Chinese Journal of Tissue Engineering Research ›› 2020, Vol. 24 ›› Issue (17): 2770-2774.doi: 10.3969/j.issn.2095-4344.2590
Previous Articles Next Articles
Biological function of circular RNA and its role in tissue repair
Shi Lei1, Sun Baolan1, Zhang Suyao1, Zhang Yuquan2
- 1Medical School of Nantong University, Nantong 226001, Jiangsu Province, China; 2Department of Obstetrics and Gynecology, Affiliated Hospital of Nantong University, Nantong 226001, Jiangsu Province, China
-
Received:2019-05-30Revised:2019-06-04Accepted:2019-07-25Online:2020-06-18Published:2020-03-30 -
Contact:Zhang Yuquan, MD, Chief physician, Department of Obstetrics and Gynecology, Affiliated Hospital of Nantong University, Nantong 226001, Jiangsu Province, China -
About author:Shi Lei, Master, Physician, Medical School of Nantong University, Nantong 226001, Jiangsu Province, China -
Supported by:the National Natural Science Foundation of China, No. 81771527
CLC Number:
Cite this article
Shi Lei, Sun Baolan, Zhang Suyao, Zhang Yuquan.
Biological function of circular RNA and its role in tissue repair [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(17): 2770-2774.
share this article
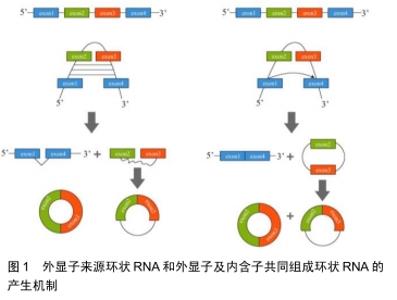
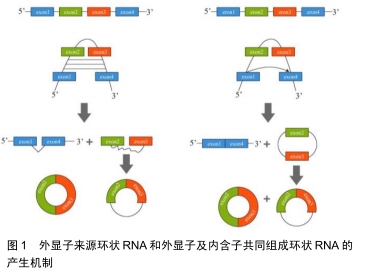
2.1 环状RNA的分类及形成机制 目前发现的环状RNA根据其来源可分为4类:外显子来源的环状RNA(exonic circRNA)、外显子及内含子共同组成的环状 RNA(exon-intron circRNA,ElciRNA)[4]、 内含子来源的环状RNA(circular intronic RNA,ciRNA)和通读环状 RNA(read-through circRNA,rt-circRNA) [2]。 2.1.1 外显子来源的环状RNA和外显子及内含子共同组成的环状RNA “内含子配对驱动的环化”和“外显子跳跃”+“套索驱动的环化”是产生这2种环状RNA的机制:①位于外显子2和外显子3侧翼的内含子之间存在互补序列,其环化成环直接通过碱基相互配对形成,最终产生外显子来源的环状RNA或外显子及内含子共同组成的环状RNA;②前体RNA发生部分折叠从而使得外显子1的3’端剪接配体与外显子4的5’端剪接受体共价结合,进而形成一个包含外显子2及外显子3的套索结构,最终环化产生外显子来源的环状RNA或外显子及内含子共同组成的环状RNA,见图1[5]。 "
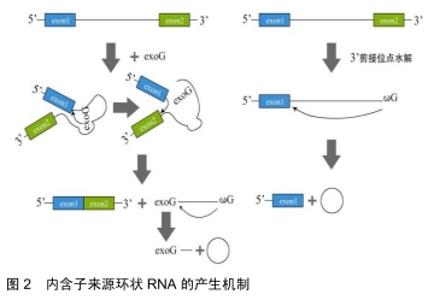
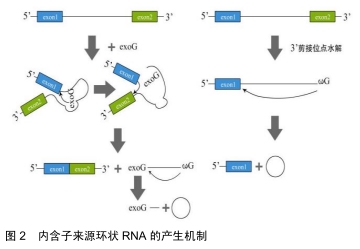
2.1.2 内含子来源的环状RNA 内含子来源的环状RNA分为Ⅰ类和Ⅱ类。Ⅰ类内含子来源的环状RNA的形成:结合在内含子上的一个外源鸟苷(exoG)攻击5’剪接位点,在第1次反应中5’外显子被切除,并且exoG与内含子变成线性连接,随后5'外显子末端的3’羟基基团攻击3’剪接位点,2个相邻的外显子相连接同时线性RNA被释放,线性内含子通过末端鸟苷(ωG)的2’-OH基团亲核攻击结合靠近3’末端的磷酸二酯键最终环化成RNA的同时释放出1个3’端尾[6]。Ⅱ类内含子来源的环状RNA的形成:环化形成需要3’端外显子的释放。内含子末端2’-OH基团攻击5’碱基位点,伴随着2’,5’磷酸二酯键形成,产生1个环状RNA和1个5’的外显子,见图2[7]。 "
