• 组织构建学术探讨 tissue construction academic discussion • 上一篇
网络医学的架构及其研究进展
黎 英1,黄文聪2
- 1广西师范学院广西高校科学计算与智能信息处理重点实验室,广西壮族自治区南宁市 530001;2广西师范学院体育学院,广西壮族自治区南宁市 530001
-
收稿日期:2016-12-12出版日期:2017-04-28发布日期:2017-05-16 -
作者简介:黎英,女,1973年生,博士,副教授,主要从事生物计算、网络医学研究。 -
基金资助:广西高校科学技术研究项目(KY2015YB188)
Architecture of network medicine and its research progress
Li Ying1, Huang Wen-cong2
- 1Guangxi Colleges and Universities Key Laboratory of Scientific Computing &
Intelligent Information Processing of Guangxi Teachers Education University, Nanning 530001, Guangxi Zhuang Autonomous Region, China; 2School of Sports of Guangxi Teachers Education University, Nanning 530001,Guangxi Zhuang Autonomous Region, China
-
Received:2016-12-12Online:2017-04-28Published:2017-05-16 -
About author:Li Ying, Ph.D., Associate professor, Guangxi Colleges and Universities Key Laboratory of Scientific Computing & Intelligent Information Processing of Guangxi Teachers Education University, Nanning 530001, Guangxi Zhuang Autonomous Region, China -
Supported by:the Scientific Research Foundation of the Higher Education Institutions of Guangxi Zhuang Autonomous Region, China, No. KY2015YB188
摘要:
.jpg) 文题释义:
网络医学:网络医学使用网络科学来诊断、预防和治疗疾病,它提供一个量化的平台来解决人类疾病的复杂性。与医学有关的网络以简单原则组织起来,这些组织原则包括度分布和枢纽、小世界现象、模式(motif)、模块、介数中心性。这些网络的节点代表生物因素(生物分子,疾病,表型等),连接(边)则代表它们的关系(物理相互作用,共享的代谢途径,共享基因,共享性状等)。
Interactome:又称为分子相互作用网络。在这类网络中,节点是代谢物和诸如蛋白质、RNA分子和基因序列这样的分子,而边是可以用大量技术来识别的物理、生物化学和功能相互作用。在Interactome中,代谢网络、蛋白质-蛋白质相互作用网络和基因调控网络提供了关于细胞系统的关键的“骨架”信息,在这个“骨架”上还可以添加功能连接的额外层(转录谱网络、表型谱网络和基因相互作用网络)来调整生物的现实表现。
文题释义:
网络医学:网络医学使用网络科学来诊断、预防和治疗疾病,它提供一个量化的平台来解决人类疾病的复杂性。与医学有关的网络以简单原则组织起来,这些组织原则包括度分布和枢纽、小世界现象、模式(motif)、模块、介数中心性。这些网络的节点代表生物因素(生物分子,疾病,表型等),连接(边)则代表它们的关系(物理相互作用,共享的代谢途径,共享基因,共享性状等)。
Interactome:又称为分子相互作用网络。在这类网络中,节点是代谢物和诸如蛋白质、RNA分子和基因序列这样的分子,而边是可以用大量技术来识别的物理、生物化学和功能相互作用。在Interactome中,代谢网络、蛋白质-蛋白质相互作用网络和基因调控网络提供了关于细胞系统的关键的“骨架”信息,在这个“骨架”上还可以添加功能连接的额外层(转录谱网络、表型谱网络和基因相互作用网络)来调整生物的现实表现。中图分类号:
引用本文
黎 英,黄文聪. 网络医学的架构及其研究进展[J]. 中国组织工程研究, doi: 10.3969/j.issn.2095-4344.2017.12.026.
Li Ying1, Huang Wen-cong2. Architecture of network medicine and its research progress[J]. Chinese Journal of Tissue Engineering Research, doi: 10.3969/j.issn.2095-4344.2017.12.026.
