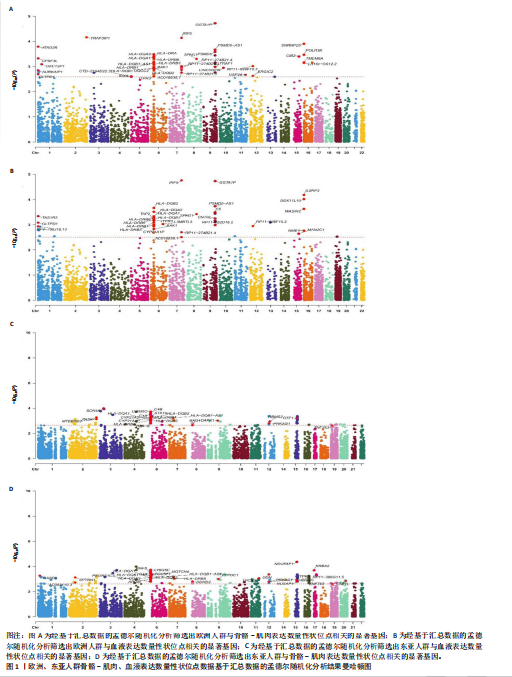
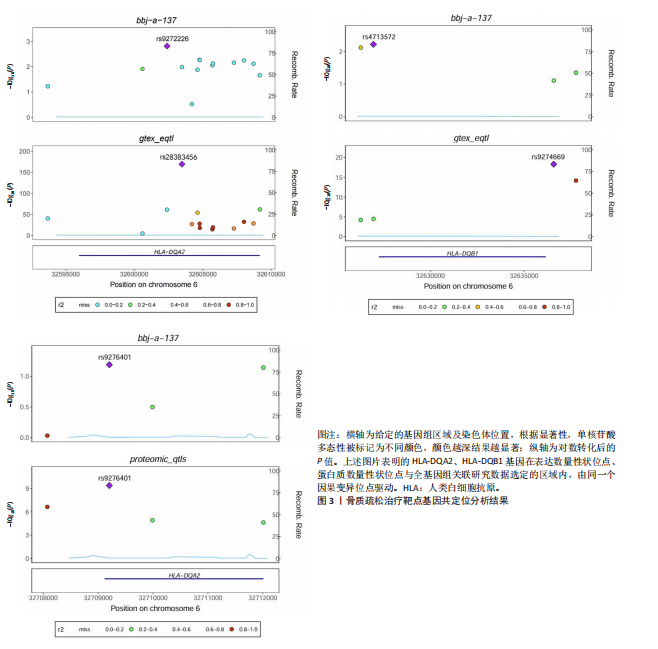
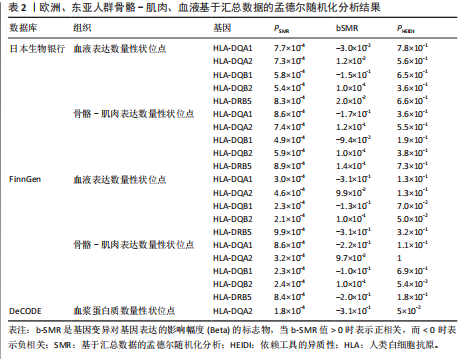
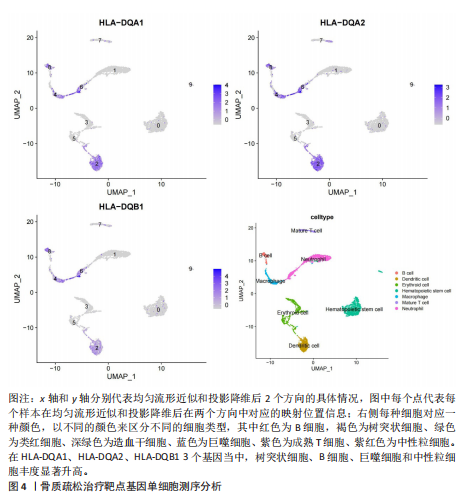
[1] TROJNIAK J, SENDERA A, BANAŚ-ZĄBCZYK A, et al. The MicroRNAs in the Pathophysiology of Osteoporosis. Int J Mol Sci. 2024;25(11):6240.