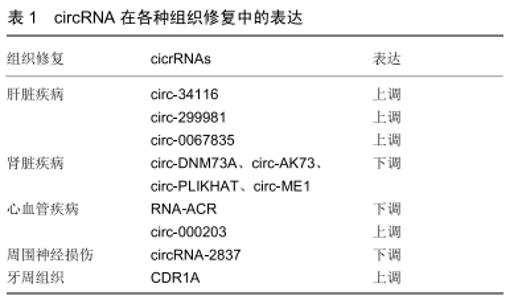
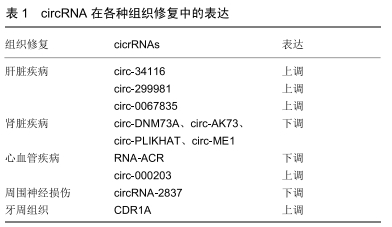
2.1.3 聚合酶Ⅱ(PolⅡ)转录通读形成的通读环状 RNA 陈玲玲课题组[8]利用4sUDRB纯化新生转录本技术,通过监测环形RNA生成与RNA聚合酶Ⅱ转录的动态偶联过程,首次证实细胞内反向剪接RNA成环与顺序剪接相比发生效率非常低,大多数RNA成环发生在转录后水平;能够发生RNA成环的基因较非成环的基因转录速度更快,这一研究结果提示外显子RNA成环效率与其所在基因的RNA聚合酶Ⅱ转录速度相偶联。 2.2 环状RNA的分子特性 2.2.1 丰富性 随着高通量测序技术的发展,环状RNA已在不同的细胞类型和物种间检测到[9]。研究发现有5.8%-23%的人类基因积极活跃转录产生环状RNA,这些环状RNA在组织和细胞间起着动态平衡作用[10-11]。MEMCZAK等[1]在CD19+、CD34+、HEK293、Neutrophil细胞系中发现2 487种环状RNA。SALZMAN[12]在人神经母细胞瘤细胞中发现4 264种环状RNA。RYBAK-WOLF等[13]在小鼠的胚胎干细胞中发现761种环状RNA。 2.2.2 稳定性 因为环状RNA的特殊结构,使其能够抵抗核糖核酸酶或核酸外切酶的降解。ZHENG等[14]对类风湿性关节炎患者和健康人血液中的单核细胞进行环状RNA表达谱分析,发现在患者和健康者中有584种显著差异的环状RNA。WU等[15]对高血压患者和健康者血浆进行了环状RNA表达谱的分析,发现有46个显著上调的环状RNA和13个显著下调的环状RNA。由于体液中含有大量的核酸酶,这些环状RNA的发现无疑证实了它的稳定性。 2.2.3 保守性 环状RNA在不同物种间也具有高度的进化保守性。在人和小鼠中有4%来自直系同源基因。在这些环状RNA中有5%-30%是完全保守的[16]。此外,在人脑中表达的环状RNA有5%-10%同样在猪脑中表达[17-18]。在大鼠和小鼠这类物种间环状RNA的保守性可高达23%[19]。哺乳动物中环状RNA的表达是相对保守的,尤其在哺乳动物的大脑组织中环状RNA保守性最高[17]。 2.3 环状RNA的主要功能 越来越多的研究表明环状RNA具有多种功能,但主要有以下4个功能。 2.3.1 作为内源竞争性RNA充当miRNA海绵 充当miRNA的海绵功能是目前对环状RNA研究比较深入的方向之一,也是环状RNA较为普遍的作用。miRNA 是一类由内源基因编码的长度约为22个核苷酸的非编码单链RNA分子,它们在动植物中参与转录后基因表达调控。miRNA可与靶基因mRNA的3'UTR相结合,起到降解或抑制mRNA翻译的作用。在宫颈癌组织中高表达的circRNA-000284可以通过吸附miR-506进而促进癌细胞增殖、迁移和侵袭[20]。JOST等[21]成功构建了人工环状RNA分子,可以吸附丙肝细胞中的miR-122,进而阻止HCV病毒蛋白形成,缓解丙型肝炎。 2.3.2 与蛋白相互作用 环状RNA与蛋白相互作用可参与多种生理活动的调控。①环状RNA与蛋白相互作用可实现对蛋白的调控。例如circ-FOXO3可与双微体2蛋白(Murine Double Minute 2,MDM2)和P53相互作用,促进MDM2介导的P53蛋白的泛素化[22];②环状RNA与蛋白的相互作用也可动态调控环状RNA的形成等过程。例如环状RNA与QKI或MBL的相互作用影响了环状RNA的形成过程。当MBI蛋白过量时,它会促进环状RNA circMBI的产生来降低自身的mRNA产量从而起到动态平衡的作用。 2.3.3 调节转录 非编码RNA可调节亲本基因的表达。研究显示内含子来源的环状RNA主要通过与RNA聚合酶Ⅱ复合体(polⅡ)结合促进基因转录。环状RNA ci-ankrd52来源于ANKRD52基因第2个内含子区域并积聚在细胞核中,ci-ankrd52干扰polⅡ的延伸并作为polⅡ的正调控因子从而起到亲本基因的顺式调节作用[2]。 2.3.4 参与蛋白翻译 LEGNINI等[23]发现环状RNA circ-ZNF609可直接翻译蛋白并参与肌肉发生过程,他们对小鼠和人成肌细胞体外分化模型及杜氏肌营养不良疾病模型进行了环状RNA表达谱分析,根据功能及表达情况,最终选择了与肌肉发生功能相关的环状RNA circ-ZNF609。基于蛋白组学分析和Western验证,证明了内源性circ-ZNF609的蛋白翻译功能。