Chinese Journal of Tissue Engineering Research ›› 2022, Vol. 26 ›› Issue (13): 2106-2113.doi: 10.12307/2022.341
Previous Articles Next Articles
Application of single-cell RNA sequencing in diagnosis and treatment of corneal and retinal diseases
Huang Chang1, Ying Peixi1, Yi Guoguo2, Fu Sheng3, Wang Yan4, Guo Xi5, Zhang Yuxi1, Fu Min6
- 1Second Clinical Medical College, Southern Medical University; 2Department of Ophthalmology, Sixth Affiliated Hospital, Sun Yat-sen University; 3Nanhua University; 4Department of Ophthalmology, Shenzhen Hospital, Southern Medical University; 5School of Rehabilitation Medicine, Southern Medical University; 6Department of Ophthalmology, Zhujiang Hospital, Southern Medical University
-
Received:2021-05-06Revised:2021-05-10Accepted:2021-06-15Online:2022-05-08Published:2021-12-20 -
Contact:Fu Min, Master, Associate chief physician, Department of Ophthalmology, Zhujiang Hospital, Southern Medical University, Guangzhou 510220, Guangdong Province, China -
About author:Huang Chang, Second Clinical Medical College, Southern Medical University, Guangzhou 510580, Guangdong Province, China Ying Peixi, Second Clinical Medical College, Southern Medical University, Guangzhou 510580, Guangdong Province, China Yi Guoguo, MD, Department of Ophthalmology, Sixth Affiliated Hospital, Sun Yat-sen University, Guangzhou 510630, Guangdong Province, China Huang Chang, Ying Peixi, and Yi Guoguo contributed equally to this article.
CLC Number:
Cite this article
Huang Chang, Ying Peixi, Yi Guoguo, Fu Sheng, Wang Yan, Guo Xi, Zhang Yuxi, Fu Min. Application of single-cell RNA sequencing in diagnosis and treatment of corneal and retinal diseases[J]. Chinese Journal of Tissue Engineering Research, 2022, 26(13): 2106-2113.
share this article
Add to citation manager EndNote|Reference Manager|ProCite|BibTeX|RefWorks
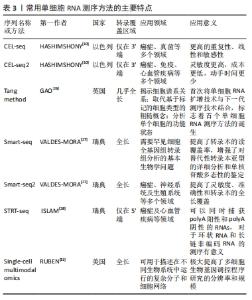
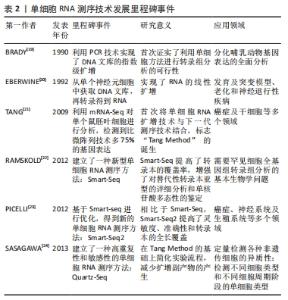
技术噪声与生物变异对于单细胞测序的数据分析提出了挑战,为了更好地克服这些问题,越来越多分析单细胞测序的工具被提出,但各自存在一定的优缺点。有研究在2013年首个开发了单细胞RNA测序方法[25]。在一篇具有里程碑意义的文章中,这种方法被称为“Tang Method”[26]。与标准微阵列实验相比,Tang Method可以更有效地检测到数千个基因和数百个新的拼接连接,使研究人员充分认识到相比于分析大量样本,分析单个细胞更为重要。几年后,有研究选择了另一种不同的方法——Smart-seq和Smart-seq2[27],试图借此显著增加转录组的覆盖率。几乎与此同时,Hashimshony等[10]利用CEL-seq方法(细胞线性扩增和测序表达),通过使用不同方法来扩增RNA,来生成足够的测序材料。在单细胞标记的反转录测序中,有研究者在单细胞领域开发了一套STRT-seq方法[28],可以使用锚定的寡核苷酸引物和鼠白血病反转录酶基酶进行反转录。在这之后,InDrop能够以极快的速度对大量来自异质群体的细胞进行测序,使鉴定出少见的细胞类型成为可能。尽管Drop seq与InDrop这2种方法是独立开发而成,但是它们却有一定的相似之处。CytoSeq,发布于2015年初,是最早的测序手段,可以实现常规的数字基因表达谱分析。应用这种方法,在不使用传统液体处理机的情况下,除了个别基因,几乎所有基因都可以从数千计的单细胞中获取。此外,MARS-seq,SCRB-seq和Seq-well也可应用于单细胞转录研究中[29]。表3总结了传统的和新一代的单细胞RNA测序技术的主要特点。"

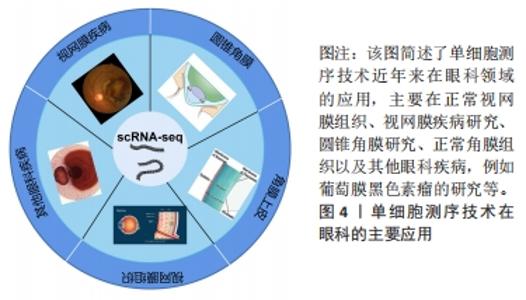
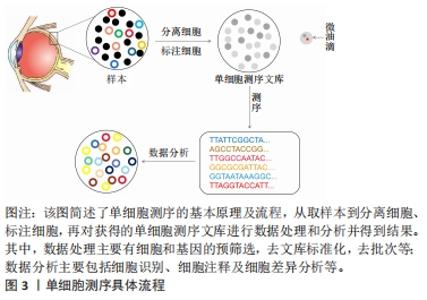
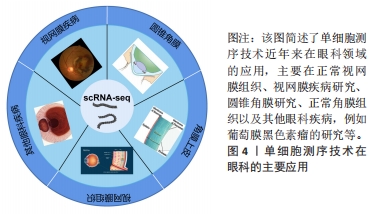
空间转录技术是通过用一个独特的位置的条形码定位在反转录引物的组织切片中,可显示高质量的RNA测序数据和组织切片的二维位置信息。空间转录技术提供定量的基因表达数据,在组织切片mRNA的分布中可视化,并使用了新型的生物信息学分析,这对于研究和诊断具有重要意义。单细胞多模态组学技术在2019年被评为年度最佳技术,成为未来研究的热点[30-31],单细胞测序技术与空间转录技术相结合的单细胞多组学技术已用于疾病的发生发展研究。单细胞测序技术是从单个细胞水平分析细胞转录组的表达谱,用于鉴定细胞特异性标记物,发现罕见细胞类型、细胞亚型,揭示细胞之间的差异表达,成为研究复杂生物体系细胞异质性的有效方法,空间转录技术对于理解组织中细胞的身份和功能至关重要,两者结合对疾病解析更具有说服力。单细胞多组学技术在研究和阐释疾病的发生发展中已有较多应用[29]。细胞和组织之间的关系是理解正常生理情况和疾病病理之间的关键。单细胞多组学技术可以解析整个空间mRNA的表达,同时可将基因表达谱映射回原来的位置。单细胞多组学技术适用于神经科学、癌症和发育生物学等多个领域的研究,为进一步了解组织和基因的表达提供了新的视角。 细胞是生命最基本的单位。对细胞功能的分析不仅本身极具趣味性,而且还能揭示细胞与周围环境之间的复杂关系。在单细胞RNA测序领域,不断发展的新技术有望产生前所未有的海量信息,并对基础研究和临床科学产生重大影响。单细胞测序技术在眼科领域的应用,见图4。近年来主要为角膜、视网膜的相关研究,这对于眼科临床治疗有重要的意义。"