[2] ANAM AK, INSOGNA K. Update on Osteoporosis Screening and Management. Med Clin North Am. 2021;105:1117-1134.
[3] MORRIS JA, KEMP JP, YOULTEN SE, et al. An atlas of genetic influences on osteoporosis in humans and mice. Nat Genet. 2019;51:258-266.
[4] ESTRADA K, STYRKARSDOTTIR U, EVANGELOU E, et al. Genome-wide meta-analysis identifies 56 bone mineral density loci and reveals 14 loci associated with risk of fracture. Nat Genet. 2012;44:491-501.
[5] KEMP JP, MORRIS JA, MEDINA-GOMEZ C, et al. Identification of 153 new loci associated with heel bone mineral density and functional involvement of GPC6 in osteoporosis. Nat Genet. 2017;49:1468-1475.
[6] PIVIDORI M, RAJAGOPAL PS, BARBEIRA A, et al. PhenomeXcan: mapping the genome to the phenome through the transcriptome. Sci Adv. 2020;6:eaba2083.
[7] GU XJ, SU WM, DOU M, et al. Expanding causal genes for Parkinson’s disease via multi-omics analysis. NPJ Parkinsons Dis. 2023;9(1):146.
[8] AL-BARGHOUTHI BM, ROSENOW WT, DU KP, et al. Transcriptome-wide association study and eQTL colocalization identify potentially causal genes responsible for human bone mineral density GWAS associations. Elife. 2022;11:e77285.
[9] SHIN HD, PARK BL, SHIN HJ, et al. Association of KCNQ1 polymorphisms with the gestational diabetes mellitus in Korean women. J Clin Endocrinol Metab. 2010;95(1):445-449.
[10] CAO M, ZHANG L, CHEN T, et al. Genetic susceptibility to gestational diabetes mellitus in a Chinese population. Front Endocrinol (Lausanne). 2020;11:247.
[11] YANG Y, HU P, ZHANG Q, et al. Single-cell and genome-wide Mendelian randomization identifies causative genes for gout. Arthritis Res Ther. 2024; 26(1):114.
[12] WU J, SUN X, WU C, et al. Single-cell transcriptome analysis reveals liver injury induced by glyphosate in mice. Cell Mol Biol Lett. 2023;28(1):11.
[13] JACOBS BM, TAYLOR T, AWAD A, et al. Summary-data-based Mendelian randomization prioritizes potential druggable targets for multiple sclerosis. Brain Commun. 2020;2(2):fcaa119.
[14] GTEX CONSORTIUM. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science. 2020;369(6509):1318-1330.
[15] LYON MS, ANDREWS SJ, ELSWORTH B, et al. The variant call format provides efficient and robust storage of GWAS summary statistics. Genome Biol. 2021;22(1):32.
[16] ISHIGAKI K, AKIYAMA M, KANAI M. Large-scale genome-wide association study in a Japanese population identifies novel susceptibility loci across different diseases. Nat Genet. 2020;52(7): 669-679.
[17] FERKINGSTAD E, SULEM P, ATLASON BA, et al. Large-scale integration of the plasma proteome with genetics and disease. Nat Genet. 2021; 53(12):1712-1721.
[18] SKRIVANKOVA VW, RICHMOND RC, WOOLF BAR, et al. Strengthening the reporting of observational studies in epidemiology using Mendelian randomization: the STROBE-MR statement. JAMA. 2021;326(16):1614-1621.
[19] WU Y, ZHANG CY, WANG L, et al. Genetic Insights of Schizophrenia via Single Cell RNA-Sequencing Analyses. Schizophr Bull. 2023;49(4):914-922.
