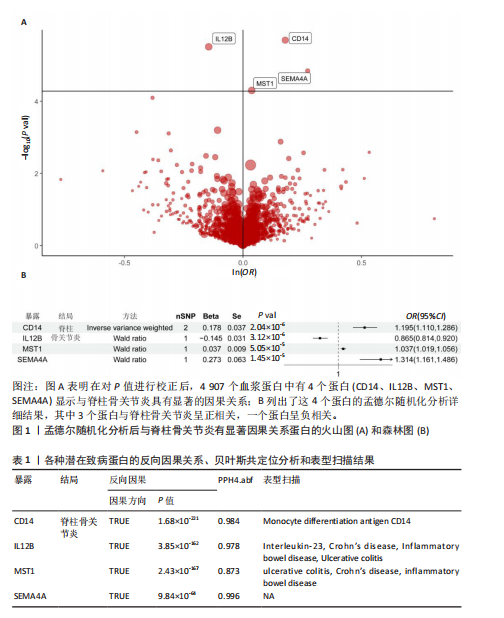
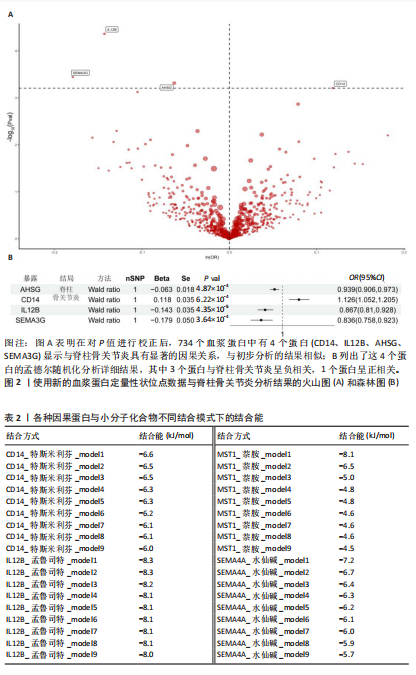
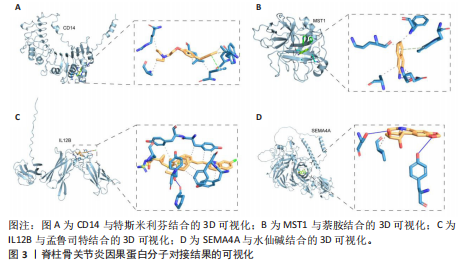
[1] FELSON DT, LAWRENCE RC, DIEPPE PA, et al. Osteoarthritis: new insights. Part 1: the disease and its risk factors. Ann Intern Med. 2000;133(8): 635-646.
[2] CHO HJ, MOREY V, KANG JY, et al. Prevalence and Risk Factors of Spine, Shoulder, Hand, Hip, and Knee Osteoarthritis in Community-dwelling Koreans Older Than Age 65 Years. Clin Orthop Relat Res. 2015;473(10):3307-3314.
[3] IORDACHE S, CURSARU A, MARINESCU A, et al. Magnetic Resonance Imaging Features and Functional Score in Patients Requiring Total Knee Arthroplasty. Cureus. 2024;16(9):e68595.
[4] LUO H, LAI Y, TANG W, et al. Mitochondrial transplantation: a promising strategy for treating degenerative joint diseases. J Transl Med. 2024; 22(1):941.
[5] TANG L, DING J, YANG K, et al. New insights into the mechanisms and therapeutic strategies of chondrocyte autophagy in osteoarthritis. J Mol Med (Berl). 2024;102(10):1229-1244.
[6] CIOROIANU GO, FLORESCU A, FLORESCU LM, et al. Knee Osteoarthritis-Current Diagnosis and Treatment Options-A Narrative Review. Curr Health Sci J. 2024;50(2):163-169.
[7] EMILSSON V, GUDMUNDSDOTTIR V, GUDJONSSON A, et al. Coding and regulatory variants are associated with serum protein levels and disease. Nat Commun. 2022;13(1):481.
[8] GUDJONSSON A, GUDMUNDSDOTTIR V, AXELSSON GT, et al. A genome-wide association study of serum proteins reveals shared loci with common diseases. Nat Commun. 2022;13(1):480.
[9] LIU F, YANG H, YANG T, et al. Dysregulated proteasome activity and steroid hormone biosynthesis are associated with mortality among patients with acute COVID-19. J Transl Med. 2024; 22(1):626.
[10] MAHIN A, SOMAN SP, MODI PK, et al. Meta-analysis of the serum/plasma proteome identifies significant associations between COVID-19 with Alzheimer’s/Parkinson’s diseases. J Neurovirol. 2024;30(1):57-70.
[11] GU Y, JIN Q, HU J, et al. Causality of genetically determined metabolites and metabolic pathways on osteoarthritis: a two-sample mendelian randomization study. J Transl Med. 2023;21(1): 357.