图2是提供“骨架”信息的3个网络,包括蛋白质相互作用网络、代谢网络和基因调控网络。
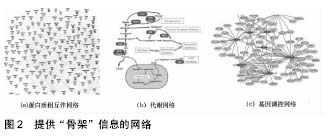
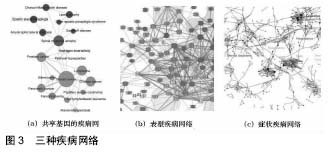
图3是3种疾病网络,分别是共享基因的疾病网络、表型疾病网络和症状疾病网络。
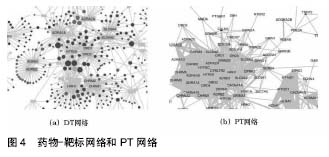
在“靶标-蛋白质网络”(Target-Protein network,PT网络)中,节点代表蛋白质,如果2种蛋白质都被至少一种相同药物靶向,那么它们被连接[92]。靶标-蛋白质网络提供了一种补充,以蛋白质为中心的药理学观点。在靶标-蛋白质网络中,394种中的305种靶标蛋白质和其他靶标蛋白质相连。多靶标药物是造成靶标-蛋白质网络高度相互连接的原因。图4显示了药物-靶标网络和PT网络。
| [1] Network medicine. From Wikipedia, the free encyclopedia. https://en.wikipedia.org/wiki/Network_medicine[2] Barabási AL, Gulbahce N, Loscalzo J. Network medicine: a network-based approach to human disease. Nat Rev Genet. 2011;12(1):56-68.[3] Barabási AL. Network medicine-from obesity to the “diseasome”. N Engl J Med. 2007;357(4):404-407.[4] Chan SY, Loscalzo J. The emerging paradigm of network medicine in the study of human disease. Circulation Res. 2012;111(3), 359-374.[5] Sanchez C, Lachaize C, Janody F, et al."Grasping at molecular interactions and genetic networks in Drosophila melanogaster using FlyNets, an Internet database". Nucleic Acids Res. 1999;27(1): 89-94. [6] Interactome.From Wikipedia, the free encyclopedia. https://en.wikipedia.org/wiki/Interactome[7] Vidal M, Cusick ME, Barabási AL.Interactome Networks and Human Disease. Cell. 2011; 144(6): 986-998.[8] Gunsalus KC, Ge H, Schetter AJ, et al.Predictive models of molecular machines involved in Caenorhabditis elegans early embryo genesis. Nature.2005;436, 861-865.[9] Rigaut G, Shevchenko A, Rutz B,et al. A generic protein purification method for protein complex characterization and proteome exploration. Nat. Biotechnol. 1999; 17:1030-1032.[10] Charbonnier S, Gallego O, Gavin AC. The social network of acell: recent advances in interactome mapping. Biotechnol Annu Rev. 2008;14:1-28.[11] Kuhner S, van Noort V, Betts MJ, et al. Proteome organization in a genome-reduced bacterium. Science.2009;326:1235-1240.[12] Seebacher J, Gavin AC. SnapShot: Protein-protein interaction networks. Cell. Cell. 2011;144(6):1000, [13] RRual JF, Venkatesan K, Hao T, et al. Towards a proteome-scale map of the human protein-protein interaction network. Nature. 2005; 437(7062):1173-1178.[14] Stelzl U, Worm U, Lalowski M, et al. A human protein-protein interaction network: a resource for annotating the proteome. Cell.2005;122(6):957-968.[15] Sun J, Zhao Z.A comparative study of cancer proteins in the human protein-protein interaction network.BMC Genomics. 2010 Dec 1;11 Suppl 3:S5.[16] Schaefer MH, Lopes TJ, Mah N, et al. Adding Protein Context to the Human Protein-Protein Interaction Network to Reveal Meaningful Interactions. PLoS Comput Biol. 2013;9(1): e1002860.[17] Rolland T, Ta?an M, Charloteaux B,et al.A Proteome-Scale Map of the Human Interactome Network.Cell. 2014;159(5):1212-1226.[18] Venkatesan K, Rual JF, Vazquez A, et al. An empirical framework for binary interactome mapping. Nat. Methods. 