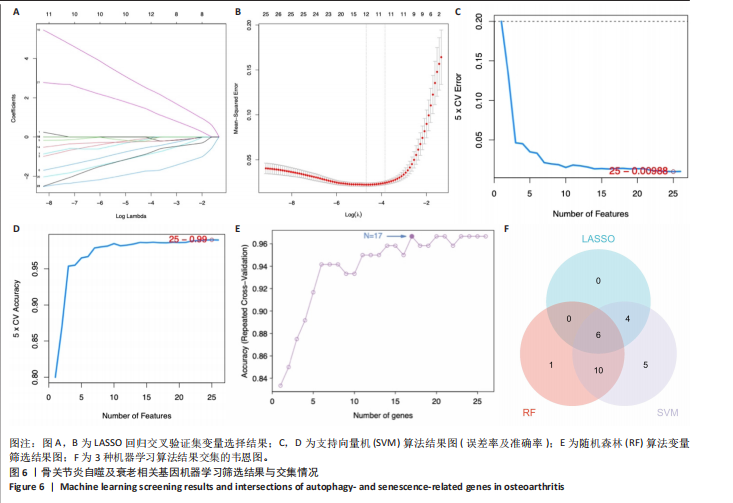
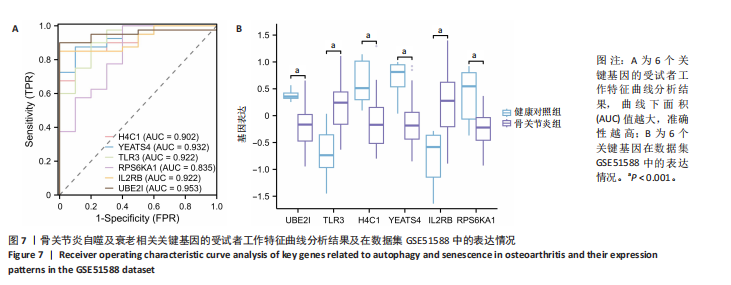
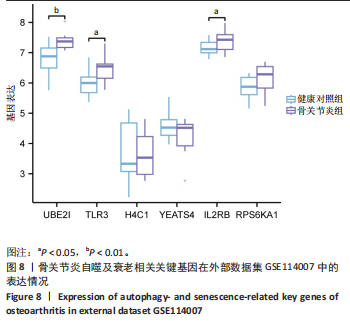
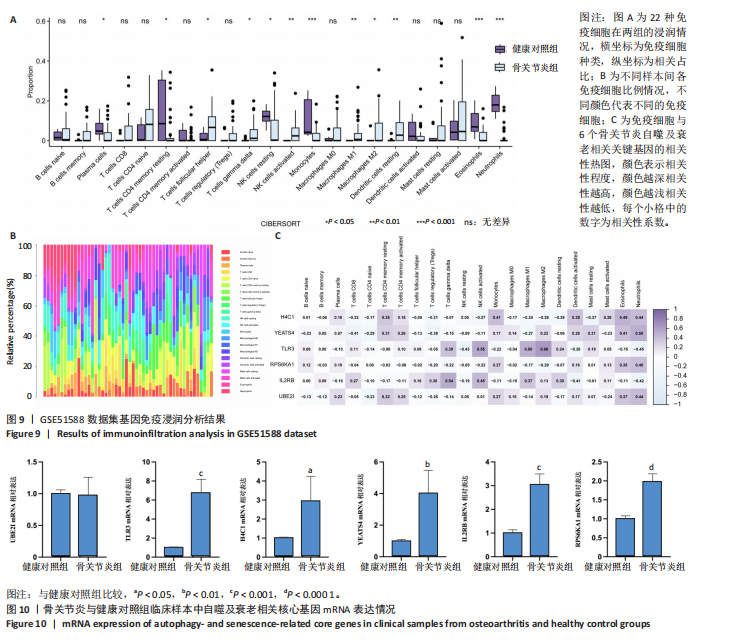
[1] XU T, LIU K, FAN J, et al. Metformin mitigates osteoarthritis progression by modulating the PI3K/AKT/mTOR signaling pathway and enhancing chondrocyte autophagy. Open Life Sci. 2024;19(1):20220922.
[2] KONG P, AHMAD RE, ZULKIFLI A, et al. The role of autophagy in mitigating osteoarthritis progression via regulation of chondrocyte apoptosis: A review. Joint Bone Spine. 2024;91(3):105642.
[3] LI J, YU G, HE Z, et al. Kojic acid alleviates osteoarthritis by attenuating inflammation and restoring impaired autophagy through regulating NF-κB and PI3K/AKT/mTOR signaling pathways: An in vivo and in vitro study. Int Immunopharmacol. 2025;147:114022.
[4] LU R, HE Z, ZHANG W, et al. Oroxin B alleviates osteoarthritis through anti-inflammation and inhibition of PI3K/AKT/mTOR signaling pathway and enhancement of autophagy. Front Endocrinol (Lausanne). 2022;13:1060721.
