Chinese Journal of Tissue Engineering Research ›› 2023, Vol. 27 ›› Issue (10): 1584-1592.doi: 10.12307/2023.313
Previous Articles Next Articles
Progress in studies on biological properties of stem cells modified by mRNA m6A methylation
Han Weiyu, Zhao Yongchao, Zhao Ranzun
- Department of Cardiology, Affiliated Hospital of Zunyi Medical University, Zunyi 563000, Guizhou Province, China
-
Received:2022-01-25Accepted:2022-06-06Online:2023-04-08Published:2022-09-09 -
Contact:Zhao Ranzun, MD, Chief physician, Professor, Doctoral supervisor, Department of Cardiology, Affiliated Hospital of Zunyi Medical University, Zunyi 563000, Guizhou Province, China -
About author:Han Weiyu, Master candidate, Department of Cardiology, Affiliated Hospital of Zunyi Medical University, Zunyi 563000, Guizhou Province, China -
Supported by:the National Natural Science Foundation of China, No. 81960066 (to ZRZ); the Basic Research Program of the Department of Science and Technology of Guizhou Province, No. ZK[2022] 671 (to ZYC)
CLC Number:
Cite this article
Han Weiyu, Zhao Yongchao, Zhao Ranzun. Progress in studies on biological properties of stem cells modified by mRNA m6A methylation[J]. Chinese Journal of Tissue Engineering Research, 2023, 27(10): 1584-1592.
share this article
Add to citation manager EndNote|Reference Manager|ProCite|BibTeX|RefWorks
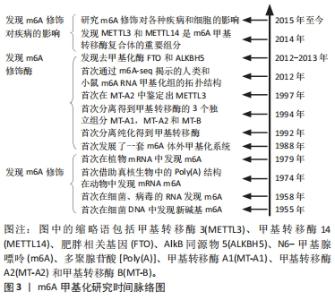
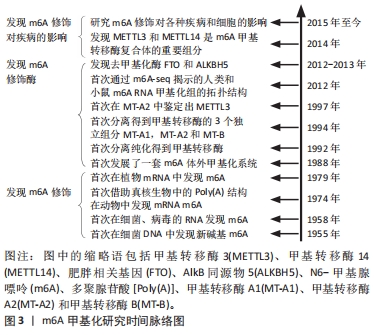
2.1 m6A甲基化调控概述 m6A是核糖核酸第6位氮原子上发生甲基化的过程,被甲基阅读蛋白识别,并在转录水平调控相关分子的表达。1955年,DUNN等[4]首次在细菌DNA中发现新碱基m6A,随后DESROSIERS等[5]又在真核生物的mRNA中发现m6A,开辟了研究m6A甲基化修饰的新方向。此后随着研究的深入,在各种原核生物、真核生物以及病毒中都陆续发现m6A甲基化修饰的存在,以及后续基因测序技术的发展,m6A甲基化修饰的相关酶分子及其功能、结构、定位以及作用等陆续被发现,逐步完善了m6A体外甲基化修饰体系。随着表观遗传学的发展以及研究的深入,多种疾病模型中也证实了m6A修饰对各种疾病的影响及其作用机制,突出了m6A对于人类疾病的治疗前景。随后的研究证明了m6A甲基化修饰在各种疾病和细胞中的重要性,见图3。"

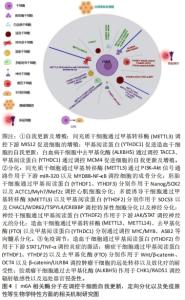
m6A是指RNA腺嘌呤第6号位N原子上的可逆性的甲基化修饰类型,是真核生物mRNA中最常见的转录后修饰,可影响修饰RNA的转录后命运,在多种生物学过程中发挥重要作用。mRNA是生物体蛋白质合成的直接模板,在基因信息表达传递中发挥重要的作用。m6A修饰位点主要富集在相对保守的RRACH序列(R代表A/G、H代表A/C/U)[6],该序列最常出现在mRNA分子的3′非翻译区(3′-UTR)、终止密码子以及长的外显子区域[7]。动态m6A修饰对mRNA的影响依赖于特定的3类酶分子,即“Writer,Eraser,Reader”,即甲基转移酶、去甲基化酶和甲基化阅读蛋白。甲基转移酶包括甲基转移酶3(methyltransferase like 3,METTL3)、甲基转移酶14(methyltransferase like 14,METTL14)、甲基转移酶16(methyltransferase like 16,METTL16)、Wilms’肿瘤相关蛋白(Wilms’ tumor associating protein,WTAP)、Vir同源m6A甲基转移酶样物(vir like m6A methyltransferase associated,VIRMA)、RNA结合基序蛋白15(RNA binding motif protein 15,RBM15)、RNA结合基序蛋白15旁系同源蛋白(RNA binding motif protein 15 paralogue,RBM15B)以及锌指CCCH域含蛋白13(zinc finger CCCH-type containing,ZC3H13)等[8-13]。去甲基化酶包括肥胖相关基因(fat mass and obesity associated gene,FTO)和AlkB同源物5(AlkB homolog 5,ALKBH5)[14-15]。甲基化阅读蛋白是指一类能识别并结合m6A修饰的mRNA进而调RNA转录后代谢的蛋白,包括YTH结构域蛋白家族(YT521-B homology,YTH)和胰岛素样生长因子结合蛋白家族(insulin-like growth factor mRNA-binding proteins,IGFBPs)以及异质核糖核蛋白等[16-17]。其中,YTH结构域蛋白家族有5个成员:YTHDC1,YTHDC2,YTHDF1,YTHDF2和YTHDF3[16]。IGFBPs家族中的IGFBP-1、-2和-3已被确定为独特的m6A阅读蛋白,含有共同的RNA结合域,可识别m6A mRNA的UGGAC序列并募集RNA稳定剂以抑制m6A修饰的mRNA的降解,以及异质核糖核蛋白A2B1(heterogeneous nuclear ribonucleoprotein A2B1,HNRNPA2B1)[18]、异质核糖核蛋白C(heterogeneous nuclear ribonucleoprotein C,HNRNPC)及异质核糖核蛋白G(heterogeneous nuclear ribonucleoprotein G,HNRNPG)等在核内外共同参与mRNA的加工调节,进而调控多种生物学过程。 一般地,m6A的形成过程由S-腺苷甲硫氨酸负责提供甲基,在甲基转移酶催化下促进m6A的形成,后者可同时接受去甲基化酶的选择性清除m6A,之后由甲基结合蛋白特异性识别m6A从而调控RNA的转录后代谢过程,如mRNA的选择性剪切、出核、翻译、稳定性和降解等过程[19-28],进而发挥调节干细胞自我更新、定向分化与免疫调节等在内的多种生物学特性[29-48],见表1。因此,m6A甲基化修饰在维持干细胞生物特性方面具有重要学意义,深入研究m6A甲基化将极大地促进对干细胞介导的再生医学的认识与发展[49-50] 。 "