2009;6: 83-90[19] Costanzo M, Baryshnikova A, Bellay J,et al. The genetic landscape of a cell. Science.2010; 327:425-431.[20] Li BQ, Huang T, Liu L,et al. Identification of Colorectal Cancer Related Genes with mRMR and Shortest Path in Protein-Protein Interaction Network. PLoS ONE.2012;7(4): e33393.[21] Paul Y, Hasija Y. Gene Prioritization by Integrated Analysis of Protein Structural and Network Topological Properties for the Protein-Protein Interaction Network of Neurological Disorders. Scientifica (Cairo). 2016;2016:9589404.[22] Shityakov S, Dandekar T, Forster C. Gene expression profiles and protein-protein interaction network analysis in AIDS patients with HIV-associated encephalitis and dementia. HIV AIDS(Auckl). 2015;7:265-276. [23] Safaei A, Rezaei Tavirani M, Arefi Oskouei A,et al. Protein-protein interaction network analysis of cirrhosis liver disease. Gastroenterol Hepatol Bed Bench. 2016;9(2):114-233.[24] Shityakov S, Dandekar T, Förster C.Gene expression profiles and protein–protein interaction network analysis in AIDS patients with HIV-associated encephalitis and dementia.HIV AIDS (Auckl). 2015;7:265-276. [25] vinayagam A,Gibson TE,Yilmazel B,et al.Controllability analysis of the directed human protein interaction network identifies disease genes and drug targets. Proc Natl Acad Sci USA. 2016;113(18):4976-4981. [26] Chapple CE, Robisson B, Spinelli L, et al. Extreme multifunctional proteins identified from a human protein interaction network.Nat Commun. 2015;6:7412. [27] Zhang C,Shen L.Functional modules analysis based on protein-protein network analysis in ankylosing spondylitis.Eur Rev Med Pharmacol Sci. 2012;16(13):1821-1827.[28] Zhao Y, Mooney SD. Functional organization and its implication in evolution of the human protein-protein interaction network.BMC Genomics. 2012;13:150.[29] Schuster S, Fell DA, Dandekar T. A general definition of metabolic pathways useful for systematic organization and analysis of complex metabolic networks. Nat. Biotechnol. 2000;18:326-332.[30] Edwards JS, Ibarra RU, Palsson BO. In silico predictions of Escherichia coli metabolic capabilities are consistent with experimental data. Nat.Biotechnol. 2001;19;125-130.[31] Jeong H, Mason SP, Barabasi AL,et al. Lethality and centrality in protein networks. Nature.2001;411:41-42.[32] Lee DS, Park J, Kay KA,et al. The implications of human metabolic network topology for disease comorbidity. Proc. Natl. Acad. Sci. USA.2008;105:9880-9885.[33] Kanehisa M, Araki M, Goto S,et al. KEGG for linking genomes to life and the environment. Nucleic Acids Res.2008;36: D480-D484.[34] Oberhardt MA, Palsson BO, Papin JA. Applications of genome-scale metabolic reconstructions. Mol Syst Biol.2009; 5:320.