[20] BAI Y, WANG J, FENG X, et al. Identification of drug targets for Sjögren’s syndrome: multi-omics Mendelian randomization and colocalization analyses. Front Immunol. 2024;15:1419363.
[21] ZHU Z, ZHANG F, HU H, et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat Genet. 2016;48(5):481-487.
[22] WU Y, ZENG J, ZHANG F, et al. Integrative analysis of omics summary data reveals putative mechanisms underlying complex traits. Nat Commun. 2018;9(1):918.
[23] HUANG X, SHEN R, ZHENG Z. Unraveling genetic threads: Identifying novel therapeutic targets for allergic rhinitis through Mendelian randomization. World Allergy Organ J. 2024; 17(7):100927.
[24] WANG Z, LI X, YANG J, et al. Single-cell RNA sequencing deconvolutes the in vivo heterogeneity of human bone marrow-derived mesenchymal stem cells. Int J Biol Sci. 2021; 17(15):4192-4206.
[25] CAO G, XUAN X, LI Y, et al. Single-cell RNA sequencing reveals the vascular smooth muscle cell phenotypic landscape in aortic aneurysm. Cell Commun Signal. 2023;21(1):113.
[26] SRIVASTAVA RK, SAPRA L, MISHRA PK. Osteometabolism: Metabolic Alterations in Bone Pathologies. Cells. 2022;11(23):3943.
[27] CHAPMAN NM, BOOTHBY MR, CHI H. Metabolic coordination of T cell quiescence and activation. Nat Rev Immunol. 2020;20(1):55-70.
[28] SCHIRMER M, SMEEKENS SP, VLAMAKIS H, et al. Linking the Human Gut Microbiome to Inflammatory Cytokine Production Capacity. Cell. 2016;167(4):1125-1136.e8.
[29] YASUDA H. Discovery of the RANKL/RANK/OPG system. J Bone Miner Metab. 2021;39(1):2-11.
[30] SRIVASTAVA S, RASOOL M. Genetics, epigenetics and autoimmunity constitute a Bermuda triangle for the pathogenesis of rheumatoid arthritis. Life Sci. 2024;357: 123075.
[31] 中山大学.中华民族人MHC(HLA)二类基因多态性及其与疾病相关性研究.2006.
[32] 北京市结核病胸部肿瘤研究所. HLA-DR、DQ基因与Ⅰ型糖尿病(IDDM)易感性研究.2002.
[33] ZHANG Z, CHEN H, WANG XP, et al. Association between major histocompatibility complex class II molecules and postmenopausal osteoporosis. Chin J Osteoporos. 2024;30(2):270-274.
[34] LIU H, HE J, BAGHERI-YARMAND R, et al. Osteocyte CIITA aggravates osteolytic bone lesions in myeloma. Nat Commun. 2022;13(1):3684.
[35] BENASCIUTTI E, MARIANI E, OLIVA L, et al. MHC class II transactivator is an in vivo regulator of osteoclast differentiation and bone homeostasis co-opted from adaptive immunity. J Bone Miner Res. 2014;29(2):290-303.
[36] WANG X, ZHANG X, HAN Y, et al. Role of the major histocompatibility complex class II protein presentation pathway in bone immunity imbalance in postmenopausal osteoporosis. Front Endocrinol (Lausanne). 2022;13:876067.
[37] SUN X, CHEN B, QI Y, et al. Multi-omics Mendelian randomization integrating GWAS, eQTL and pQTL data revealed GSTM4 as a potential drug target for migraine. J Headache Pain. 2024;25(1):117.
[38] MA Y, ZHOU Y, JIANG D, et al. Integration of human organoids single-cell transcriptomic profiles and human genetics repurposes critical cell type-specific drug targets for severe COVID-19. Cell Prolif. 2024;57(3):e13558.
|