2.2 m6A在干细胞生物学特性方面的调控作用 2.2.1 m6A调控干细胞增殖与自我更新 干细胞的增殖与自我更新是维持细胞群体稳定性的重要特性之一,正常干细胞自我更新与增殖受程序性控制,增殖和分化维持平衡有序状态,干细胞分为胚胎干细胞与成体干细胞,成体干细胞中典型代表是肿瘤干细胞,肿瘤干细胞的失控无序的自我更新与增殖是恶性肿瘤重要机制之一,而非肿瘤性干细胞在恶性疾病的发生发展的恶劣微环境中不能耐受导致增殖受抑制,影响疾病治疗及预后。先前研究已阐明动态m6A修饰可广泛地参与多种干细胞的产生和维持,控制疾病的进展和治疗的有效性[51] ,但尚未阐明具体作用机制。 m6A甲基化修饰与多种干细胞的增殖与自我更新关系密切,可通过调控增殖相关蛋白、转录因子以及DNA复制过程中解旋酶复合体的表达等促进细胞的自我更新与增殖。CUI等[52]研究发现,METTL3,METTL14和FTO可通过动态调控胶质瘤干细胞的m6A甲基化在GSCs的增殖中发挥关键作用,敲除METTL3或METTL14将改变胶质瘤干细胞解整合素样金属蛋白酶19(A Disintegrin And Metalloprotease 19,ADAM19)和富含丝氨酸和精氨酸的剪切因子(serine- and arginine-rich splicing factors,SRSF)的表达[53],进而促进干细胞的自我更新。同样,高表达的ALKBH5可促进叉头框转录因子(forkhead box transcription factor M1,FOXM1)的未成熟mRNA去甲基化,进而上调其蛋白表达以维持胶质瘤干细胞的自我更新和成瘤能力[54]。 然而,抑制FTO,ALKBH5或过表达METTL3则可发挥抑制胶质瘤干细胞的功能,机制上与促进胶质瘤干细胞分化的关键转录因子SRY-like HMG box(SOX2)的甲基化并降低其表达相关。WU等[55]研究发现,早衰综合征患者的间充质干细胞中METTL3和m6A甲基化的水平均显著降低,过表达METTL3将通过IGF2BP2识别和促进m6A甲基化水平保持MIS12 mRNA稳定转录而抑制间充质干细胞的过早衰老,因此METTL3通过调控MIS12的m6A甲基化水平成为阻断人间充质干细胞衰老所必须的分子。SHENG等[56]最新研究发现,YTHDC1是人急性髓系白血病细胞增殖和生存所必需的,Ythdc1缺失将显著阻断小鼠体内急性髓系白血病的发生和白血病干细胞的自我复制与更新。YTHDC1对于正常骨髓造血功能和造血干/祖细胞维持自我更新至关重要,基因敲除Ythdc1将促进DNA复制过程中的解旋酶复合体的关键组分之一微小染色体维持蛋白(minichromosome maintenance proteins 4,MCM4)稳定性下降和表达减少,从而显著地抑制急性髓系白血病细胞增殖。SHEN等[57]近期研究表明,在急性髓系白血病中去甲基化酶ALKBH5是致癌基因诱导的细胞永生化和白血病发生所必需,可通过m6A转录后调节转录相关酸性卷曲样蛋白3(transforming acidic coiled-coil proteins,TACC3)的表达与多种肿瘤预后相关的关键致癌基因的表达,从而促进白血病干细胞的自我更新和急性髓系白血病的发生与进展,但有趣的是ALKBH5并不影响正常造血功能。 造血干细胞的自我更新和分化涉及大量mRNA的生成和降解,与其在正常和应激状态下细胞质中YTHDF2促进m6A依赖的mRNA降解相关[58]。BAI等[59]研究发现在大肠癌中YTHDF1的表达显著升高,介导大肠癌细胞中Wnt/β-catenin通路的激活并增强了肿瘤干细胞的增殖活性。综上,相关研究揭示了甲基转移酶、去甲基化酶和甲基阅读蛋白等通过影响m6A水平进而调控多种靶基因mRNA的甲基化修饰,表明m6A在干细胞的再生增殖与维持细胞干性中发挥重要作用。 因此,m6A甲基化修饰通过多种信号通路调控干细胞的自我更新与增殖。此外,m6A甲基化修饰酶可通过调控细胞周期转换、促进有丝分裂的过程、促进DNA复制过程中解旋酶复合体的表达以及促进转录后翻译等,从而促进干细胞的增殖等过程。现阶段关于m6甲基化修饰调控干细胞的研究主要集中在肿瘤干细胞、造血干细胞等种类的干细胞[52-75] ,还需以不同种类的干细胞为研究对象去进行更深层次的研究,见表2。 "

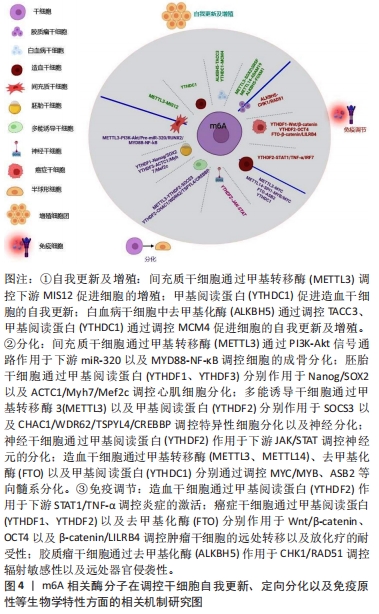
2.2.2 m6A调控干细胞的定向分化 干细胞的另一重要特征是具有多向或定向分化潜能,可产生不同类型的子代细胞,这是其发挥病理生理调控功能的最重要特征之一。近年研究发现,m6A甲基化修饰在调控干细胞的多向及与定向分化中亦扮演了重要的角色[76]。通过定向诱导分化,可弥补传统治疗带来的不足。 多项研究显示,间充质干细胞的成骨分化过程受到m6A的动态调节。TIAN等[30]研究显示骨髓源间充质干细胞中敲除METTL3将降低骨形成相关基因Runx2和Osterix的表达及碱性磷酸酶活性的降低,伴随间充质干细胞的成骨分化障碍以及骨矿化和矿化结节的形成减少。不仅如此,敲除METTLE3还会诱导间充质干细胞成骨分化过程中血管内皮生长因子(vascular endothelial growth factor,VEGF)的mRNA水平降低,表现为VEGF-164和VEGF-188两种mRNA剪接变体降低为主,而不影响VEGF-120水平。其中,VEGF-164主要负责调节间充质干细胞增殖和向软骨细胞分化,而VEGF-188则更有效地促进间充质干细胞的成骨分化。这些发现将有助于理解转录组的m6A动态调控在骨髓间充质干细胞成骨分化中的作用,为骨再生治疗提供新的可能策略。 YU等[60]研究从另一个机制阐述了m6A在骨髓源间充质干细胞成骨分化中的作用,发现METTL3通过增加m6A甲基化水平促进MYD88的表达,诱导核转录因子κB的激活并抑制骨形成。因此,METTL3是骨髓源间充质干细胞成骨分化进程中的一个关键抑制因子,以METTL3为靶点介导m6A的动态调节基因表达或是再生医学和骨组织工程领域中极有前景的策略。然而也有不同的研究结论,同期另一项研究却发现,骨质疏松症患者和去卵巢小鼠均出现METTL3的表达降低,导致骨髓间充质干细胞的成骨分化抑制,骨量减少,其作用机制可能是METTL3增强成骨分化和骨形成的关键转录基因RUNX2(runt-related transcription factor 2)和pre-miR-320的m6A甲基化水平实现的[61]。 WU等[62]研究也显示,间充质干细胞中METTL3过表达可保护小鼠免受雌激素缺乏引起的骨质疏松,相反地METTL3功能丧失将降低m6A调控的重要下游通路甲状旁腺激素(parathyroid hormone,PTH)/甲状旁腺激素受体1(parathyroid hormone receptor-1,Pth1r)信号轴中等位基因Pth1r的翻译效率,从而破坏PTH诱导的成骨和成脂反应,导致间充质干细胞的成骨分化潜能不足和骨形成受损,这些研究结果提示过表达METTL3可能是未来治疗人类骨质疏松症的一种新方法。因此,m6A在干细胞中成骨分化作用结果差异可能与研究靶基因不同有关,在不同干细胞中的确切作用与机制仍需深入研究。m6A可能也参与间充质干细胞向脂肪分化的调节。有研究显示,与健康者骨髓间充质干细胞相比,急性髓系白血病患者的骨髓间充质干细胞向脂肪增殖分化能力增强,深入分析发现可能与急性髓系白血病来源的间充质干细胞中METTL3表达下调后降低蛋白激酶mRNA的m6A甲基化水平,使蛋白激酶表达增加,诱导间充质干细胞的脂肪生成作用增强,这可能也是导致急性髓系白血病细胞化疗耐药性的重要原因之一[63]。因此m6A修饰靶向蛋白激酶调控间充质干细胞向脂肪分化可能为克服急性髓系白血病化疗耐药性提供一种全新的治疗思路与策略。 有研究发现原癌基因MYC是造血干细胞中m6A的直接靶点,在METTL3缺失的造血干细胞在诱导分化时不能上调MYC的表达,导致的造血干细胞的分化障碍,因此m6A调控MYC是造血干细胞对称性和非对称性分化的决定性标志[77]。此外,该靶点除了METTL3修饰外,作为甲基转移酶复合体的另一个关键组分也参与MYC的修饰调节, WENG等[64]研究发现,m6A甲基转移酶复合物的关键成分METTL14在正常造血干/祖细胞 (HSPCs)和携带t(11q23)、t(15;17) 或t(8;21)基因突变的急性髓系白血病细胞中高表达而在髓细胞分化过程中却相反出现表达下调。沉默METTL14将促进正常HSPCs和急性髓系白血病细胞的终末髓系分化并抑制急性髓系白血病细胞的存活/增殖,进一步研究揭示其机制可能是METTL14通过m6A修饰调节MYB和MYC(两者均为正常造血干细胞和急性髓系白血病自我更新和分化的主要调节因子)的mRNA靶点发挥其致癌作用。这些结果揭示了骨髓生成和白血病发生中的METTL14- MYB/ MYC信号轴的重要作用,提示METTL14调节的m6A修饰在骨髓正常造血和恶性增殖中的关键作用。 除甲基转移酶外,去甲基化酶也通过影响m6A水平参与干细胞定向分化功能调节。研究发现FTO高表达于急性髓系白血病中,其增强白血病癌基因介导的细胞分化和白血病发生,并抑制全反式维甲酸诱导的急性髓系白血病细胞分化。