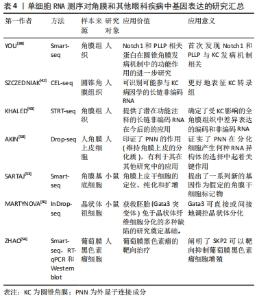
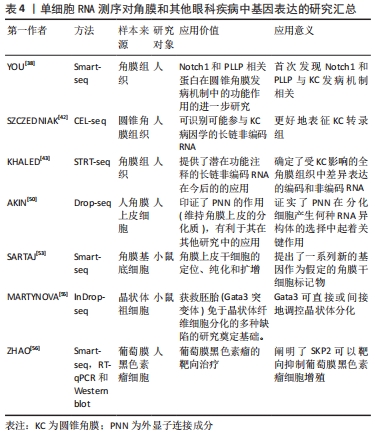
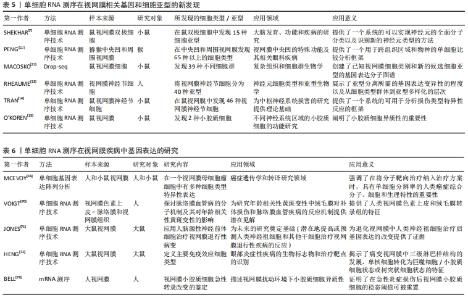
2.2 单细胞RNA测序技术在角膜中的应用 单细胞RNA测序技术是一项相对新兴的技术,正引领测序领域飞速发展。该技术提供了一种新方法来表征具有多重异质性的细胞系统且可用于分析单细胞转录组[32]。 确切地说,所有与角膜相关的单细胞测序研究均局限于圆锥角膜、角膜上皮细胞基因表达及干细胞研究、角膜内皮细胞功能障碍这几项研究中[33]。 2.3 单细胞RNA测序技术在圆锥角膜中的应用 圆锥角膜是一种与角膜胶原组织结构变化有关的退行性角膜疾病,可导致严重视力损害,进而使患者生活质量下降[34]。该疾病可使角膜间质逐渐变薄,导致角膜变为锥形、光学变形、甚至丧失视觉功能[35]。圆锥角膜的发病机制与遗传和环境因素有关[36],该疾病多发病于青春期,呈双侧性,但双眼的发病时间与疾病严重程度可不一致。典型的组织病理学变化为鲍曼层断裂和角膜基质变薄,角膜中央或旁中央呈锥型扩张,圆锥顶端的角膜基质变薄最为明显。同时,角膜层的结构变化也会导致光学像差,进而导致视觉功能丧失[37]。 YOU等[38]通过Smart-seq测序方法研究圆锥角膜患者的组织,发现在患者组织中的Hedgehog,Wnt和Notch1信号通路相关基因下调,血浆磷脂和Notch1基因蛋白表达显著降低(P < 0.05)。血浆磷脂是在人角膜上皮细胞中发现的一种新型蛋白,在其他组织上皮细胞的分化中起着重要作用[39]。BYKHOVSKAYA团队[40]使用CEL-seq的测序方式,研究人类圆锥角膜组织并提出了相关候选基因,包括VSX1,HGF和mir-184.5-7。高密度寡核苷酸芯片显示,与正常角膜上皮细胞相比,圆锥角膜上皮细胞中有56个基因异常表达[41],其中ACTN1,SERPINB1,LOX,SLPI,DSG3,FHL2,ANXA1,LGALS1,RPL23,S100A2和d-GPNMB在正常角膜上皮细胞中也有表 达[42]。科学研究表明圆锥角膜可能影响角膜基质中HSP90、核心蛋白聚糖、纤连蛋白、铁蛋白重链和角蛋白的表达[35]。 在前人的研究基础上,KHALED等[43]使用STRT-seq测序方式,研究并强调了在圆锥角膜患者组织中,CTGF,SFRP1,AQP5,lnc-WNT4-2:1和lncALDH3A2-2:1基因的表达增高。 根据以往的研究,圆锥角膜处于非炎症状态,其治疗方案应首选角膜移植和角膜交联。然而,根据最近的研究结果,在圆锥角膜中某些炎症因子(如白细胞介素6、肿瘤坏死因子α和白细胞介素1受体)升高。圆锥角膜患者上皮细胞白细胞介素6、PPARG和HLA表达下调[44],推测它们可能参与了圆锥角膜的发病过程[45]。其中,Ⅰ型单纯疱疹病毒感染对白细胞介素6表达的变化有重要影响[46],白细胞介素6在神经变性、眼内炎症和角膜血管生成中也起重要作用。并且根据PPARG激动剂罗格列酮在视网膜母细胞瘤细胞中的作用表明,罗格列酮可抑制细胞生长、转移和侵袭,促进细胞凋亡[47],具有抗肿瘤作用[48]。这些发现对圆锥角膜的后续临床治疗都具有一定的指导作用。 2.4 单细胞RNA测序技术在角膜上皮细胞中的应用 Pinin (PNN)是一种外显子连接成分[49],在角膜上皮细胞分化中起重要作用,是角膜上皮表型的稳定剂。AKIN团队[50]通过RNA测序发现,PNN可敲除大量与细胞迁移和细胞外基质重塑过程相关的上调基因,如MMPs,ADAMs,HAS2,LAMA3,CXCRs和UNC5C。此外,被敲除的基因还包括IGFBP5,FGD3,FGFR2,PAX6,RARG和SOX10[51]。近年来,有多项研究对角膜上皮干细胞标志物进行了选择。其中,一项研究使用单细胞RNA测序和可诱导的转基因小鼠脉冲追逐模型(K5Tta×treg-h2bgfp)对慢循环角膜细胞进行了定位、纯化和鉴定[52]。 SARTAJ团队[53]通过Smart-seq证实Krt15(角蛋白15为一种上皮干细胞表达的祖细胞标记物)、Fzd7和Krt17在角膜缘基底细胞中高表达,而Sox9,Actn1(细胞骨架蛋白)和Anxa3(属于膜联蛋白家族)的表达也延伸到少数基底上皮祖细胞[54]。Fzd7和Tcf3(二者皆为表皮干细胞标记物)在42 d后上调,干细胞相关基因干扰素诱导的跨膜蛋白1和Krt15也同样上调。但是截至目前,单细胞RNA测序技术在角膜领域的应用相对受限[38,42-43,50,53,55-56],见表4。"
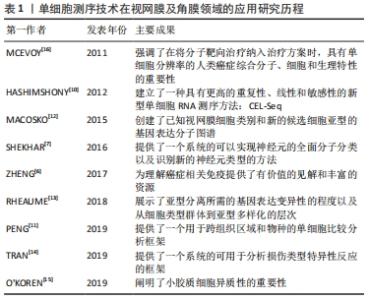
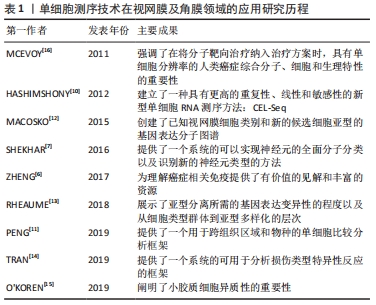
2.5 单细胞RNA测序技术在视网膜中的应用 对细胞类型特异性转录组的研究拓展了研究者对包括视网膜在内的异质组织中细胞功能的理解[57]。所有类型的细胞都有不同的功能和协调,以允许视觉感知和调节,因此,在细胞类型或单细胞水平上理解转录组将扩大与疾病相关的研究[58]。在视网膜中,许多细胞亚型,特别是稀有细胞的转录组图谱通常被大量RNA序列所掩盖。基于细胞表面标记的分选和纯化方法尚未被开发来丰富所有类型的细胞[59],因此,单细胞测序是获得视网膜中每种细胞类型的转录组的最有效和最公正的方法。作为一种具有强大性能的工具,单细胞RNA测序技术可以通过生成单细胞转录组图来解码复杂组织的异质 性[60]。因此,该方法可用于识别细胞特异性标记物,发现罕见细胞类型和细胞亚型,揭示细胞间的表达差异,是研究复杂生物系统细胞异质性的有效方法[61]。近年来,单细胞测序已陆续应用于研究小鼠、人类胎儿、灵长类动物和成人视网膜,对其细胞亚型以及相关的疾病基因通路有了相应的研究,这对于眼科临床治疗有着重要的意义。 众所周知,视网膜是一种易于获取和观察的组织,具有一定程度的神经元多样性,是研究细胞发育、神经元分化和神经系统多样性的理想模型[62]。此外,视网膜由小胶质细胞、无长突细胞、神经节细胞、双极细胞、感光细胞和视网膜色素上皮细胞等多种类型的细胞组成[63],对于如此复杂的组织结构,基于细胞异质性,传统的批量细胞RNA测序常检测细胞种群平均值来说明重要单细胞的变化,不能准确反映细胞间的特异性,但在疾病的发生发展中,单个细胞的特异性和异质性可能是最为关键的因素。与传统方法相比,单细胞RNA测序技术可以准确反映单个视网膜细胞的基因表达,在预防疾病中发挥重要作用[64]。细胞在高糖环境中的变化是一个动态的过程,同一细胞在不同阶段的基因表达也是不同的[65]。然而,单细胞RNA测序技术可以通过RNA在单细胞水平上的差异表达来分析转录异质性,并揭示给定复杂组织中先前未知的细胞类型或细胞状态。因此,单细胞RNA测序技术是获得视网膜各细胞类型转录组最有效的方法[65]。近年来,这项技术在眼科领域,特别是在视网膜方面的应用不断增加[66-73]。表5,6总结了近年来单细胞RNA测序技术在正常视网膜组织及视网膜相关疾病中的应用情况。"