[35] Duarte NC, Becker SA, Jamshidi N, et al.Global reconstruction of the human metabolic network based on genomic and bibliomic data. Proc. Natl. Acad. Sci. USA.2007; 104:1777-1782. [36] Karlstädt A, Fliegner D, Kararigas G, et al. CardioNet: a human metabolic network suited for the study of cardiomyocyte metabolism. BMC Syst Biol.2012;6:114. [37] Sahoo S,Franzson L,Jonsson JJ,et al. A compendium of inborn errors of metabolism mapped onto the human metabolic network. Mol Biosyst. 2012;8(10):2545-2558.[38] Mo ML,Palsson BO.Understanding human metabolic physiology: a genome-to-systems approach. Trends Biotechnol. 2009;27:37-44.[39] Kell DB, Goodacre R. Metabolomics and systems pharmacology: why and how to model the human metabolic network for drug discovery. Drug Discov Today.2014;19(2):171-182. [40] Galhardo M,Sinkkonen L,Berninger P,et al.Integrated analysis of transcript-level regulation of metabolism reveals disease-relevant nodes of the human metabolic network. Nucleic Acids Res.2014;42(3):1474-1496.[41] Hu J,Locasale JW,Bielas JH,et al.Heterogeneity of tumor-induced gene expression changes in the human metabolic network. Nat Biotechnol.2013;31(6):522-529[42] Wu M, Chan C.Prediction of therapeutic microRNA based on the human metabolic network.Bioinformatics. 2014;30(8): 1163-1171.[43] Megchelenbrink W, Katzir R, Lu X.Synthetic dosage lethality in the human metabolic network is highly predictive of tumor growth and cancer patient survival.Proc Natl Acad Sci USA. 2015;112(39):12217-12222.[44] Gebauer J, Schuster S, de Figueiredo LF, et al. Detecting and investigating substrate cycles in a genome-scale human metabolic network.FEBS J.2012;279(17):3192-3202[45] Bordbar A, Palsson BO.Using the reconstructed genome-scale human metabolic network to study physiology and pathology. J Intern Med. 2012;271(2):131-141.[46] Zhu C,Byers KJ,McCord RP,et al.High-resolution DNA-binding specificity analysis of yeast transcription factors. Genome Res.2009;19:556–566.[47] Deplancke B, Dupuy D, Vidal M, et al. A Gatewaycompatible yeast one-hybrid system. Genome Res. 2004;14, 2093–2101.[48] Lee TI, Rinaldi NJ, Robert F,et al. Transcriptional regulatory networks inSaccharomyces cerevisiae. Science.2002;298, 799-804.[49] Walhout AJ. Unraveling transcription regulatory networks by proteinDNA and protein-protein interaction mapping. Genome Res. 2006;16:1445-1454.[50] Reece-Hoyes JS, Deplancke B, Shingles J,et al. A compendium ofCaenorhabditis elegansregulatory transcription factors: a resource for mapping transcription regulatory networks. Genome Biol. 2005;6:R110.[51] Vaquerizas JM, Kummerfeld SK, Teichmann SA, et al. A census of human transcription factors: function, expression and evolution. Nat. Rev. Genet. 2009; 10:252-263.[52] Grove CA, De Masi F, Barrasa MI, et al. A multiparameter network reveals extensive divergence betweenC. elegansbHLH transcription factors. Cell.2009;138: 314-327.