[1] 任云晓,肖茹丹,娄晓敏,等.基因编辑技术及其在基因治疗中的应用[J].遗传,2019,41(1):18-28.
[2] MALI P, YANG L, ESVELT KM, et al. RNA-guided human genome engineering via Cas9 prashant. Science. 2013;339(6121):823-826.
[3] 陈怡李,姚书忠.CRISPR/Cas9基因编辑技术的应用研究进展[J].国际生殖健康/计划生育杂志,2017,36(6):482-487.
[4] 丁晓,史晨辉,孟德峰,等.骨关节炎关键基因与治疗药物的生物信息学筛选[J].中国实验方剂学杂志,2019,25(9):189-196.
[5] 杨春艳,王磊,穆登彩,等.基因编辑技术在疾病治疗中的研究进展[J].中国生物工程杂志,2019,39(11):87-95.
[6] CAI M, YANG Y. Targeted genome editing tools for disease modeling and gene therapy. Curr Gene Ther. 2014;14(1):2-9.
[7] CONG L, RAN FA, COX D, et al. Multiplex genome engineering using CRISPR/Cas systems. Science. 2013;339(6121):819-823.
[8] 牛煦然,尹树明,陈曦,等.基因编辑技术及其在疾病治疗中的研究进展[J].遗传,2019,41(7):582-598.
[9] ANGUELA XM, HIGH KA. Entering the modern era of gene therapy. Annu Rev Med. 2019;70:273-288.
[10] YOSHIDA K, TREEN N. TALEN-Based Knockout System. Adv Exp Med Boil. 2018;1029:131-139.
[11] 何景进,王慧,付雪梅.基因编辑技术研究进展[J].发育医学电子杂志, 2019,7(3):235-240.
[12] 曲道峰,沈杨,张聪聪,等.沙门氏菌CRISPR位点的结构特征比较[J].微生物学报,2018,58(2):209-2018.
[13] CONG L, RAN FA, COX D, et al. Multiplex genome engineering using CRISPR/Cas systems. Science. 2013;339:819-823.
[14] 吴璐,王磊,任远,等.基因组编辑技术研究进展[J].生物技术通报, 2014(11):84-90.
[15] 玉婷,董金玲,冯力元,等.内源性过表达Nestin对NIH3T3细胞增殖、迁移及MAPK信号传导通路的影响[J].第三军医大学学报,2020, 42(2):125-132.
[16] 王昕.基因编辑技术创新构建大鼠药物代谢模型及其应用[J].中国药理学与毒理学杂志,2019,33(10):814.
[17] NIU YY, SHEN B, CUI YQ, et al. Generation of gene-modified cynomolgus monkey via Cas9/RNA-mediated gene targeting in onecell embryos. Cell. 2014;156:836-843.
[18] 孙明尧,白玮,梁宏玉,等.CRISPR-Cas9技术在医药领域中的应用[J].沈阳药科大学学报,2019, 36(12):1145-1152.
[19] RICHTER H, RANDAU L, PLAGENS A. Exploiting CRISPR-Cas:interference mechanisms and applications. Int J Mol Sci. 2013;14(7):14518-14531.
[20] 陶洋,李金城,廖奇.CRISPR相关的生物信息学基础[J].中国生物化学与分子生物学报, 2016,32(10):1067-1076.
[21] MA DJ, LI Y, XIAO W, et al. Achyranthes bidentata extract protects chondrocytes functions through suppressing glycolysis and apoptosis via MAPK/AKT signaling axis. Am J Transl Res. 2020;12(1):142-152.
[22] 王继成,易智.miRNA与骨关节炎病理发展过程的相关性[J].中国组织工程研究,2019,23(24):3875-3881.
[23] 顾晓东,车先达,李鹏翠,等.骨关节炎基因治疗的研究进展[J].中华骨与关节外科杂志,2019,12(5):396-398.
[24] 詹子睿,邵增务.骨关节炎基因治疗进展[J].中国中医骨伤科杂志, 2004,12(3):57-59.
[25] 邵进,张岩,王治,等.低氧诱导因子-1α参与骨发育及骨代谢调控的研究进展[J].中国骨质疏松杂志,2015,21(3):349-355.
[26] 田大川,李海乐,肖大伟,等.联合应用软骨再生支架与突变型HIF-1α修饰BMSCs分泌的外泌体对晚期软骨缺损修复的促进作用[J].吉林大学学报,2018,44(2):216-222.
[27] ALLERSTORFER D, LONGATO S, SCHWARZER C, et al. VEGF and its role in the early development of the long bone epiphysis. J Anat. 2010;216(5):611-624.