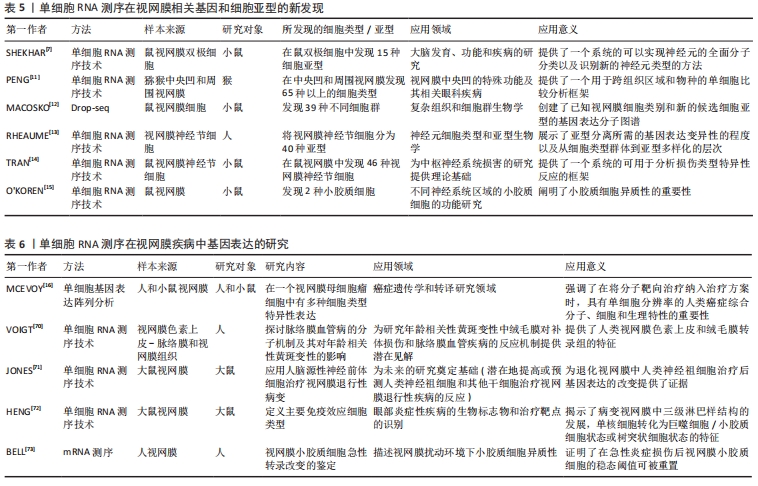

2.6 单细胞RNA测序技术在正常视网膜组织中的应用 细胞是生物结构和功能的基本单位,其类型和状态差异较大。每一个细胞都是生物系统形态和功能的基石,充分了解细胞类型及其功能特征对于确定其对正常组织功能和疾病状态进展的作用至关重要。单细胞RNA测序技术是一种从单个细胞的角度解决细胞异质性的技术。视网膜是一个高度不均匀的组织,估计其中含有100多种神经细胞亚型[66]。仅测量平均转录组图谱会使细胞功能的惟一性变得模糊不定,不能揭示促进疾病发展的因素。细胞的惟一性正是维持正常组织功能的关键。 单细胞RNA测序技术可在不同的动物模型中发现新细胞亚型。有学者在对灵长类猕猴的研究中,发现中央凹和周围视网膜中含有超过65种细胞类型,大多数类型对应不同的区域,但在比例和基因表达上存在差异[11]。此外,MACOSKO团队[12]在对小鼠视网膜细胞的研究中,也发现了39个不同的细胞群。 有研究在原有30种视网膜神经节细胞亚型的基础上,利用单细胞RNA测序技术将视网膜神经节细胞分为40种[13,67]。值得注意的是,最近有研究者还根据电生理特性鉴定了40种小鼠视网膜神经节细胞[68],表明这40种视网膜神经节细胞亚型之间的分子差异有可能基于其不同的电生理功能。此外,TRAN等[14]发现小鼠视网膜中有46种视网膜神经节细胞。从以上对视网膜神经节细胞亚型的研究可以看出,随着单细胞RNA测序技术的不断应用和发展,将会有越来越多的细胞亚型被发现。 SHEKHAR等[7]在小鼠体内发现了15种视网膜双极细胞亚型。O’KOREN等[15]在对小鼠视网膜小胶质细胞的研究中发现了两种类型的小胶质细胞,分别位于外丛状层和内丛状层,两种类型的小胶质细胞在稳定状态下存在功能差异,而在疾病状态下,小胶质细胞向视网膜下间隙大规模迁移,粘附于视网膜色素上皮顶部,保护视网膜外层和视网膜色素上皮细胞层的结构完整性。 通过对正常视网膜组织的单细胞RNA测序研究,不难看出越来越多的新细胞亚型不断被发现,这些发现揭示了对组织细胞更深层次的认识,也为后续疾病的诊断和治疗提供了新的视角。 2.7 单细胞RNA测序技术在视网膜疾病中的应用 单细胞RNA测序技术在视网膜疾病中也有较多应用,特别是在眼部肿瘤的靶向治疗方面。肿瘤的分子和细胞特征可以指示肿瘤细胞的来源,为靶向治疗提供依据。视网膜母细胞瘤是一种婴幼儿眼内的恶性肿瘤,关于肿瘤细胞起源的争论已经持续了约一个世纪。最近,MCEVOY等[16]对视网膜母细胞瘤患者和小鼠的肿瘤细胞进行了单细胞基因表达阵列分析,发现单个视网膜母细胞瘤细胞具有多种细胞类型特异性表达,这一现象在正常的视网膜发育过程中并不存在,表明控制视网膜发育和构建不同细胞类型的信号通路在肿瘤发展过程中受到干扰。这一机制可能为视网膜母细胞瘤的治疗提供新的靶点。 此外,单细胞RNA测序技术还可以通过分析相关疾病的发病机制,发现新的生物学靶点和标记物,来研究和治疗视网膜退行性疾病。NEWMAN等[69]发现了新的老年性黄斑变性生物标志物和发病机制的基因表达特征,说明视网膜色素上皮脉络膜细胞性炎症反应是老年黄斑变性的核心特征。老年黄斑变性可能是一种具有共同免疫反应过程的单一疾病。调节免疫活动的基因以及许多其他新发现,都可能为老年黄斑变性导向治疗和诊断提供极具前景的新靶点。同样,VOIGT等[70]对视网膜色素上皮和脉络膜进行单细胞RNA测序,探索脉络膜血管疾病的分子机制及其对老年黄斑变性的影响,为老年黄斑变性的治疗提供潜在思路。JONES等[71]利用人脑源性神经前体细胞治疗视网膜退行性病变,其中单细胞RNA测序技术可用于增强或预测治疗反应。这些发现可促进对视网膜的进一步分析。同时,它们也表明单细胞RNA测序技术在某些复杂的遗传性疾病细胞类型的研究中有非常高的潜力。 此外,单细胞RNA测序技术在眼科免疫性疾病的研究中也发挥了重要作用。在世界范围内,视网膜自身免疫性炎症性疾病仍是致盲的主要原因。单细胞RNA测序技术已成为研究自发性、慢性和进行性自身免疫性葡萄膜炎视网膜细胞类型和基因表达模式的一种全面、客观的新方法。HENG等[72]用小鼠建立了自身免疫性葡萄膜炎视网膜炎模型,确定了主要的免疫效应细胞类型,揭示了病变视网膜中三级淋巴结构的发育,并表征了单核细胞向巨噬细胞/小胶质细胞或树突状细胞状态的转变。结果表明,基本上所有视网膜细胞类型都对γ干扰素有反应,它可能是疾病过程的一个组成部分;同时,也提示Müller胶质细胞可能是抗原提呈细胞。BELL等[73]研究了炎症刺激期间视网膜小胶质细胞的免疫稳态,利用mRNA测序技术来识别视网膜小胶质细胞的急性转录变化。 总体而言,单细胞RNA测序技术在视网膜研究中发挥了关键作用。利用这种技术,可以发现新的细胞亚型,识别单个视网膜细胞亚型的基因标记,辅助研究和定位与特定视觉功能相关的靶点,从而更深入地了解细胞功能和细胞异质性的来源,探索建立维持细胞多样性的基因网络。 2.8 单细胞RNA测序技术在其他眼科疾病中的应用 近年来,除了在视网膜和角膜上的应用外,单细胞RNA测序技术还被应用于晶状体和葡萄膜疾病。研究发现,表达Gata3转基因的胚胎在晶状体纤维细胞分化过程中明显表现出多种缺陷[74]。利用单细胞RNA测序技术发现GATA3可直接通过调控CdKn1b/p27和CdKn1c/p57的表达[55],以及直接或间接调控Bfsp2和DNA酶Ⅱβ的转录来调节晶状体的分化。 有研究对葡萄膜黑色素瘤这一高度转移性癌症进行研究,采用单细胞测序的方法对8个原发样本和3个转移样本进行分析,发现肿瘤细胞表现出新的亚克隆基因组复杂性和转录状态[75]。这种复杂的肿瘤和免疫细胞生态系统为葡萄膜黑色素瘤生物学提供了新的见解。值得一提的是,在一项针对葡萄膜黑色素瘤的靶向治疗研究中发现,通过单细胞RNA测序发现, SKP2在葡萄膜黑色素瘤细胞中高表达[56],敲除SKP2可抑制葡萄膜黑色素瘤细胞的增殖[76]。此外,SKP2被认为是一种与多种恶性肿瘤相关的致癌基因[77]。SKP2可抑制葡萄膜黑色素瘤细胞的增殖[78],这项新发现为葡萄膜黑色素瘤靶向治疗提供了新的选择和可能。"

| [1] CHEN G, NING B, SHI T. Single-cell RNA-seq technologies and related computational data analysis. Front Genet. 2019;10:317. [2] LIU Y, HU J, LiU D, et al. Single-cell analysis reveals immune landscape in kidneys of patients with chronic transplant rejection. Theranostics. 2020;10(19):8851-8862. [3] DING S, CHEN X, SHEN K. Single-cell RNA sequencing in breast cancer: Understanding tumor heterogeneity and paving roads to individualized therapy. Cancer Commun. 