[53] Cawley S, Bekiranov S, Ng HH, et al. Unbiased mapping of transcription factor binding sites along human chromosomes 21 and 22 points to widespread regulation of noncoding RNAs. Cell.2004;116:499-509.[54] Gustafsson M, Gawel DR, Alfredsson L,et al.A validated gene regulatory network and GWAS identifies early regulators of T cell-associated diseases. Sci Transl Med. 2015;7(313):313ra178.[55] He C, Gao H, Fan X, et al. Identification of a novel miRNA-target gene regulatory network in osteosarcoma by integrating transcriptome analysis. Int J Clin Exp Pathol. 2015; 8(7):8348-8357.[56] Carson MB, Gu J, Yu G, et al. Identification of cancer-related genes and motifs in the human gene regulatory network. IET Syst Biol. 2015;9(4):128-134.[57] Mohammadnia A, Yaqubi M, Fallahi H. Predicting transcription factors in human alcoholic hepatitis from gene regulatory network. Eur Rev Med Pharmacol Sci. 2015;19(12):2246-2253.[58] Kousa YA, Schutte BC.Toward an orofacial gene regulatory network. Dev Dyn. 2016;245(3):220-232.[59] Tang WW, Dietmann S, Irie N,et al. A Unique Gene Regulatory Network Resets the Human Germline Epigenome for Development. Cell. 2015;161(6):1453-1467.[60] Yang X, Jia M, Li Z, et al. Bioinformatics analysis of aggressive behavior of breast cancer via an integrated gene regulatory network.J Cancer Res Ther.2014;10(4):1013-1018.[61] Xu X, Li H.Integrated microRNA gene analysis of coronary artery disease based on miRNA and gene expression profiles.Mol Med Rep.2016;13(4):3063-3073.[62] Ruvkun G, Wightman B,Ha I. The 20 years it took to recognize the importance of tiny RNAs. Cell.2004;116;S93-S96.[63] Martinez NJ, Ow MC, Barrasa MI,et al. AC. elegans genome-scale microRNA network contains composite feedback motifs with high flux capacity. Genes Dev. 2008;22: 2535-2549.[64] Zhang XD, Song J, Bork P, et al.The exploration of network motifs as potential drug targets from post-translational regulatory networks.Scientific Reports.2016;6:20558.[65] Zamani-Ahmadmahmudi M, Najafi A, Nassiri SM, et al. Reconstruction of canine diffuse large B-cell lymphoma gene regulatory network: detection of functional modules and hub genes. J Comp Pathol. 2015;152(2-3):119-130.[66] Gubelmann C, Schwalie PC, Raghav SK, et al. Identification of the transcription factor ZEB1 as a central component of the adipogenic gene regulatory network. Elife. 2014;3:e03346.[67] Janky R, Verfaillie A, Imrichová H, et al. iRegulon: from a gene list to a gene regulatory network using large motif and track collections. PLoS Comput Biol. 2014;10(7):e1003731.[68] Dusonchet J, Li H, Guillily M, et al. A Parkinson's disease gene regulatory network identifies the signaling protein RGS2 as a modulator of LRRK2 activity and neuronal toxicity. Hum Mol Genet.2014;23(18):4887-4905.[69] Ravasz E,Somera AL,Mongru DA,et al. Hierarchical organization of modularity in metabolic networks. Science. 