PAMUDURTI等[24]在果蝇的大脑组织中发现部分环状RNA可以结合到核糖体上并经过一系列的研究证实部分环状RNA可翻译蛋白。现在已经有越来越多的实验直接或间接证实环状RNA具有翻译的功能。 2.4 环状RNA在组织修复过程中作用的研究进展 2.4.1 肝脏疾病 ZHOU等[25]发现mmu-circ-34116在小鼠肝纤维化模型中是显著上调的,敲低mmu-circ-34116后α-平滑肌肌动蛋白(α-SMA)的表达显著增加,增加肝纤维化的进展。OU等[26]对非酒精性脂肪性肝炎的小鼠肝脏组织进行了环状谱的全面分析,研究发现共有450个环状RNA表达失调,298个环状RNA表达上调,152个环状RNA表达下调,其中circ-29981被鉴定为一种显著差异表达的环状RNA。circRNA- miRNA通路相互作用分析结果显示circRNA-29981是肝星状细胞激活的潜在调节因子,肝星状细胞是肝纤维化的主要细胞类型。ZHU等[27]发现在LX-2细胞中circ-0067835的表达显著增加,敲除circ-0067835后LX-2细胞增殖明显下降,导致G1停滞,并促进凋亡,进一步的实验发现circ-0067835通过充当mir-155的海绵来促进foxo3a的表达来调节肝纤维化的进展,circ-0067835可作为肝纤维化患者的潜在治疗靶点。 2.4.2 肾脏疾病 氯沙坦对急性肾损伤有抑制作用,但其潜在的分子机制仍不清楚。为了解氯沙坦对急性肾损伤的治疗作用,FANG等[28]通过缺血再灌注法建立大鼠急性肾损伤模型,发现大鼠缺血再灌注肾脏中存在广泛差异表达的环状RNA,circ-DNMT3A、circ-AKT3、circ-PLEKHA7和circ-ME1在急性肾损伤大鼠中表达量显著下调,而经氯沙坦预处理后的急性肾损伤大鼠中这几种环状RNA的表达量可恢复。虽然这几种环状RNA的具体作用机制不明确,但一系列研究显示这些环状RNA在急性肾损伤中起着重要的作用。K?LLING等[29]发现肾损伤患者血液中的ciRs-126表达呈现显著上调,ciRs-126可作为miR-126的海绵吸附体,而miR-126已被证实可以维持缺氧状态下内皮细胞的稳定性,故敲低ciRs-126后可上调miR-126的表达从而起到保护肾脏的作用。 2.4.3 心血管疾病 最新的研究发现环状RNA ACR通过调节PINK1/FAM65B通路抑制自噬,从而减轻心肌缺血再灌注损伤,ACR通过直接与dnmt3b结合并阻断dnmt3b介导的pink1启动子的DNA甲基化来激活pink1表达,PINK1可抑制自噬,在PINK1转基因小鼠中发现心肌梗死灶缩小。pink1可继续调控下游靶基因fam65b,而磷酸化FAM65B抑制心脏自噬和细胞死亡[30]。有研究表明在心肌纤维化的小鼠模型中circ-000203的表达显著上调,进一步研究显示上调的circ-000203可吸附miR-26b-5p,从而解除miR-26b-5p对心肌成纤维细胞中纤维化基因(如Col1a2、CTGF)的抑制作用,最终加速心肌纤维化[31]。circRNA HRCR可作为miR-223的海绵体,抑制miR-223对含胱冬肽酶富集功能域的凋亡抑制因子(ARC)的正向调控作用,来减缓心力衰竭[32]。 2.4.4 周围神经损伤 ZHOU等[33]研究发现circRNA- 2837在大鼠坐骨神经损伤模型中是显著下调的,下调circRNA-2837后可诱导体内自噬减轻坐骨神经损伤。下调circRNA-2837后检测自噬相关蛋白LC3-II和p62,发现二者呈现负相关状态,这一研究验证了其通过自噬发挥作用。CAO等[34]发现circ-013779,circ-008008,circ-003724在大鼠坐骨神经慢性损伤模型中存在显著差异,这一发现提示研究这几种环状RNA可能在坐骨神经治疗中起着重要的作用,具体机制有待进一步研究。 2.4.5 牙周组织 GU等[35]研究发现CDR1as在牙周韧带干细胞成骨分化中起着重要的作用。在成骨分化过程中CDR1A显著上调,而MIR-7显著下调。作为一种mir-7抑制剂,触发了gdf5上调和随后的smad1/5/8、p38-mapk磷酸化以促进成骨分化。该研究提供了对成骨分化机制的新理解,并提出了促进骨形成的一种潜在方法。牙周韧带干细胞在成骨培养基中培养第0,3,7,14天这4个时间点观察环状RNA表达谱的变化,研究发现在第3天有118个环状RNA表达差异,在第7天有128个环状RNA表达差异,在第14天有139个环状RNA表达差异。在胞质或膜结合囊泡和细胞外基质中,差异表达的环状RNA的宿主基因丰富,表明它们调节细胞外囊泡的生物发生具有潜在作用[36]。 