2020;40(8):329-344. [4] ZHANG Y, WANG D, PENG M, et al. Single-cell RNA sequencing in cancer research. J Exp Clin Cancer Res. 2021;40(1):81. [5] SUN YF, GUO W, XU Y, et al. Circulating tumor cells from different vascular sites exhibit spatial heterogeneity in epithelial and mesenchymal composition and distinct clinical significance in hepatocellular carcinoma. Clin Cancer Res. 2018;24(3):547-559. [6] ZHENG C, ZHENG L, YOO JK, et al. Landscape of infiltrating T cells in liver cancer revealed by single-cell sequencing. Cell. 2017;169(7):1342-1356. [7] SEHKHAR K, LAPAN SW, WHITNEY IE, et al. Comprehensive classification of retinal bipolar neurons by single-cell transcriptomics. Cell. 2016; 166(5):1308-1323. [8] KAPLAN N, WANG J, WRAY B, et al. Single-cell RNA transcriptome helps define the limbal/corneal epithelial stem/early transit amplifying cells and how autophagy affects this population. Invest Ophthalmol Vis Sci. 2019;60(10):3570-3583. [9] KAMEISHI S, UMEMOTO T, MATSUZAKI Y, et al. Characterization of rabbit limbal epithelial side population cells using RNA sequencing and single-cell qRT-PCR. Biochem Biophys Res Commun. 2016;473(3):704-709. [10] HASHIMSHONY T, WAGNER F, SHER N, et al. CEL-Seq: single-cell RNA-Seq by multiplexed linear amplification. Cell Rep. 2012;2:666-673. [11] PENG YR, WANG Y, HERRMANN D, et al. Molecular classification and comparative taxonomics of foveal and peripheral cells in primate retina. Cell. 2019;176(5):1222-1237. [12] MACOSKO EZ, SATIJA R, NEMESH J, et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell. 2015;161(5):1202-1214. [13] Rheaume BA, Jereen A, Bolisetty M, et al. Single cell transcriptome profiling of retinal ganglion cells identifies cellular subtypes. Nat Commun. 2018;9(1):2759. [14] TRAN NM, SHEKHAR K, WHITNEY IE, et al. Single-cell profiles of retinal ganglion cells differing in resilience to injury reveal neuroprotective genes. Neuron. 2019;104(6):1039-1055. [15] O’KOREN EG, YU C, KLINGEBORN M, et al. Microglial function is distinct in different anatomical locations during retinal homeostasis and degeneration. Immunity. 2019;50(3):723-737. [16] MCEVOY J, FLORES-OTERO J, ZHANG J, et al. Coexpression of normally incompatible developmental pathways in retinoblastoma genesis. Cancer Cell. 2011;20(2):260-275. [17] HWANG B, LEE JH, BANG D. Single-cell RNA sequencing technologies and bioinformatics pipelines. Exp Mol Med. 2018;50(8):1-14. [18] YE Y, SONG H, ZHANG J, et al. Understanding the biology and pathogenesis of the kidney by single-cell transcriptomic analysis. Kidney Dis (Basel). 2018;4(4):214-225. [19] BRADY G, ISCOVE N. Representative in vitro cDNA amplification from individual hemopoietic cells and colonies. Methods Mol Cell Biol. 1990;2(1):17-25. [20] EBERWINE J, MIYASHIRO K, CAO Y, et al. Analysis of gene expression in single live neurons. Proc Natl Acad Sci U S A. 1992;89(7):3010-3014. [21] TANG F, WANG Y, NORDMAN E, et al. mRNA-Seq whole-transcriptome analysis of a single cell. Nat Methods. 2009;6(5):377-382. [22] RAMSKOLD D, WANG YC, LI R, et al. Full-length mRNA-Seq from single-cell levels of RNA and individual circulating tumor cells. Nat Biotechnol. 