因此,FTO可能在急性髓系白血病发生中扮演关键的致癌作用。LI等[65]研究发现,FTO在t(11q23)/MLL重排、t(15;17)/PML-RARA、FLT3-ITD和/或NPM1突变的急性髓系白血病细胞中高表达;敲除FTO可通过减少抗白血病基因锚蛋白重复序列和SOCS盒包含2(Ankyrin repeat and SOCS box containing 2,ASB2)和维甲酸受体α(Retinoic acid receptor A gene ,RARA)转录mRNA的m6A水平并增强其表达,进而显著降低白血病干细胞的自我更新和髓系分化,抑制其集落形成的能力,这些研究结果显示了m6A甲基化及其调节的相应蛋白质表达在肿瘤发生中的重要性,不仅深刻揭示了白血病的发生发展与恶化重要机制,也为白血病治疗及其药物反应提供了有意义地研究策略。此外,WANG等[66]新近研究发现,肿瘤坏死因子TNF-ɑ处理的间充质干细胞内FTO水平表达降低,进而降低FTO对同源域蛋白Nanog mRNA去甲基化作用,使其m6A甲基化水平显著增加而导致mRNA转录和蛋白表达降低,结果是显著抑制了间充质干细胞分化为汗腺细胞的能力,这提示通过调控间充质干细胞的m6A水平实现细胞定向分化调节可能是再生医学和组织工程学研究与应用的新策略。 YTH家族的阅读蛋白调控m6A在干细胞分化过程中同样扮演着不可或缺的重要角色。WANG等[67]研究发现YTHDF1和YTHDF3在调控胚胎干细胞的分化中发挥着不同的功能,敲除YTHDF3基因将导致胚胎干细胞多能性丧失,表现为心肌细胞特异性基因如肌细胞增强因子2C(myocyte enhancer factor 2C,Mef2c)、心肌激动蛋白α1(recombinant Actin Alpha 1,ACTC1)和肌球蛋白重链7(recombinant Myosin Heavy Chain 7,Myh7)等表达上调,即胚胎干细胞向心肌细胞分化;相反地,YTHDF1基因功能缺失则引起心肌细胞分化严重受损,伴有心肌特异性基因和多潜能性基因(如Nanog和SOX2)均表达下调,且YTHDF3似乎部分通过抑制YTHDF1作用实现细胞分化调节,因此YTHDF1和YTHDF3在决定干细胞命运和定向分化中相互关联又协同配合发挥不同的作用。HECK等[68]研究发现,YTHDF2高表达于诱导多能干细胞(induced pluripotent stem cells,iPSCs)中,但在神经分化过程中表达下调,进一步RNA测序发现YTHDF2通过直接调节m6A修饰而抑制与神经发育相关的特异性基因的mRNA,包括阳离子转运调控样蛋白1(cation transport regulator-like protein,CHAC1)、WD40重复蛋白62(WD40-repeat protein 62,WDR62)、睾丸特异蛋白Y编码的样蛋白4(testis?specific protein Y-encoded-like 4,TSPYL4)和CREB结合蛋白(CREB binding protein,CREBBP)等,而剔除YTHDF2基因表达则促进神经特异性基因的mRNA转录和蛋白表达。METTL3表达缺失则抑制YTHDF1介导的JAK2信号和阻断YTHDF2依赖的细胞因子信号转导抑制分子(suppressors of cytokine signaling 3,SOCS3)mRNA的衰减,进而抑制诱导多能干细胞的自我更新能力并诱导特异性细胞分化[69]。神经干细胞中也发现了类似的现象,神经干细胞中YTHDF2表达上调将抑制JAK/STAT信号级联反应,有助于神经保护和轴突生长[70]。总之,YTH家族阅读蛋白亦通过调控多种不同作用靶基因mRNA中m6A水平实现干细胞分化,通过调控m6A修饰维持干细胞的分化能力为干细胞介导的再生医学提供了一个新的潜在治疗靶点。 因此,m6A在调控干细胞定向分化过程中发挥着关键作用,尤其是间充质干细胞的成骨分化,可作为间充质干细胞分化的标志物。但目前的研究大多主要针对间充质干细胞的成骨分化方面的研究,其对于成脂肪,成软骨等其它分化方向的研究,以及其它种类的干细胞的研究较少,且具体机制研究较浅。此外,对于多能诱导干细胞,胚胎干细胞等干细胞向心肌细胞和神经细胞等永久性细胞的定向分化,对再生医学的发展有着极大的推动意义,突出了干细胞定向分化对于恶性疾病的治疗方面的重要作用,但目前针对这方面的研究较少。因此需进一步探讨m6A甲基化各种蛋白酶对干细胞定向分化为心肌细胞、神经细胞等永久细胞的作用及其相关机制,见图4。 "

2.2.3 m6A调控干细胞的免疫学特性 免疫调节得益于干细胞的免疫调节作用,干细胞利用其旁分泌效应,可以通过分泌一些炎症相关因子,抑制免疫反应,该特性可以用于多种疾病的治疗以及预后的改善。m6A甲基化可以通过靶向免疫检查点、干扰素调节因子以及肿瘤坏死因子的表达等,抑制免疫应答,调节免疫稳态的平衡。新的研究进展表明,m6A甲基化与干细胞的免疫调节特性关系密切,并在其调节中发挥着重要作用。 有研究表明,m6A不仅可能参与干细胞的免疫调节,而且涉及肿瘤干细胞免疫逃逸过程,而其进一步的运动和迁徙能力又促进了干细胞的转移。其中FTO作为m6A的重要去甲基化酶在多种肿瘤干细胞的免疫逃逸中起关键作用[71]。急性髓系白血病治疗应用阿扎胞苷等低甲基化药物诱导的药物抵抗可能与免疫检查点基因细胞程序性死亡受体1(programmed cell death 1,PD-1)、细胞程序性死亡配体1(programmed cell death 1 ligand,PD-L1)和细胞程序性死亡配体2(programmed cell death 1 ligand,PD-L2)的表达引起继发性免疫逃逸有关;进一步研究显示这些免疫检查点基因表达上调系去甲基化酶FTO增加导致m6A水平降低引起,与其他m6A调节酶ALKBH5、METTL3和METTL14无关[72]。SU等[71]研究亦发现抑制FTO会增加人急性髓系白血病细胞对T细胞毒性敏感,克服低甲基化诱导的免疫逃逸,且伴随细胞中白细胞免疫球蛋白样受体B4(leukocyte immunoglobulin (Ig)-like receptor B4,LILRB4)表达降低,因此FTO/m6A/LILRB4轴可能是急性髓系白血病发生药物抵抗和免疫逃逸的重要机制与作用靶点。此外,FTO亦高表达于LSCs/LICs,药物抑制或基因敲除FTO将显著抑制这些细胞的自我更新能力,这与FTO正性调节MYC和CEBPA基因有关[71]。同样作为去甲基化酶酶分子ALKBH5在免疫调节方面也发挥着重要作用。胶质母细胞瘤的复发也被认为是胶质母细胞瘤干细胞引起的,这种干细胞具有显著的化疗耐受性,从而导致胶质母细胞瘤对常规的放射化疗治疗具有抵抗性和较强的侵袭性,这是该肿瘤的高复发性的重要原因。抑制m6A RNA的去甲基化酶ALKBH5可通过控制同源修复来提高其辐射敏感性并抑制其侵袭性,伴相关的调节同源重组的基因表达下降,包括细胞周期检查点激酶1(cell cycle checkpoint kinase 1,CHK1)和DNA双链断裂修复蛋白RAD51等[73]。因此,在免疫逃避方面,肿瘤干细胞通过对m6A修饰的调节有力地控制细胞的转移和侵袭,对减缓疾病的进展以及为患者争取更多的治疗时间,增加预后效果有重要的临床意义。 同样作为甲基阅读蛋白在免疫方面也起着重要作用,Zhang等[74]研究发现,在肝癌细胞中敲除和过表达YTHDF2将导致八聚体结合转录因子(octamer-binding transcription factor,OCT4)mRNA 5′-非翻译区m6A甲基化水平的降低和升高,从而导致OCT4蛋白表达降低和升高而影响肿瘤干细胞的表型;进一步研究发现,YTHDF2缺失通过降低OCT4 mRNA的m6A水平减轻了裸鼠原位移植后的肿瘤负荷,并抑制肿瘤肺转移,这些研究结果表明,YTHDF2通过调控OCT4 mRNA的m6A甲基化,促进肿瘤干细胞的肝脏表型和肿瘤转移,进而降低肝细胞癌患者的生存率。此外,m6A还可能通过调节多个基因表达而影响干细胞表型。Mapperley等[75]研究发现YTHDF2缺失显著影响造血干细胞和多系统造血功能,表现为幼年小鼠来源的YTHDF2缺陷的造血干细胞存在连续的细胞移植失败,并伴有大量m6A调控的炎症相关因子如STAT1转录因子、肿瘤坏死因子α和干扰素调节因子7等的mRNA的转录水平升高,即YTHDF2缺失将显著激活细胞中促炎症通路,因此以调节m6A为主的YTHDF2系造血干细胞炎症通路的抑制因子,在细胞中发挥抗炎症损伤作用。 综上所述,m6A在干细胞的免疫调节方面起着非常重要的作用,尤其是肿瘤干细胞的免疫逃逸以及对放化疗的耐受性,导致其远处转移,m6A甲基化的修饰治疗有望使得肿瘤干细胞的侵袭性降低,进而恶性程度降低。现阶段针对m6A甲基化修饰对干细胞的免疫调节特性的研究尚不全面,对于T细胞的活化、树突状细胞的形成、淋巴细胞的增殖以及巨噬细胞的吞噬功能等方面的影响还处于初级阶段,有望进一步深入探索其作用机制,为实现免疫相关恶性疾病的治疗提供坚实的理论依据以及临床实践。此外,间充质干细胞是天生的免疫豁免细胞,这是其免疫调节的大前提,同时具有定向分化的功能,因此,间充质干细胞对于干细胞移植治疗恶性疾病,推动再生医学的发展有着重要作用。但是目前针对干细胞通过免疫调控靶向治疗恶性疾病方面存在争议,细胞移植后的生物安全性问题不容忽视[78-80]。 "