2.4.6 其他 HUANG等[37]对处理和未处理的缺血再灌注小鼠小肠进行环状RNA表达谱的分析,发现circ-012412和circ-016863在未处理的缺血再灌注小鼠小肠中显著下调,而在处理的缺血再灌注小肠中显著上调。由此推测这2种环状RNA可能是防止肠道缺血再灌注损伤的保护因素。SONG等[38]对椎间盘退变中的髓核组织进行高通量测序,发现circ-104670在椎间盘退变中的髓核组织高表达。干扰circ-104670可抑制髓核细胞的凋亡,并可吸附miR-17-3p调控下游MMP-2表达。由此推断circ-104670在椎间盘退变中起关键作用,抑制椎间盘的退变。课题组用人脐带间充质干细胞修复损伤的子宫内膜间质细胞,在子宫内膜间质细胞中发现大量差异表达的环状RNA,其中circ-0111659表达显著上调,生物信息学分析发现与之结合的miR-17-5p/20b-5p/ 93-5p与VEGF均有结合位点。由此推测circ-0111659可通过miR-17-5p/20b-5p/93-5p调控VEGF的表达起到修复损伤内膜的作用[39]。以上大量研究证实circRNA在组织修复中有重要作用,见表1。 "
| [1] MEMCZAK S, JENS M, ELEFSINIOTI A, et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013;495 (7441):333-338. [2] ZHANG Y, ZHANG XO, CHEN T, et al. Circular intronic long noncoding RNAs. Mol Cell. 2013;51(6):792-806. [3] LI Z, HUANG C, BAO C, et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol. 2015;22(3):256-264. [4] JECK WR, SORRENTINO JA, WANG K, et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA. 2013; 19(2):141-157. [5] PETKOVIC S, MÜLLER S. RNA circularization strategies in vivo and in vitro. Nucleic Acids Res. 2015;43(4):2454-2465. [6] CECH TR. Self-splicing of group I introns. Annu Rev Biochem. 1990; 59:543-568. [7] LI-POOK-THAN J, BONEN L. Multiple physical forms of excised group II intron RNAs in wheat mitochondria. Nucleic Acids Res. 2006;34(9): 2782-2790. [8] ZHANG Y, XUE W, LI X, et al. The Biogenesis of Nascent Circular RNAs. Cell Rep. 2016;15(3):611-624. [9] BACHMAYR-HEYDA A, REINER AT, AUER K, et al. Correlation of circular RNA abundance with proliferation--exemplified with colorectal and ovarian cancer, idiopathic lung fibrosis, and normal human tissues. Sci Rep. 2015;5:8057. [10] CONN SJ, PILLMAN KA, TOUBIA J, et al. The RNA binding protein quaking regulates formation of circRNAs. Cell. 2015;160(6): 1125-1134. [11] KELLY S, GREENMAN C, COOK PR, et al. Exon Skipping Is Correlated with Exon Circularization. J Mol Biol. 2015;427(15): 2414-2417. [12] SALZMAN J. Circular RNA Expression: Its Potential Regulation and Function. Trends Genet. 