2012;38(3):374. [23] PICELLI S, BJORKLUND AK, WINBERG G, et al. Full-length RNA-seq from single cells using Smart-seq2. Nat Protoc. 2012;9(1):171-181. [24] SASAGAWA Y, HAYASHIT, DANNO H, et al. Quartz-Seq: a highly reproducible and sensitive single-cell RNA sequencing method, reveals non-genetic gene-expression heterogeneity. Genome Biol. 2013;14(4):31. [25] SEE P, CHEN J, GINHOUX F. A single-cell sequencing guide for immunologists. Front Immunol. 2018;9:2425. [26] GAO S. Data analysis in single-cell transcriptome sequencing. Methods Mol Biol. 2018;1754:311-326. [27] VALDES-MORA F, HANDLER K, LAW AMK, et al. Single-cell transcriptomics in cancer immunobiology: the future of precision oncology. Front Immunol. 2018;9:2582. [28] ISLAM S, MOLINER A, ZAJAC P, et al. Highly multiplexed and strand-specific single-cell RNA 5’end sequencing. Nat Protoc. 2012;7(5):813-828. [29] HASHIMSHONY T, SENDEROVICH N, AVITAL G, et al. CEL-Seq2: sensitive highly-multiplexed single-cell RNA-Seq. Genome Biol. 2016;17:77. [30] Method of the Year 2019: single-cell multimodal omics. Nat Methods. 2020;17(1):1. [31] RUBEN M, DALIA B, FLORIAN W, et al. Integrating microarray-based spatial transcriptomics and single-cell RNA-seq reveals tissue architecture in pancreatic ductal adenocarcinomas. Nat Biotech. 2020;38(3):333-342. [32] SKORCZYK-WERNER A, CHIANG WC, WAWROCKA A, et al. Autosomal recessive cone-rod dystrophy can be caused by mutations in the ATF6 gene. Eur J Hum Genet. 2017;25(11):1210-1216. [33] LEE EJ, CHIANG WJ, KROEGER H, et al. Multiexon deletion alleles of ATF6 linked to achromatopsia. JCI Insight. 2020. doi:10.1172/jci.insight.136041. [34] YOU J, HODGE C, WEN L, et al. Tear levels of SFRP1 are significantly reduced in keratoconus patients. Mol Vis. 2013;19:509. [35] HAYES S, BOOTE C, TUFY SJ, et al. A study of corneal thickness, shape and collagen organisation in keratoconus using videokeratography and X-ray scattering techniques. Exp Eye Res. 2007;84(3):423-434. [36] KANDEL H, PESUDOVSK, WATSON SL. Measurement of quality of life in keratoconus. Cornea. 2020;39(3):386-393. [37] FERRARI G, RAMA P. The keratoconus enigma: a review with emphasis on pathogenesis. Ocul Surf. 2020;18(3):363-373. [38] YOU J, CORLEY SM, WEN L, et al. RNA-Seq analysis and comparison of corneal epithelium in keratoconus and myopia patients. Sci Rep. 2018; 8(1):389. [39] NAJMI H, MOBARKI Y, MANIA K, et al. The correlation between keratoconus and eye rubbing: a review. Int J Ophthalmol. 2019;12(11):1775-1781. [40] BYKHOVSKAYA Y, RABINOWITA YS. Update on the genetics of keratoconus. Exp Eye Res. 2021;202:108398. [41] SLOVIN S, CARISSIMO A, PANARIELLO F, et al. Single-cell rna sequencing analysis:a step-by-step overview. Methods Mol Biol. 2021;2284:343-365. [42] SZCZESNIAK MW, KABZA M, KAROLAK JA, et al. KTCNlncDB-a first platform to investigate lncRNAs expressed in human keratoconus and non-keratoconus corneas. Database. 