| [1] ALMOTIRI A, ALZAHRANI H, MENENDEZ-GONZALEZ JB, et al. Zeb1 modulates hematopoietic stem cell fates required for suppressing acute myeloid leukemia. J Clin Invest. 2021;131(1):e129115. [2] WANG W, MAO J, CHEN Y, et al. Naringin promotes osteogenesis and ameliorates osteoporosis development by targeting JAK2/STAT3 signalling. Clin Exp Pharmacol Physiol. 2022;49(1):113-121. [3] YU T, QI X, ZHANG L, et al. Dynamic reprogramming and function of RNA N6-methyladenosine modification during porcine early embryonic development. Zygote. 2021;29(6):417-426. [4] DUNN DB, SMITH JD. Occurrence of a new base in the deoxyribonucleic acid of a strain of Bacterium coli. Nature. 1955;175(4451):336-337. [5] DESROSIERS R, FRIDERICI K, ROTTMAN F. Identification of methylated nucleosides in messenger RNA from Novikoff hepatoma cells. Proc Natl Acad Sci U S A. 1974;71(10):3971-3975. [6] ZHANG C, CHEN Y, SUN B, et al. m(6)A modulates haematopoietic stem and progenitor cell specification. Nature. 2017,549(7671):273-276. [7] JIAN D, WANG Y, JIAN L, et al. METTL14 aggravates endothelial inflammation and atherosclerosis by increasing FOXO1 N6-methyladeosine modifications. Theranostics. 2020;10(20):8939-8956. [8] ARCIDIACONO OA, KREJčí J, BáRTOVá E. The distinct function and localization of METTL3/METTL14 and METTL16 enzymes in cardiomyocytes. Int J Mol Sci. 2020;21(21):8139. [9] NANCE DJ, SATTERWHITE ER, BHASKAR B, et al. Characterization of METTL16 as a cytoplasmic RNA binding protein. PLoS One. 2020;15(1): e0227647. [10] PING XL, SUN BF, WANG L, et al. Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Res. 2014;24(2):177-189. [11] ZHU W, WANG JZ, WEI JF, et al. Role of m6A methyltransferase component VIRMA in multiple human cancers (Review). Cancer Cell Int. 2021;21(1):172. [12] WANG X, TIAN L, LI Y, et al. RBM15 facilitates laryngeal squamous cell carcinoma progression by regulating TMBIM6 stability through IGF2BP3 dependent. J Exp Clin Cancer Res. 2021;40(1):80. [13] YANG Y, HSU PJ, CHEN YS, et al. Dynamic transcriptomic m6A decoration:writers, erasers, readers and functions in RNA metabolism. Cell Res. 2018;28(6):616-624. [14] JIA G, FU Y, ZHAO X, et al. N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat Chem Biol. 2011; 7(12):885-887. [15] GUO X, LI K, JIANG W, et al. RNA demethylase ALKBH5 prevents pancreatic cancer progression by posttranscriptional activation of PER1 in an m6A-YTHDF2-dependent manner. Mol Cancer. 2020;19(1):91. [16] HAZRA D, CHAPAT C, GRAILLE M. m⁶A mRNA destiny: chained to the rhYTHm by the YTH-containing proteins. Genes (Basel). 2019; 10(1):49. [17] HE L, LI H, WU A, et al. Functions of N6-methyladenosine and its role in cancer. Mol Cancer. 2019;18(1):176. [18] LIU Y, SHI SL. The roles of hnRNP A2/B1 in RNA biology and disease. Wiley Interdiscip Rev RNA. 2021;12(2):e1612. [19] LIU N, ZHOU KI, PARISIEN M, et al. N6-methyladenosine alters RNA structure to regulate binding of a low-complexity protein. Nucleic Acids Res. 2017;45(10):6051-6063. [20] HAUSSMANN IU, BODI Z, SANCHEZ-MORAN E, et al. m6A potentiates Sxl alternative pre-mRNA splicing for robust Drosophila sex determination. Nature. 