[28] 马笃军,彭力平,余阗,等.牛膝总皂苷对骨关节炎模型兔软骨修复及低氧诱导因子1信号通路的影响[J].中国组织工程研究, 2019,23(27): 4332-4337.
[29] BOBACZ K, GRUBER R, SOLEIMAN A, et al. Expression of bone morphogenetic protein 6 in healthy and osteoarthritic human articular chondrocytes and stimulation of matrix synthesis in vitro. Arthritis Rheum. 2003;48(9):2501-2508.
[30] 于斐,曾晖,雷鸣,等.SIRT1基因敲除对骨关节炎小鼠VEGF/AKT信号通路的影响[J].中国骨质疏松杂志,2016,22(1):30-40.
[31] BEENBAUM F. Osteoarthritis as an inflammatory disease (osteoarthritis is not osteoarthrosis). Osteoarthritis Cartilage. 2013;21(3):16-21.
[32] 石继祥,纪斌,虞陆超,等.补肾活血方对骨关节炎小鼠局部淋巴管结构与分布的影响[J].中华中医药学刊,2018,12(12):3000-3004.
[33] 张伟.张蜀茂.陈跃.淋巴系统显像的现状及研究进展[J].广东医学, 2018,39(12):3441-3447.
[34] 韩海慧,王晓赟,梁倩倩,等.淋巴管系统与骨性关节炎的相关性研究进展[J].中华中医药杂志,2019,34(10):4727-4729, 4730.
[35] 张武强,石继祥.骨性关节炎中淋巴管的研究进展[J].世界最新医学信息文摘,2019,19(66):106-107.
[36] GUO R, ZHOU Q, PROULX ST, et al. Inhibition of lymphangiogenesis and lymphatic drainage via vascular endothelial growth factor receptor 3 blockade increases the severity of inflammation in a mouse model of chronic inflammatory arthritis. Arthritis Rheum. 2009;60(9):2666-2676.
[37] LIANG QQ, SHI Q, WOODR W, et al. Peri-articular lymphatic system and“Bi” theory of Chinese medicine in the pathogenesis and treatment of arthritis. Chin J Integr Med. 2015;21(9):648-655.
[38] ZHOU Q, WOOD R, SCHWA R, et al. Near-infrared lymphatic imaging demonstrates the dynamics of lymph flow and lymphangiogenesis.during the acute versus chronic phases of arthritis in mice. Arthritis Rheum. 2010;62(7):1881-1889.
[39] ZISCHEWSKI J, FISCHER R, BORTESI L. Detection of on-target and off-target mutations generated by CRISPR/Cas9 and other sequence-specific nucleases. Biotechnol Adv. 2016;35(1):95-104.
[40] 黄丕铂.基因编辑婴儿事件及其对科技创新管理的影响[J].科技中国, 2019,2:35-38.
[41] HSU PD, LANDER ES, ZHANG F. Development and applications of CRISPR-Cas9 for genome engineering. Cell. 2014;157(6):1262-1278.
[42] 杨伟,李施施,张瑄,等.基于诱导多能干细胞的基因编辑和细胞治疗[J].中国细胞生物学学报,2015,37(1):90-99.
[43] ZHANG X, LI S, YANG W, et al. Patient-specificinduced pluripotent stem cell models in mitochondrial diseases. Curr Stem Cell Res Ther. 2014;9(2):134-140.
[44] INOUE H, YAMANAKA S. The use of induced pluripotent stem cells in drug development. Clin Pharmacol Ther. 2011;89(5):655-661.
[45] 郑武,谷峰.CRISPR/Cas9的应用及脱靶效应研究进展[J].遗传,2015, 37(10):1003-1010.
[46] LOWRY WE, QUAN WL. Roadblocks en route to the clinical application of induced pluripotent stem cells. J Cell Sci. 2010;123(Pt 5):643-651.
[47] 周琪.干细胞如何创造生命的奇迹[J].科学中国人,2019(12):38-42.
[48] 张庆金,关若洪,张安文,等.miR-410可促进骨髓间充质干细胞定向分化为软骨细胞[J].中国组织工程研究,2019,23(1):13-17.
[49] 马笃军,彭力平,王立新,等.牛膝醇提物诱导兔骨髓间充质干细胞软骨定向分化的实验研究[J].中国中医骨伤科杂志2017,25(2):6-11.
[50] 陈锦富,耿倚云,王大平.定向诱导间充质干细胞成软骨分化相关活性因子研究进展[J]. 组织工程与重建外科杂志, 2018,14(4):227-230.