2017. doi:10.1093/database/baw168. [43] KHALED ML, BYKHOVSKAYA Y, YABLONSKI SER, et al. Differential expression of coding and long noncoding rnas in keratoconus-affected corneas. Invest Ophthalmol Vis Sci. 2018;59(7):2717-2728. [44] SHETTY R, SATHYANARAYANAMOORTHY A, RAMACHANDRA RA, et al. Attenuation of lysyl oxidase and collagen gene expression in keratoconus patient corneal epithelium corresponds to disease severity. Mol Vis. 2015;21:12-25. [45] MERCER TR, GERHARDT DJ, DINGER ME, et al. Targeted RNA sequencing reveals the deep complexity of the human transcriptome. Nat Biotechnol. 2011;30(1):99-104. [46] PERTEA M, PERTEA GM, ANTONESCU CM, et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat Biotechnol. 2015;33(3):290-295. [47] BHENDE SD, VaARANASI MB, ABBULU K. A sensitive HPTLC method for the estimation of glibenclamide, rosiglitazone maleate and metformin hydrochloride from a multicomponent dosage form. J Chromatogr Sci. 2020;58(5):418-426. [48] MANABE A, FURUKAWA C, HASEGAWA H, et al. Upregulation of transient receptor potential melastatin 6 channel expression by rosiglitazone and all-trans-retinoic acid in erlotinib-treated renal tubular epithelial cells. J Cell Physiol. 2019;234(6):8951-8962. [49] NOTARA M, SHORTT AJ, O’CALAGHAN AR, et al. The impact of age on the physical and cellular properties of the human limbal stem cell niche. Age. 2013;35(2):289-300. [50] AKIN D, NEWMAN JR, MCINTYRE LM, et al. RNA-seq analysis of impact of PNN on gene expression and alternative splicing in corneal epithelial cells. Mol Vis. 2016;22:40-60. [51] TANG T, YANG L, CAO Y, et al. LncRNA AATBC regulates Pinin to promote metastasis in nasopharyngeal carcinoma. Mol Oncol. 2020;14(9):2251-2270. [52] OKUNURA N, KAY EP, NAKAHARA M, et al. Inhibition of TGF-β signaling enables human corneal endothelial cell expansion in vitro for use in regenerative medicine. PLoS One. 2013;8(2):e58000. [53] SARTAJ R, ZHANG C, WAN P, et al. Characterization of slow cycling corneal limbal epithelial cells identifies putative stem cell markers. Sci Rep. 2017;7(1):3793. [54] DENG SX, LEE WB, HAMMERSMIT KM, et al. Descemet membrane endothelial keratoplasty: safety and outcomes: a report by the american academy of ophthalmology. Ophthalmology. 2018;125(2):295-310. [55] MARTYNOVA E, ZHAO Y, XIE Q, et al. Transcriptomic analysis and novel insights into lens fibre cell differentiation regulated by Gata3. Open Biol. 2019;9(12):190220. [56] ZHAO H, PAN H, WANG H, et al. SKP2 targeted inhibition suppresses human uveal melanoma progression by blocking ubiquitylation of p27. Onco Targets Ther. 2019;12:4297-4308. [57] MAYER S, VELMESHEV D, MAYER A, et al. Multimodal single-cell analysis reveals physiological maturation in the developing human neocortex. Neuron. 2019;102(1):143-158. [58] LIANG Q, OWEN L, SHAKOOR A, et al. Single-nuclei RNA-seq on human retinal tissue provides improved transcriptome profiling. Nat Commun. 2019;10(1):5743. [59] HARSHINI C, VASUDHARANI D. Molecular mechanisms mediating diabetic retinal neurodegeneration: potential research avenues and therapeutic targets. J Mol Neurosci. 2018;66(3):445-461. [60] TANG F, BARBACIORU C, BAO S, et al. Tracing the derivation of embryonic stem cells from the inner cell mass by single-cell RNA-Seq analysis. Cell Stem Cell. 2010;6(5):468-478. [61] PAPALEXI E, SATIJA R. Single-cell RNA sequencing to explore immune cell heterogeneity. Nat Rev Immunol. 