[12] TERCIC D, BOZIC B. The basis of the synovial fluid analysis. Clin Chem Lab Med. 2001;39(12):1221-1226.
[13] PIERCE BL, BURGESS S. Efficient design for Mendelian randomization studies: subsample and 2-sample instrumental variable estimators. Am J Epidemiol. 2013;178(7):1177-1184.
[14] HARTWIG FP, DAVIES NM, HEMANI G, et al. Two-sample Mendelian randomization: avoiding the downsides of a powerful, widely applicable but potentially fallible technique. Int J Epidemiol. 2016;45(6):1717-1726.
[15] LIN J, ZHOU J, XU Y. Potential drug targets for multiple sclerosis identified through Mendelian randomization analysis. Brain. 2023;146(8): 3364-3372.
[16] REAY WR, CAIRNS MJ. Advancing the use of genome-wide association studies for drug repurposing. Nat Rev Genet. 2021;22(10):658-671.
[17] EMDIN CA, KHERA AV, KATHIRESAN S. Mendelian Randomization. JAMA. 2017;318(19):1925-1926.
[18] FERKINGSTAD E, SULEM P, ATLASON BA, et al. Large-scale integration of the plasma proteome with genetics and disease. Nat Genet. 2021; 53(12):1712-1721.
[19] ZHENG J, HABERLAND V, BAIRD D, et al. Phenome-wide Mendelian randomization mapping the influence of the plasma proteome on complex diseases. Nat Genet. 2020;52(10):1122-1131.
[20] BOER CG, HATZIKOTOULAS K, SOUTHAM L, et al. Deciphering osteoarthritis genetics across 826,690 individuals from 9 populations. Cell. 2021; 184(24):6003-6005.
[21] ZHANG J, DUTTA D, KÖTTGEN A, et al. Plasma proteome analyses in individuals of European and African ancestry identify cis-pQTLs and models for proteome-wide association studies. Nat Genet. 2022;54(5):593-602.
[22] WANG Q, DAI H, HOU T, et al. Dissecting Causal Relationships Between Gut Microbiota, Blood Metabolites, and Stroke: A Mendelian Randomization Study. J Stroke. 2023;25(3):350-360.
[23] YAN W, JIANG M, HU W, et al. Causality Investigation between Gut Microbiota, Derived Metabolites, and Obstructive Sleep Apnea: A Bidirectional Mendelian Randomization Study. Nutrients. 2023;15(21):4544.
[24] BOWDEN J, DEL GRECO MF, MINELLI C, et al. Assessing the suitability of summary data for two-sample Mendelian randomization analyses using MR-Egger regression: the role of the I2 statistic. Int J Epidemiol. 2016;45(6):1961-1974.
[25] CHENG J, YANG L, YE Y, et al. Mendelian Randomisation Analysis of Causal Association between Lifestyle, Health Factors, and Keratoconus. Bioengineering (Basel). 2024; 11(3):221.
[26] BURGESS S, BUTTERWORTH A, THOMPSON SG. Mendelian randomization analysis with multiple genetic variants using summarized data. Genet Epidemiol. 2013;37(7):658-665.
[27] HEMANI G, TILLING K, DAVEY SMITH G. Orienting the causal relationship between imprecisely measured traits using GWAS summary data. PLoS Genet. 2017;13(11):e1007081.
[28] CULLELL N, GALLEGO-FÁBREGA C, CÁRCEL-MÁRQUEZ J, et al. ICA1L Is Associated with Small Vessel Disease: A Proteome-Wide Association Study in Small Vessel Stroke and Intracerebral Haemorrhage. Int J Mol Sci. 2022;23(6):3161.
[29] GIAMBARTOLOMEI C, VUKCEVIC D, SCHADT EE, et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 2014;10(5):e1004383.
[30] KAMAT MA, BLACKSHAW JA, YOUNG R, et al. PhenoScanner V2: an expanded tool for searching human genotype-phenotype associations. Bioinformatics. 2019;35(22):4851-4853.
[31] BURLEY SK, BHIKADIYA C, BI C, et al. RCSB Protein Data Bank: powerful new tools for exploring 3D structures of biological macromolecules for basic and applied research and education in fundamental biology, biomedicine, biotechnology, bioengineering and energy sciences. Nucleic Acids Res. 2021;49(D1):D437-d451.
[32] MONTGOMERY SB, DERMITZAKIS ET. From expression QTLs to personalized transcriptomics. Nat Rev Genet. 2011;12(4):277-282.
[33] OPPMANN B, LESLEY R, BLOM B, et al. Novel p19 protein engages IL-12p40 to form a cytokine, IL-23, with biological activities similar as well as distinct from IL-12. Immunity. 2000;13(5):715-725.
[34] TENG MW, BOWMAN EP, MCELWEE JJ, et al. IL-12 and IL-23 cytokines: from discovery to targeted therapies for immune-mediated inflammatory diseases. Nat Med. 2015;21(7):719-729.