2002;297:1551-1555.[70] Menche J, Sharma A, Kitsak M, et al.Uncovering Disease-disease Relationships Through the Incomplete Interactome. Science.2015;347, 1257601.[71] 冀俊忠,刘志军,刘红欣,等. 蛋白质相互作用网络功能模块检测的研究综述[J].自动化学报, 2014, 40(4):577-593.[72] Kemmeren P, van Berkum NL, Vilo J, et al. Protein interaction verification and functional annotation by integrated analysis of genome-scale data. Mol. Cell.2002; 9:1133-1143.[73] Stuart JM, Segal E, Koller D,et al. A gene-coexpression network for global discovery of conserved genetic modules. Science.2003;302:249-255.[74] Ge H, Walhout AJ, Vidal M. Integrating ‘omic’ information: a bridge between genomics and systems biology. Trends Genet. 2003;19: 551-560.[75] Boulton SJ, Gartner A, Reboul J, et al. Combined functional genomic maps of theC. elegansDNA damage response. Science. 2002;295:127-131.[76] Mani R, St Onge RP, Hartman JL,et al. Defining genetic interaction. Proc. Natl. Acad. Sci. USA.2008;105: 3461-3466.[77] Boone C, Bussey H, Andrews BJ. Exploring genetic interactions and networks with yeast. Nat. Rev. Genet. 2007; 8:437-449.[78] Rai S, Bhatnagar S. Hyperlipidemia, Disease Associations, and Top 10 Potential Drug Targets: A Network View. OMICS. 2016; 20(3):152-168.[79] Network medicine. From Wikipedia, the free encyclopedia. https://en.wikipedia.org/wiki/Network_medicine#Diseasome.[80] Goh KI, Cusick ME, Valle D, et al. The human disease network. Proc Natl Acad Sci U S A. 2007;104(21):8685-8690.[81] Lee DS, Park J, Kay KA, et al. The implications of human metabolic network topology for disease comorbidity. PNAS. 2008; 105:9880-9885.[82] Lu M, Zhang Q, Deng M, et al. An Analysis of Human MicroRNA and Disease Associations. PLoS One. 2008;3(10): e3420. [83] Yuan D, Cui X, Wang Y, et al. Enrichment Analysis Identifies Functional MicroRNA-Disease Associations in Humans. PLoS One.2015;10(8): e0136285. [84] Rzhetsky A, et al. Probing genetic overlap among complex human phenotypes. PNAS. 2007;104:11694-11699.[85] Hidalgo C, Blumm N, Barabási AL, et al.A Dynamic Network Approach for the Study of Human Phenotypes. Plos Computational Biology. 2009; 5 e1000353.[86] van Driel MA, Bruggeman J, Vriend G, et al. A text-mining analysis of the human phenome. Eur J Hum Genet. 2006; 14(5):535-542.[87] Zhou X, Menche J, Barabási AL, et al.Human symptoms-disease network.Nat. Commun. 2014;5:4212.[88] Liu YI, Wise PH, Butte AJ. The "etiome": identification and clustering of human disease etiological factors. BMC Bioinformatics. 2009; 10:S14.[89] Yang J, Wu SJ, Dai WT, et al. The human disease network in terms of dysfunctional regulatory mechanisms. Biol Direct. 2015;10:60.[90] Yang J, Wu SJ, Yang SY, et al.DNetDB: The human disease network database based on dysfunctional regulation mechanism. BMC Syst Biol. 2016;10(1):36.[91] Goh KI,Choi IG. Exploring the human diseasome: the human disease network. Brief Funct Genomics. 2012;11(6):533-542.[92] Yildirim MA, Goh KI, Cusick ME, et al. Drug-target network. Nat Biotechnol.2007; 25:1119-1126. |