2018;18(1):35-45. [62] MASLAND RH. The neuronal organization of the retina. Neuron. 2012; 76(2):266-280. [63] HARTL D, KREBS AR, JUTTNER J, et al. Cis-regulatory landscapes of four cell types of the retina. Nucleic Acids Res. 2017;45(20):11607-11621. [64] POTTER SS. Single-cell RNA sequencing for the study of development, physiology and disease. Nat Rev Nephrol. 2018;14(8):479-492. [65] LIEVENS B, HALLSWORTH JE, POZO MI, et al. Microbiology of sugar-rich environments:diversity, ecology and system constraints. Environ Microbiol. 2015;17(2):278-298. [66] DHARMAT R, KIM S, LI Y, et al. Single-cell capture, RNA-seq, and transcriptome analysis from the neural retina. Methods Mol Biol. 2020; 2092:159-186. [67] JOSHUA R, RICHARD H, MASLAND. The types of retinal ganglion cells: current status and implications for neuronal classification. Annu Rev Neurosci. 2015;38:221-246. [68] BADEN T, BERENS P, FRANKE K, et al. The functional diversity of retinal ganglion cells in the mouse. Nature. 2016;529(7586):345-350. [69] NEWMAN AM, GALLO NB, HANCOX LS, et al. Systems-level analysis of age-related macular degeneration reveals global biomarkers and phenotype-specific functional networks. Genome Med. 2012;4(2):16. [70] VOIGT AP, MULFAUL K, MULLIN NK, et al. Single-cell transcriptomics of the human retinal pigment epithelium and choroid in health and macular degeneration. Proc Natl Acad Sci U S A. 2019;116(48):24100-24107. [71] JONES MK, LU B, SAHIZADEH M, et al. Gene expression changes in the retina following subretinal injection of human neural progenitor cells into a rodent model for retinal degeneration. Mol Vis. 2016;22:472-490. [72] HENG JS, HACKETT SF, STEIN-O’BRIEN GL, et al. Comprehensive analysis of a mouse model of spontaneous uveoretinitis using single-cell RNA sequencing. Proc Natl Acad Sci. 2019;116(52):26734-26744. [73] BELL OH, COPLAND DA, WARD A, et al. Single eye mRNA-Seq reveals normalisation of the retinal microglial transcriptome following acute inflammation. Front Immunol. 2019;10:3033. [74] YOUNG MD, WAKEFILED MJ, SMYTH GK, et al. Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol. 2010;11(2):R14. [75] Durante MA, Rodriguez DA, Kurtenbach S, et al. Single-cell analysis reveals new evolutionary complexity in uveal melanoma. Nat Commun. 2020;11(1):496. [76] BULLARD JH, PUEDOM E, HANSEN KD, et al. Evaluation of statistical methods for normalization and differential expression in mRNA-Seq experiments. BMC Bioinformatics. 2010;11:94. [77] CHEN G, LI R, SHI L, et al. Revealing the missing expressed genes beyond the human reference genome by RNA-Seq. BMC Genomics. 2011;12:590. [78] PENG Z, CHENG Y, TAN BC, et al. Comprehensive analysis of RNA-Seq data reveals extensive RNA editing in a human transcriptome. Nat Biotechnol. 2012;30(3):253-260. [79] GORDON-SHAAG A, MILLODOT M, SHNEOR E, et al. The genetic and environmental factors for keratoconus. Biomed Res Int. 2015; 2015:795738. [80] KARAMICHOS D, ZIESKE JD, SEJERSEN H, et al. Tear metabolite changes in keratoconus. Exp Eye Res. 2015;132:1-8. |
| [1] | Cai Xiaoxuan, Lü Yingnian, Qi Yi. Long-circulating lipsomes: application and mechanism [J]. Chinese Journal of Tissue Engineering Research, 2022, 26(16): 2613-2617. |