[5] 薛庆云,王坤正,裴福兴,等.中国40岁以上人群原发性骨关节炎患病状况调查[J].中华骨科杂志,2015,35(12):1206-1212.
[6] CHEN Y, PAN X, ZHAO J, et al. Icariin alleviates osteoarthritis through PI3K/Akt/mTOR/ULK1 signaling pathway. Eur J Med Res. 2022;27(1):204.
[7] LIU Y, ZHANG Z, LI T, et al. Senescence in osteoarthritis: from mechanism to potential treatment. Arthritis Res Ther. 2022;24(1):174.
[8] JEON OH, KIM C, LABERGE RM, et al. Local clearance of senescent cells attenuates the development of post-traumatic osteoarthritis and creates a pro-regenerative environment. Nat Med. 2017;23(6):775-781.
[9] SHAO Y, ZHANG H, GUAN H, et al. PDZK1 protects against mechanical overload-induced chondrocyte senescence and osteoarthritis by targeting mitochondrial function. Bone Res. 2024;12(1):41.
[10] CHEN Z, TANG M, WU Z, et al. Increased Rab1a accelerates osteoarthritis by inhibiting autophagy via activation of the mTORC1-S6K pathway. J Adv Res. 2025;75:607-619.
[11] LIU Z, WANG T, SUN X, et al. Autophagy and apoptosis: regulatory factors of chondrocyte phenotype transition in osteoarthritis. Hum Cell. 2023;36(4):1326-1235.
[12] PAN Y, YANG Y, FAN M, et al. Progranulin regulation of autophagy contributes to its chondroprotective effect in osteoarthritis. Genes Dis. 2023;10(4):1582-1595.
[13] 李文铭,李永航,严才平,等.增强自噬的可注射水凝胶微球改善软骨微环境抗软骨细胞衰老[J]. 中国组织工程研究,2025,29(22):4710-4719.
[14] FENG K, YE T, XIE X, et al. ESC-sEVs alleviate non-early-stage osteoarthritis progression by rejuvenating senescent chondrocytes via FOXO1A-autophagy axis but not inducing apoptosis. Pharmacol Res. 2024;209:107474.
[15] BINVIGNAT M, PEDOIA V, BUTTE A J, et al. Use of machine learning in osteoarthritis research: a systematic literature review. RMD Open. 2022;8(1): e001998.
[16] XU W, SUN Y, ZHAO S, et al. Identification and validation of autophagy-related genes in primary open-angle glaucoma. BMC Med Genomics. 2023;16(1):287.
[17] LI ZH, LI SP, LI YH, et al. Identification of aging-related biomarkers for intervertebral disc degeneration in whole blood samples based on bioinformatics and machine learning. Front Immunol. 2025;16:1565945.
[18] AVELAR RA, ORTEGA JG, TACUTU R, et al. A multidimensional systems biology analysis of cellular senescence in aging and disease. Genome Biol. 2020;21(1):91.
[19] SUN M, YANG Q, HU C, et al. Identification and Validation of Autophagy-Related Genes in Sepsis-Induced Acute Respiratory Distress Syndrome and Immune Infiltration. J Inflamm Res. 2022;15:2199-2212.
[20] DING JT, HONG FF, YANG SL. Roles of autophagy in rheumatoid arthritis. Clin Exp Rheumatol. 2022;40(11):2179-2187.
[21] CHENG L, CHANG S, TAN Y, et al. Platelet-rich plasma combined with isometric quadriceps contraction regulates autophagy in chondrocytes via the PI3K/AKT/mTOR pathway to promote cartilage repair in knee osteoarthritis. Regen Ther. 2025;28:81-89.
[22] TANG L, DING J, YANG K, et al. New insights into the mechanisms and therapeutic strategies of chondrocyte autophagy in osteoarthritis. J Mol Med (Berl). 2024; 102(10):1229-1244.
[23] LEE DY, BAHAR ME, KIM CW, et al. Autophagy in Osteoarthritis: A Double-Edged Sword in Cartilage Aging and Mechanical Stress Response: A Systematic Review. J Clin Med. 2024;13(10):3005.
[24] MUSUMECI G, CASTROGIOVANNI P, TROVATO FM, et al. Biomarkers of Chondrocyte Apoptosis and Autophagy in Osteoarthritis. Int J Mol Sci. 2015; 16(9):20560-20575.
[25] ANDREI C, MIHAI DP, NITULESCU GM, et al. Modulating Autophagy in Osteoarthritis: Exploring Emerging Therapeutic Drug Targets. Int J Mol Sci. 2024; 25(24):13695.
[26] LV X, ZHAO T, DAI Y, et al. New insights into the interplay between autophagy and cartilage degeneration in osteoarthritis. Front Cell Dev Biol. 2022;10:1089668.
[27] WU J, HUANG S, YU Y, et al. Human adipose and synovial-derived MSCs synergistically attenuate osteoarthritis by promoting chondrocyte autophagy through FoxO1 signaling. Stem Cell Res Ther. 2024;15(1):261.