2016;540(7632):301-304. [21] CHEN ZH, CHEN TQ, ZENG ZC, et al. Nuclear export of chimeric mRNAs depends on an lncRNA-triggered autoregulatory loop in blood malignancies. Cell Death Dis. 2020;11(7):566. [22] WESTMARK CJ, MALONEY B, ALISCH RS, et al. FMRP regulates the nuclear export of Adam9 and Psen1 mRNAs: secondary analysis of an N6-methyladenosine dataset. Sci Rep. 2020;10(1):10781. [23] LIU T, WEI Q, JIN J, et al. The m6A reader YTHDF1 promotes ovarian cancer progression via augmenting EIF3C translation. Nucleic Acids Res. 2020;48(7):3816-3831. [24] YU J, CHEN M, HUANG H, et al. Dynamic m6A modification regulates local translation of mRNA in axons. Nucleic Acids Res. 2018;46(3): 1412-1423. [25] SONG P, FENG L, LI J, et al. β-catenin represses miR455-3p to stimulate m6A modification of HSF1 mRNA and promote its translation in colorectal cancer. Mol Cancer. 2020;19(1):129. [26] ZHANG L, WAN Y, ZHANG Z, et al. IGF2BP1 overexpression stabilizes PEG10 mRNA in an m6A-dependent manner and promotes endometrial cancer progression. Theranostics. 2021;11(3):1100-1114. [27] PENG J, ZHENG H, LIU F, et al. The m6A methyltransferase METTL3 affects autophagy and progression of nasopharyngeal carcinoma by regulating the stability of lncRNA ZFAS1. Infect Agent Cancer. 2022; 17(1):1. [28] LI B, ZHU L, LU C, et al. circNDUFB2 inhibits non-small cell lung cancer progression via destabilizing IGF2BPs and activating anti-tumor immunity. Nat Commun. 2021;12(1):295. [29] HUANG H, WANG Y, KANDPAL M, et al. FTO-dependent N 6-methyladenosine modifications inhibit ovarian cancer stem cell self-renewal by blocking cAMP signaling. Cancer Res. 2020;80(16):3200-3214. [30] TIAN C, HUANG Y, LI Q, et al. Mettl3 regulates osteogenic differentiation and alternative splicing of vegfa in bone marrow mesenchymal stem cells. Int J Mol Sci. 2019;20(3):551. [31] CHEN J, WANG C, FEI W, et al. Epitranscriptomic m6A modification in the stem cell field and its effects on cell death and survival. Am J Cancer Res. 2019;9(4):752-764. [32] PENDLETON KE, CHEN B, LIU K, et al. The U6 snRNA m(6)A methyltransferase METTL16 regulates SAM synthetase intron retention. Cell. 2017;169(5):824-835.e14. [33] YUE Y, LIU J, CUI X, et al. VIRMA mediates preferential m(6)A mRNA methylation in 3’UTR and near stop codon and associates with alternative polyadenylation. Cell Discov. 2018;4:10. [34] TRAN NT, SU H, KHODADADI-JAMAYRAN A, et al. The AS-RBM15 lncRNA enhances RBM15 protein translation during megakaryocyte differentiation. EMBO Rep. 2016;17(6):887-900. [35] WAN C, BORGESON B, PHANSE S, et al. Panorama of ancient metazoan macromolecular complexes. Nature. 2015;525(7569):339-344. [36] ZHENG G, DAHL JA, NIU Y, et al. Sprouts of RNA epigenetics: the discovery of mammalian RNA demethylases. RNA Biol. 2013;10(6):915-918. [37] KIM H, LEE YS, KIM SM, et al. RNA demethylation by FTO stabilizes the FOXJ1 mRNA for proper motile ciliogenesis. Dev Cell. 2021;56(8):1118-1130.e6. [38] ZHENG G, DAHL JA, NIU Y, et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol Cell. 2013;49(1):18-29. [39] ROUNDTREE IA, LUO GZ, ZHANG Z, et al. YTHDC1 mediates nuclear export of N6-methyladenosine methylated mRNAs. Elife. 2017;6:e31311. [40] XIAO W, ADHIKARI S, DAHAL U, et al. Nuclear m(6)A Reader YTHDC1 Regulates mRNA Splicing. Mol Cell. 2016;61(4):507-519. [41] MAO Y, DONG L, LIU XM, et al. m(6)A in mRNA coding regions promotes translation via the RNA helicase-containing YTHDC2. Nat Commun. 2019;10(1):5332. [42] DIXIT D, PRAGER BC, GIMPLE RC, et al. The RNA m6A reader YTHDF2 maintains oncogene expression and is a targetable dependency in glioblastoma stem cells. Cancer Discov. 