| [2] | Gong Yubo, Zhang Wenqian, Guo Xiaohua, Yan Jin, Song Feilong, Shi Yuanyuan, Luo Ling, Zhao Jun, Zhao Hongwei. Methazolamide effects on intraocular pressure, retinal and choroidal thickness in rats with tail suspension for 2 weeks [J]. Chinese Journal of Tissue Engineering Research, 2022, 26(14): 2238-2242. |
| [3] | Li Jianyi, Liu Zhiyuan, Deng Chengliang. Function and hot spot of adipose-derived stem cell exosomes in tissue regeneration and repair [J]. Chinese Journal of Tissue Engineering Research, 2022, 26(13): 2093-2098. |
| [4] | Li Shanshan, Guo Xiaoxiao, You Ran, Yang Xiufen, Zhao Lu, Chen Xi, Wang Yanling. Photoreceptor cell replacement therapy for retinal degeneration diseases [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(7): 1116-1121. |
| [5] | Yang Xin, Jin Zhe, Feng Xu, Lu Bing. The current situation of knowledge and attitudes towards organ, eye tissue, body donation of residents in Shenyang [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(5): 779-784. |
| [6] | Liu Jingchen, Li Mingjun, Sun Shuguang, Li Cairui. Application of acellular tissue in ophthalmology [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(34): 5530-5536. |
| [7] | Liu Rixu, Lü Wenbo, Gao Wenshan. Research and clinical application of calcium phosphate cement modification [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(34): 5537-5543. |
| [8] | Zhang Jichao, Dong Wanpeng, Dong Yuefu, Zhang Zhen, Li Jiayi, Ren Meng. Parameters of finite element model of the knee joint [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(30): 4781-4786. |
| [9] | Yang Menglu, Zhang Na, Wang Fangyuan, Liu Jianguo. Application of chitosan in nano drug delivery system [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(28): 4546-4552. |
| [10] | Zhu Bikang, Luo Shanchao, Luo Shixing. Progress and main application of fish skin collagen in tissue engineering [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(28): 4561-4566. |
| [11] | Zhu Bin, Yan Yiguo. Application of single-cell RNA sequencing technology in the study of skeletal system biology [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(26): 4204-4210. |
| [12] | Wang Xuan, Zhou Chao, Zhang Yingzi. Research, application and development of human amniotic epithelial cells in the field of obstetrics and gynecology [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(25): 4070-4075. |
| [13] | Li Shanshan, You Ran, Guo Xiaoxiao, Zhao Lu, Wang Yanling, Chen Xi. Advances in the mechanisms of optic nerve regeneration [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(23): 3740-3745. |
| [14] | Xu Bin, Yang Xiushu, Liu Xuan, Wang Zhenxing. Changes of intestinal epithelial cells and their apoptotic factors Caspase-3, Bax and Bcl-2 under urinary environment [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(20): 3173-3177. |
| [15] | Yan Xiurui, Tao Jin, Liang Xueyun. Mechanism by which exosomes from human fetal placental mesenchymal stem cells protect lung epithelial cells against oxidative stress injury [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(19): 2994-2999. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||