[51] KOZHEMYAKINA E, LASSAR AB, ZELZER E. A pathway to bone: signaling molecules and transcription factors involved in chondrocyte development and maturation. Development. 2015;142(5):817-831.
[52] CORREA D, SOMOZA RA, LIN P, et al. Sequential exposure to fibroblast growth factors (FGF)2,9 and 18 enhances hMSC chondrogenic differentiation. Osteoarthritis Cartilage. 2015;23(3):443-453.
[53] 李大伟,孙一,姜翠萍,等.重组人骨形态发生蛋白-2质粒转染诱导人脐血间充质干细胞软骨分化研究[J].中华老年骨科与康复电子杂志, 2019,5(2):75-81.
[54] YIN C, ZHANG T, QU X, et al. In vivo excision of HIV-1 provirus by saCas9 and multiplex single-guide RNAs in animal models. Mol Ther. 2017;25(5):1168-1186.
[55] KC M, STEER CJ. A new era of gene editing for the treatment of human diseases. Swiss Med Wkly. 2019;149:w20021.
[56] 陈曦,陈亮,李大力.基因治疗在临床应用中的研究进展[J].生物工程学报,2019,35(12):2295-2307.
[57] TSAI SQ, JOUNG JK. Defining and improving the genome-wide specificities of CRISPR-Cas9 nucleases. Nat Rev Genet. 2016;17(5):300-312. |
 最终要把基因编辑产品应用于临床,目前还是存在很多尚需解决的问题,主要有:①选择合适的目的基因;② 如何调节治疗基因的长期表达;③ 如何使用安全的递送载体转移基因;④如何降低脱靶效应,脱靶效应严重制约了基因编辑技术的广泛应用[56];⑤伦理学问题。随着新工具的开发和利用,人类的思想也逐步开阔,那些技术上的问题都将会得到解决,例如利用基因测序选择疾病特定目的基因序列,对干细胞进行基因修饰维持特定基因持续表达,利用生物信息学的方法筛选非同源或少相似性靶基因位点,以减少脱靶效率[57]。技术的进步会对伦理的发展提供更加高的要求,促使科学工作者更加遵守伦理纲常,合理使用基因编辑工具造福于人类。
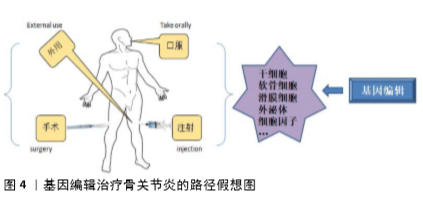
骨关节炎的发病因素较为复杂,循证医学研究已证实多学科技术联合治疗关节炎可明显提高疗效,值得进一步的尝试研究。随着基因组学技术的迅速发展,基因编辑技术已经成为基因研究必不可少的手段之一,因此,基因组编辑技术的不断更新发展是必然的趋势。与传统的基因治疗不同,基因编辑技术可实现精确的靶基因插入、敲除及编辑。基因编辑技术可在离体细胞上进行,也可通过载体递送的方式在体内实现原位基因组的编辑。因此,基因编辑技术为骨关节炎基因治疗方案提供了更多的新途径,将来必定在预防及治疗骨关节炎方面起到举足轻重的作用。
最终要把基因编辑产品应用于临床,目前还是存在很多尚需解决的问题,主要有:①选择合适的目的基因;② 如何调节治疗基因的长期表达;③ 如何使用安全的递送载体转移基因;④如何降低脱靶效应,脱靶效应严重制约了基因编辑技术的广泛应用[56];⑤伦理学问题。随着新工具的开发和利用,人类的思想也逐步开阔,那些技术上的问题都将会得到解决,例如利用基因测序选择疾病特定目的基因序列,对干细胞进行基因修饰维持特定基因持续表达,利用生物信息学的方法筛选非同源或少相似性靶基因位点,以减少脱靶效率[57]。技术的进步会对伦理的发展提供更加高的要求,促使科学工作者更加遵守伦理纲常,合理使用基因编辑工具造福于人类。
骨关节炎的发病因素较为复杂,循证医学研究已证实多学科技术联合治疗关节炎可明显提高疗效,值得进一步的尝试研究。随着基因组学技术的迅速发展,基因编辑技术已经成为基因研究必不可少的手段之一,因此,基因组编辑技术的不断更新发展是必然的趋势。与传统的基因治疗不同,基因编辑技术可实现精确的靶基因插入、敲除及编辑。基因编辑技术可在离体细胞上进行,也可通过载体递送的方式在体内实现原位基因组的编辑。因此,基因编辑技术为骨关节炎基因治疗方案提供了更多的新途径,将来必定在预防及治疗骨关节炎方面起到举足轻重的作用。