2021;11(2):480-499. [43] HOU G, ZHAO X, LI L, et al. SUMOylation of YTHDF2 promotes mRNA degradation and cancer progression by increasing its binding affinity with m6A-modified mRNAs. Nucleic Acids Res. 2021;49(5):2859-2877. [44] SHI H, WANG X, LU Z, et al. YTHDF3 facilitates translation and decay of N6-methyladenosine-modified RNA. Cell Res. 2017;27(3):315-328. [45] HUANG H, WENG H, SUN W, et al. Recognition of RNA N6-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat Cell Biol. 2018;20(3):285-295. [46] YING Y, MA X, FANG J, et al. EGR2-mediated regulation of m(6)A reader IGF2BP proteins drive RCC tumorigenesis and metastasis via enhancing S1PR3 mRNA stabilization. Cell Death Dis. 2021;12(8):750. [47] YIN M, CHENG M, LIU C, et al. HNRNPA2B1 as a trigger of RNA switch modulates the miRNA-mediated regulation of CDK6. iScience. 2021; 24(11):103345. [48] HUANG GZ, WU QQ, ZHENG ZN, et al. M6A-related bioinformatics analysis reveals that HNRNPC facilitates progression of OSCC via EMT. Aging (Albany NY). 2020;12(12):11667-11684. [49] HUANG XT, LI JH, ZHU XX, et al. HNRNPC impedes m6A-dependent anti-metastatic alternative splicing events in pancreatic ductal adenocarcinoma. Cancer Lett. 2021;518:196-206. [50] ZHOU KI, SHI H, LYU R, et al. Regulation of co-transcriptional pre-mRNA splicing by m(6)A through the low-complexity protein hnRNPG. Mol Cell. 2019;76(1):70-81.e9. [51] MA Z, JI J. N6-methyladenosine (m6A) RNA modification in cancer stem cells. Stem Cells. 2020. doi: 10.1002/stem.3279. [52] CUI Q, SHI H, YE P, et al. m6A RNA methylation regulates the self-renewal and tumorigenesis of glioblastoma stem cells. Cell Rep. 2017; 18(11):2622-2634. [53] LI F, YI Y, MIAO Y, et al. N6-methyladenosine modulates nonsense-mediated mrna decay in human glioblastoma. Cancer Res. 2019;79(22): 5785-5798. [54] ZHANG S, ZHAO BS, ZHOU A, et al. m6A demethylase ALKBH5 maintains tumorigenicity of glioblastoma stem-like cells by sustaining FOXM1 expression and cell proliferation program. Cancer Cell. 2017;31(4):591-606.e6. [55] WU Z, SHI Y, LU M, et al. METTL3 counteracts premature aging via m6A-dependent stabilization of MIS12 mRNA. Nucleic Acids Res. 2020; 48(19):11083-11096. [56] SHENG Y, WEI J, YU F, et al. A critical role of nuclear m6A reader YTHDC1 in leukemogenesis by regulating MCM complex-mediated DNA replication. Blood. 2021;138(26):2838-2852. [57] SHEN C, SHENG Y, ZHU AC, et al. RNA demethylase ALKBH5 selectively promotes tumorigenesis and cancer stem cell self-renewal in acute myeloid leukemia. Cell Stem Cell. 2020;27(1):64-80.e9. [58] LI Z, QIAN P, SHAO W, et al. Suppression of m6A reader Ythdf2 promotes hematopoietic stem cell expansion. Cell Res. 2018;28(9):904-917. [59] BAI Y, YANG C, WU R, et al. YTHDF1 regulates tumorigenicity and cancer stem cell-like activity in human colorectal carcinoma. Front Oncol. 2019;9:332. [60] YU J, SHEN L, LIU Y, et al. The m6A methyltransferase METTL3 cooperates with demethylase ALKBH5 to regulate osteogenic differentiation through NF-κB signaling. Mol Cell Biochem. 2020;463(1-2):203-210. [61] YAN G, YUAN Y, HE M, et al. m6A methylation of precursor-miR-320/RUNX2 controls osteogenic potential of bone marrow-derived mesenchymal stem cells. Mol Ther Nucleic Acids. 2020;19:421-436. [62] WU Y, XIE L, WANG M, et al. Mettl3-mediated m6A RNA methylation regulates the fate of bone marrow mesenchymal stem cells and osteoporosis. Nat Commun. 