| [1] | 姚晓玲, 彭建城, 许岳荣, 杨志东, 张顺聪. 可变角度零切迹前路椎间融合内固定系统治疗脊髓型颈椎病:30个月随访[J]. 中国组织工程研究, 2022, 26(9): 1377-1382. |
| [2] | 张璟琳, 冷 敏, 朱博恒, 汪 虹. 干细胞源外泌体促进糖尿病创面愈合的机制及应用[J]. 中国组织工程研究, 2022, 26(7): 1113-1118. |
| [3] | 安维政, 何 萧, 任 帅, 刘建宇. 肌源干细胞在周围神经再生中的潜力[J]. 中国组织工程研究, 2022, 26(7): 1130-1136. |
| [4] | 何云影, 李玲婕, 张舒淇, 李雨舟, 杨 生, 季 平. 聚丙烯酸/琼脂糖三维培养构建细胞球的方法[J]. 中国组织工程研究, 2022, 26(4): 553-559. |
| [5] | 何冠宇, 徐宝山, 杜立龙, 张同星, 霍振鑫, 申 力. 天然丝素蛋白构建仿生取向微通道纤维环支架[J]. 中国组织工程研究, 2022, 26(4): 560-566. |
| [6] | 陈晓旭, 罗雅馨, 毕浩然, 杨 琨. 脱细胞支架制备及其在组织工程和再生医学中的应用[J]. 中国组织工程研究, 2022, 26(4): 591-596. |
| [7] | 康坤龙, 王新涛. 生物支架材料促进骨髓间充质干细胞成骨分化的研究热点[J]. 中国组织工程研究, 2022, 26(4): 597-603. |
| [8] | 沈佳华, 付 勇. 基于石墨烯的纳米材料可否在干细胞领域应用[J]. 中国组织工程研究, 2022, 26(4): 604-609. |
| [9] | 张 通, 蔡金池, 袁志发, 赵海燕, 韩兴文, 王文己. 基于透明质酸的复合水凝胶修复骨关节炎软骨损伤:应用与机制[J]. 中国组织工程研究, 2022, 26(4): 617-625. |
| [10] | 李 辉, 陈良龙. 植骨材料在脊柱结核治疗中的应用与特征[J]. 中国组织工程研究, 2022, 26(4): 626-630. |
| [11] | 高仓健, 杨 振, 刘舒云, 李 浩, 付力伟, 赵天元, 陈 威, 廖志垚, 李品学, 眭 翔, 郭全义. 静电纺丝技术在肩袖损伤修复中的应用[J]. 中国组织工程研究, 2022, 26(4): 637-642. |
| [12] | 关 健, 贾燕飞, 张葆鑫, 赵国中. 4D生物打印在组织工程的应用[J]. 中国组织工程研究, 2022, 26(3): 446-455. |
| [13] | 刘家利, 索海瑞, 杨翰, 王玲, 徐铭恩. 填充结构对3D打印聚己内酯支架力学性能的影响[J]. 中国组织工程研究, 2022, 10(16): 2612-2617. |
| [14] | 黄波, 陈明学, 彭礼庆, 罗旭江, 李获, 王皓, 田沁玉, 鲁晓波, 刘舒云, 郭全义. 可注射性甲基丙烯酸酐改性明胶/软骨源性基质微粒复合水凝胶支架的制备及生物相容性[J]. 中国组织工程研究, 2022, 10(16): 2600-2606. |
| [15] | 房晓阳, 唐田, 王楠, 钱宇章, 谢林. 椎间盘全层纤维环修复与再生治疗[J]. 中国组织工程研究, 2022, 26(10): 1582-1587. |
中国组织工程研究杂志出版内容重点:组织构建;骨细胞;软骨细胞;细胞培养;成纤维细胞;血管内皮细胞;骨质疏松;组织工程
1.4 质量评价 符合纳入标准的92篇文献中,文献[1-4]探讨了网络医学的概念和作用;文献[5-78]探讨了分子相互作用网络的研究现状,文献[79-90]探讨了疾病网络的研究现状,文献[91-92]探讨了药物网络的研究现状。文献检索流程图见图1。
.jpg)
中国组织工程研究杂志出版内容重点:组织构建;骨细胞;软骨细胞;细胞培养;成纤维细胞;血管内皮细胞;骨质疏松;组织工程
中国组织工程研究杂志出版内容重点:组织构建;骨细胞;软骨细胞;细胞培养;成纤维细胞;血管内皮细胞;骨质疏松;组织工程
.jpg) 文题释义:
网络医学:网络医学使用网络科学来诊断、预防和治疗疾病,它提供一个量化的平台来解决人类疾病的复杂性。与医学有关的网络以简单原则组织起来,这些组织原则包括度分布和枢纽、小世界现象、模式(motif)、模块、介数中心性。这些网络的节点代表生物因素(生物分子,疾病,表型等),连接(边)则代表它们的关系(物理相互作用,共享的代谢途径,共享基因,共享性状等)。
Interactome:又称为分子相互作用网络。在这类网络中,节点是代谢物和诸如蛋白质、RNA分子和基因序列这样的分子,而边是可以用大量技术来识别的物理、生物化学和功能相互作用。在Interactome中,代谢网络、蛋白质-蛋白质相互作用网络和基因调控网络提供了关于细胞系统的关键的“骨架”信息,在这个“骨架”上还可以添加功能连接的额外层(转录谱网络、表型谱网络和基因相互作用网络)来调整生物的现实表现。
中国组织工程研究杂志出版内容重点:组织构建;骨细胞;软骨细胞;细胞培养;成纤维细胞;血管内皮细胞;骨质疏松;组织工程
文题释义:
网络医学:网络医学使用网络科学来诊断、预防和治疗疾病,它提供一个量化的平台来解决人类疾病的复杂性。与医学有关的网络以简单原则组织起来,这些组织原则包括度分布和枢纽、小世界现象、模式(motif)、模块、介数中心性。这些网络的节点代表生物因素(生物分子,疾病,表型等),连接(边)则代表它们的关系(物理相互作用,共享的代谢途径,共享基因,共享性状等)。
Interactome:又称为分子相互作用网络。在这类网络中,节点是代谢物和诸如蛋白质、RNA分子和基因序列这样的分子,而边是可以用大量技术来识别的物理、生物化学和功能相互作用。在Interactome中,代谢网络、蛋白质-蛋白质相互作用网络和基因调控网络提供了关于细胞系统的关键的“骨架”信息,在这个“骨架”上还可以添加功能连接的额外层(转录谱网络、表型谱网络和基因相互作用网络)来调整生物的现实表现。
中国组织工程研究杂志出版内容重点:组织构建;骨细胞;软骨细胞;细胞培养;成纤维细胞;血管内皮细胞;骨质疏松;组织工程
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||