[28] LORENZO-GÓMEZ I, NOGUEIRA-RECALDE U, GARCÍA-DOMÍNGUEZ C, et al. Defective chaperone-mediated autophagy is a hallmark of joint disease in patients with knee osteoarthritis. Osteoarthritis Cartilage. 2023;31(7):919-933.
[29] KHALEQUE MA, KIM JH, TANVIR MAH, et al. Significance of Necroptosis in Cartilage Degeneration. Biomolecules. 2024;14(9):1192.
[30] WANG H, ZHOU J, LU Y, et al. The interplay between autophagy and programmed cell death in osteoarthritis: insights into mechanisms and therapeutic targets. Mol Cell Biochem. 2025;480(8):4627-4646.
[31] DIEKMAN BO, LOESER RF. Aging and the emerging role of cellular senescence in osteoarthritis. Osteoarthritis Cartilage. 2024;32(4):365-371.
[32] TAMATTA R, PAI V, JAISWAL C, et al. Neuroinflammaging and the Immune Landscape: The Role of Autophagy and Senescence in Aging Brain. Biogerontology. 2025;26(2):52.
[33] XU F, LI Z, LIU T, et al. The role of cellular senescence in the pathogenesis of Rheumatoid Arthritis: Focus on IL-6 as a target gene. Cytokine. 2024;184:156762.
[34] DALMAO-FERNÁNDEZ A, HERMIDA-GÓMEZ T, NOGUEIRA-RECALDE U, et al. Mitochondrial Role on Cellular Apoptosis, Autophagy, and Senescence during Osteoarthritis Pathogenesis. Cells. 2024;13(11):976.
[35] SILVA R. The dichotomic role of cytokines in aging. Biogerontology. 2024;26(1):17.
[36] KANDHAYA-PILLAI R, YANG X, TCHKONIA T, et al. TNF-α/IFN-γ synergy amplifies senescence-associated inflammation and SARS-CoV-2 receptor expression via hyper-activated JAK/STAT1. Aging Cell. 2022;21(6):e13646.
[37] STOLBERG-STOLBERG J, BOETTCHER A, SAMBALE M, et al. Toll-like receptor 3 activation promotes joint degeneration in osteoarthritis. Cell Death Dis. 2022; 13(3):224.
[38] QIAN Y, CHU G, ZHANG L, et al. M2 macrophage-derived exosomal miR-26b-5p regulates macrophage polarization and chondrocyte hypertrophy by targeting TLR3 and COL10A1 to alleviate osteoarthritis. J Nanobiotechnology. 2024;22(1):72.
[39] WAN Z, WANG X, FU Z, et al. Toll-like receptor activation regulates the paracrine effect of adipose-derived mesenchymal stem cells on reversing osteoarthritic phenotype of chondrocytes. Mol Biol Rep. 2024;51(1):550.
[40] ZHOU D, LUO Y, LI F, et al. Exploring the mechanisms of PANoptosis in osteoarthritis and the therapeutic potential of andrographolide through bioinformatics and single-cell analysis. Biol Direct. 2025;20(1):41.
[41] HOU Z, WANG M, CAO S. Inhibition of Toll-Like Receptor 3 Relieves Osteoarthritis by Suppression of Cartilage Degradation, Nuclear Factor Kappa B-Mediated Inflammation, and Activation of Autophagy. Cartilage. 2025: 19476035251317713. doi: 10.1177/19476035251317713.
[42] YANG Y, YUAN S, CHE M, et al. Genetic analysis of the relation between IL2RA/IL2RB and rheumatoid arthritis risk. Mol Genet Genomic Med. 2019;7(7):e00754.
[43] HU M, DENG Y, BAI Y, et al. Identifying Key Biomarkers Related to Immune Response in the Progression of Diabetic Kidney Disease: Mendelian Randomization Combined With Comprehensive Transcriptomics and Single-Cell Sequencing Analysis. J Inflamm Res. 2025;18:949-972.
[44] LIU M, WU C, WU C, et al. Immune cells differentiation in osteoarthritic cartilage damage: friends or foes? Front Immunol. 2025;16:1545284.
[45] RUYSSEN-WITRAND A, LUKAS C, NIGON D, et al. Association of IL-2RA and IL-2RB genes with erosive status in early rheumatoid arthritis patients (ESPOIR and RMP cohorts). Joint Bone Spine. 2014;81(3):228-234.
[46] ARNOLD CR, WOLF J, BRUNNER S, et al. Gain and loss of T cell subsets in old age--age-related reshaping of the T cell repertoire. J Clin Immunol. 2011;31(2):137-146.
[47] PUA HH, GUO J, KOMATSU M, et al. Autophagy is essential for mitochondrial clearance in mature T lymphocytes. J Immunol. 2009;182(7):4046-4055.
[48] NIKOLICH-ŽUGICH J. The twilight of immunity: emerging concepts in aging of the immune system. Nat Immunol. 2018;19(1):10-19.
|