2018,9(1):4772. [63] PAN ZP, WANG B, HOU DY, et al. METTL3 mediates bone marrow mesenchymal stem cell adipogenesis to promote chemoresistance in acute myeloid leukaemia. FEBS Open Bio. 2021;11(6):1659-1672. [64] WENG H, HUANG H, WU H, et al. METTL14 inhibits hematopoietic stem/progenitor differentiation and promotes leukemogenesis via mRNA m6A modification. Cell Stem Cell, 2018;22(2):191-205.e9. [65] LI Z, WENG H, SU R, et al. FTO plays an oncogenic role in acute myeloid leukemia as a N6-methyladenosine RNA demethylase. Cancer Cell. 2017;31(1):127-141. [66] WANG Y, WANG R, YAO B, et al. TNF-α suppresses sweat gland differentiation of MSCs by reducing FTO-mediated m6A-demethylation of Nanog mRNA. Sci China Life Sci. 2020;63(1):80-91. [67] WANG S, ZHANG J, WU X, et al. Differential roles of YTHDF1 and YTHDF3 in embryonic stem cell-derived cardiomyocyte differentiation. RNA Biol. 2021;18(9):1354-1363. [68] HECK AM, RUSSO J, WILUSZ J, et al. YTHDF2 destabilizes m6A-modified neural-specific RNAs to restrain differentiation in induced pluripotent stem cells. RNA. 2020;26(6):739-755. [69] WU R, LIU Y, ZHAO Y, et al. m6A methylation controls pluripotency of porcine induced pluripotent stem cells by targeting SOCS3/JAK2/STAT3 pathway in a YTHDF1/YTHDF2-orchestrated manner. Cell Death Dis. 2019;10(3):171. [70] LI M, ZHAO X, WANG W, et al. Ythdf2-mediated m6A mRNA clearance modulates neural development in mice. Genome Biol. 2018;19(1):69. [71] SU R,DONG L, LI Y, et al. Targeting FTO suppresses cancer stem cell maintenance and immune evasion. Cancer Cell. 2020;38(1):79-96.e11. [72] STAHL M, GOLDBERG AD. Immune checkpoint inhibitors in acute myeloid leukemia:novel combinations and therapeutic targets. Curr Oncol Rep. 2019;21(4):37. [73] KOWALSKI-CHAUVEL A, LACORE MG, ARNAUDUC F, et al. The m6A RNA demethylase ALKBH5 promotes radioresistance and invasion capability of glioma stem cells. Cancers (Basel). 2020;13(1):40. [74] ZHANG C, HUANG S, ZHUANG H, et al. YTHDF2 promotes the liver cancer stem cell phenotype and cancer metastasis by regulating OCT4 expression via m6A RNA methylation. Oncogene. 2020;39(23):4507-4518. [75] MAPPERLEY C, VAN DE LAGEMAAT LN, LAWSON H, et al. The mRNA m6A reader YTHDF2 suppresses proinflammatory pathways and sustains hematopoietic stem cell function. J Exp Med. 2021;218(3) e20200829. [76] SUN W, ZHANG B, BIE Q, et al. The role of RNA methylation in regulating stem cell fate and function-focus on m6A. Stem Cells Int. 2021;2021:8874360. [77] CHENG Y, LUO H, IZZO F, et al. m6A RNA methylation maintains hematopoietic stem cell identity and symmetric commitment. Cell Rep. 2019;28(7):1703-1716.e6. [78] LALU MM, MAZZARELLO S, ZLEPNIG J, et al. Safety and efficacy of adult stem cell therapy for acute myocardial infarction and ischemic heart failure (SafeCell Heart): a systematic review and meta-analysis. Stem Cells Transl Med. 2018,7(12):857-866. [79] GAO L, XU W, LI T, et al. Stem cell therapy: a promising therapeutic method for intracerebral hemorrhage. Cell Transplant. 2018;27(12): 1809-1824. [80] KOLAJA K. Stem cells and stem cell-derived tissues and their use in safety assessment. J Biol Chem. 2014,289(8):4555-4561. [81] TSUCHIYA K, YOSHIMURA K, IWASHITA Y, et al. m6A demethylase ALKBH5 promotes tumor cell proliferation by destabilizing IGF2BPs target genes and worsens the prognosis of patients with non-small-cell lung cancer. Cancer Gene Ther. 2022. doi: 10.1038/s41417-022-00451-8. [82] HU Y, GONG C, LI Z, et al. Demethylase ALKBH5 suppresses invasion of gastric cancer via PKMYT1 m6A modification. Mol Cancer. 2022; 21(1):34. [83] CHEN Y, ZHAO Y, CHEN J, et al. ALKBH5 suppresses malignancy of hepatocellular carcinoma via m6A-guided epigenetic inhibition of LYPD1. Mol Cancer. 2020;19(1):123. [84] CAI W, JI Y, HAN L, et al. METTL3-dependent glycolysis regulates dental pulp stem cell differentiation. J Dent Res. 2022;101(5):580-589. [85] HOU J, WANG Z, LI H, et al. Gene signature and identification of clinical trait-related m6A regulators in pancreatic cancer. Front Genet. 2020;11:522. [86] CHEN L, WANG X. Relationship between the genetic expression of WTAP and bladder cancer and patient prognosis. Oncol Lett. 2018; 16(6):6966-6970. [87] DENG H, TANG F, ZHOU M, et al. Identification and validation of n6-methyladenosine-related biomarkers for bladder cancer: implications for immunotherapy. Front Oncol. 2022;12:820242. [88] CHEN F, WANG Q, ZHOU Y. The construction and validation of an RNA binding protein-related prognostic model for bladder cancer. BMC Cancer. 2021;21(1):244. |