[35] EK WE, KARLSSON T, HÖGLUND J, et al. Causal effects of inflammatory protein biomarkers on inflammatory diseases. Sci Adv. 2021;7(50): eabl4359.
[36] WRIGHT SD, RAMOS RA, TOBIAS PS, et al. CD14, a receptor for complexes of lipopolysaccharide (LPS) and LPS binding protein. Science. 1990;249(4975): 1431-1433.
[37] SETOGUCHI M, NASU N, YOSHIDA S, et al. Mouse and human CD14 (myeloid cell-specific leucine-rich glycoprotein) primary structure deduced from cDNA clones. Biochim Biophys Acta. 1989; 1008(2):213-222.
[38] ZANONI I, TAN Y, DI GIOIA M, et al. By Capturing Inflammatory Lipids Released from Dying Cells, the Receptor CD14 Induces Inflammasome-Dependent Phagocyte Hyperactivation. Immunity. 2017;47(4):697-709.e693.
[39] FURNKRANZ A, SCHOBER A, BOCHKOV VN, et al. Oxidized phospholipids trigger atherogenic inflammation in murine arteries. Arterioscler Thromb Vasc Biol. 2005;25(3):633-638.
[40] SHARYGIN D, KONIARIS LG, WELLS C, et al. Role of CD14 in human disease. Immunology. 2023;169(3):260-270.
[41] FUENTELSAZ-ROMERO S, BARRIO-ALONSO C, GARCÍA CAMPOS R, et al. The Macrophage Reprogramming Ability of Antifolates Reveals Soluble CD14 as a Potential Biomarker for Methotrexate Response in Rheumatoid Arthritis. Front Immunol. 2021;12:776879.
[42] NAKAMIZO S, DUTERTRE CA, KHALILNEZHAD A, et al. Single-cell analysis of human skin identifies CD14+ type 3 dendritic cells co-producing IL1B and IL23A in psoriasis. J Exp Med. 2021;218(9):e20202345.
[43] WONG SS, OSHANSKY CM, GUO XJ, et al. Activated CD4(+) T cells and CD14(hi)CD16(+) monocytes correlate with antibody response following influenza virus infection in humans. Cell Rep Med. 2021;2(4):100237.
[44] SHI Z, ZHOU Z. MST kinases in innate immune signaling. Cell Stress. 2017;2(1):4-13.
[45] PRASKOVA M, XIA F, AVRUCH J. MOBKL1A/MOBKL1B phosphorylation by MST1 and MST2 inhibits cell proliferation. Curr Biol. 2008;18(5): 311-321.
[46] ABUKAR Y, RAMCHANDRA R, HOOD SG, et al. Increased cardiac sympathetic nerve activity in ovine heart failure is reduced by lesion of the area postrema, but not lamina terminalis. Basic Res Cardiol. 2018;113(5):35.
[47] LI T, WEN Y, LU Q, et al. MST1/2 in inflammation and immunity. Cell Adh Migr. 2023;17(1):1-15.
[48] ZONG F, ZHAO Y. Alkaloid leonurine exerts anti-inflammatory effects via modulating MST1 expression in trophoblast cells. Immun Inflamm Dis. 2021;9(4):1439-1446.
[49] ZHOU C, YAO S, FU F, et al. Morroniside attenuates nucleus pulposus cell senescence to alleviate intervertebral disc degeneration via inhibiting ROS-Hippo-p53 pathway. Front Pharmacol. 2022; 13:942435.
[50] LIU C, CHU X, BIAO Y, et al. Association between lipid-lowering agents with intervertebral disc degeneration, sciatica and low back pain: a drug-targeted mendelian randomized study and cross-sectional observation. Lipids Health Dis. 2024; 23(1):327.
[51] QUAN M, LV H, LIU Z, et al. MST1 Suppresses Disturbed Flow Induced Atherosclerosis. Circ Res. 2022;131(9):748-764.
[52] IYER AS, CHAPOVAL SP. Neuroimmune Semaphorin 4A in Cancer Angiogenesis and Inflammation: A Promoter or a Suppressor? Int J Mol Sci. 2018; 20(1):124.
[53] NKYIMBENG-TAKWI E, CHAPOVAL SP. Biology and function of neuroimmune semaphorins 4A and 4D. Immunol Res. 2011;50(1):10-21.
[54] KUMANOGOH A, MARUKAWA S, SUZUKI K, et al. Class IV semaphorin Sema4A enhances T-cell activation and interacts with Tim-2. Nature. 2002; 419(6907):629-633.
[55] WANG L, SONG G, ZHENG Y, et al. Expression of Semaphorin 4A and its potential role in rheumatoid arthritis. Arthritis Res Ther. 2015; 17(1):227.
[56] KUMANOGOH A, KIKUTANI H. Immune semaphorins: a new area of semaphorin research. J Cell Sci. 2003;116(Pt 17):3463-3470
|