duncan TM. Spotlighting motors and controls of single FoF1-ATP synthase. Biochem Soc Trans. 2013;41(5): 1219-1226.
[52] Fu Y, Zhu Y. Ectopic ATP synthase in endothelial cells: a novel cardiovascular therapeutic target. Curr Pharm Des. 2010; 16(37): 4074-4079.
[53] Krysko DV, Vandenabeele P. From regulation of dying cell engulfment to development of anti-cancer therapy. Cell Death Differ. 2008;15(1):29-38.
[54] Subramanian K, Naik VD, Sathishkumar K, et al. Interactive effects of in vitro binge-like alcohol and ATP on umbilical endothelial nitric oxide synthase post-translational modifications and redox modulation. Reprod Toxicol. 2014;43:94-101.
[55] Barker RN, Erwig LP, Hill KS, et al. Antigen presentation by macrophages is enhanced by the uptake of necrotic, but not apoptotic, cells. Clin Exp Immunol. 2002;127(2):220-225.
[56] Oh YS, Lee YJ, Park K, et al. Treatment with glucokinase activator, YH-GKA, increases cell proliferation and decreases glucotoxic apoptosis in INS-1 cells. Eur J Pharm Sci. 2014;51: 137-145.
[57] De Marchi U, Thevenet J, Hermant A, et al. Calcium co-regulates oxidative metabolism and ATP synthase- dependent respiration in pancreatic beta cells. J Biol Chem. 2014;289(13):9182-9194. |