| [1] | Nong Fuxiang, Jiang Zhixiong, Li Yinghao, Xu Wencong, Shi Zhilan, Luo Hui, Zhang Qinglang, Zhong Shuang, Tang Meiwen. Bone cement augmented proximal femoral nail antirotation for type A3.3 intertrochanteric femoral fracturalysis [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(在线): 1-10. |
| [2] | Dang Yi, Du Chengyan, Yao Honglin, Yuan Nenghua, Cao Jin, Xiong Shan, Zhang Dingmei, Wang Xin. Hormonal osteonecrosis and oxidative stress [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(9): 1469-1476. |
| [3] | Pan Zhongjie, Qin Zhihong, Zheng Tiejun, Ding Xiaofei, Liao Shijie. Targeting of non-coding RNAs in the pathogenesis of the osteonecrosis of the femoral head [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(9): 1441-1447. |
| [4] | Cai Zhihao, Xie Zhaoyong. Femoral neck anteversion measurement assessment: how to establish a unified method and standard [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(9): 1448-1454. |
| [5] | Wang Ji, Zhang Min, Yang Zhongya, Zhang Long. A review of physical activity intervention in type 2 diabetes mellitus with sarcopenia [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(8): 1272-1277. |
| [6] | Nie Chenchen, Su Kaiqi, Gao Jing, Fan Yongfu, Ruan Xiaodi, Yuan Jie, Duan Zhaoyuan, Feng Xiaodong. The regulatory role of circular RNAs in cerebral ischemia-reperfusion injury [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(8): 1286-1291. |
| [7] | Gao Yu, Han Jiahui, Ge Xin. Immunoinflammatory microenvironment after spinal cord ischemia-reperfusion injury [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(8): 1300-1305. |
| [8] | Xu Xingxing, Wen Chaoju, Meng Maohua, Wang Qinying, Chen Jingqiao, Dong Qiang. Carbon nanomaterials in oral implant [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(7): 1062-1070. |
| [9] | Li Cheng, Zheng Guoshuang, Kuai Xiandong, Yu Weiting. Alginate scaffold in articular cartilage repair [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(7): 1080-1088. |
| [10] | Xu Cong, Zhao He, Sun Yan. Regeneration of facial nerve injury repaired by biomaterial nerve conduits [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(7): 1089-1095. |
| [11] | Chen Shisong, Liu Xiaohong, Xu Zhiyun. Current status and prospects of bioprosthetic heart valves [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(7): 1096-1102. |
| [12] | Lu Di, Zhang Cheng, Duan Rongquan, Liu Zongxiang. Osteoinductive properties of calcium phosphate ceramic bone repair materials [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(7): 1103-1109. |
| [13] | Shi Yehong, Wang Cheng, Chen Shijiu. Early thrombosis and prevention of small-diameter blood vessel prosthesis [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(7): 1110-1116. |
| [14] | Tang Haotian, Liao Rongdong, Tian Jing. Application and design of piezoelectric materials for bone defect repair [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(7): 1117-1125. |
| [15] | Xue Ting, Zhang Xinri, Kong Xiaomei. Mesenchymal stem cell therapy for pneumoconiosis using nanomaterials combined with multi-modal molecular imaging [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(7): 1133-1140. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||