2016;32(5):309-316. [13] RYBAK-WOLF A, STOTTMEISTER C, GLAŽAR P, et al. Circular RNAs in the Mammalian Brain Are Highly Abundant, Conserved, and Dynamically Expressed. Mol Cell. 2015;58(5):870-885. [14] ZHENG F, YU X, HUANG J, et al. Circular RNA expression profiles of peripheral blood mononuclear cells in rheumatoid arthritis patients, based on microarray chip technology. Mol Med Rep. 2017;16(6): 8029-8036. [15] WU N, JIN L, CAI J. Profiling and bioinformatics analyses reveal differential circular RNA expression in hypertensive patients. Clin Exp Hypertens. 2017;39(5):454-459. [16] GUO JU, AGARWAL V, GUO H, et al. Expanded identification and characterization of mammalian circular RNAs. Genome Biol. 2014; 15(7):409. [17] BARRETT SP, SALZMAN J. Circular RNAs: analysis, expression and potential functions. Development. 2016;143(11):1838-1847. [18] VENØ MT, HANSEN TB, VENØ ST, et al. Spatio-temporal regulation of circular RNA expression during porcine embryonic brain development. Genome Biol. 2015;16:245. [19] YOU X, VLATKOVIC I, BABIC A, et al. Neural circular RNAs are derived from synaptic genes and regulated by development and plasticity. Nat Neurosci. 2015;18(4):603-610. [20] MA HB, YAO YN, YU JJ, et al. Extensive profiling of circular RNAs and the potential regulatory role of circRNA-000284 in cell proliferation and invasion of cervical cancer via sponging miR-506. Am J Transl Res. 2018;10(2):592-604. [21] JOST I, SHALAMOVA LA, GERRESHEIM GK, et al. Functional sequestration of microRNA-122 from Hepatitis C Virus by circular RNA sponges. RNA Biol. 2018;15(8):1032-1039. [22] DU WW, FANG L, YANG W, et al. Induction of tumor apoptosis through a circular RNA enhancing Foxo3 activity. Cell Death Differ. 2017;24(2): 357-370. [23] LEGNINI I, DI TIMOTEO G, ROSSI F, et al. Circ-ZNF609 Is a Circular RNA that Can Be Translated and Functions in Myogenesis. Mol Cell. 2017;66(1):22-37. [24] PAMUDURTI NR, BARTOK O, JENS M, et al. Translation of CircRNAs. Mol Cell. 2017;66(1):9-21. [25] ZHOU Y, LV X, QU H, et al. Differential expression of circular RNAs in hepatic tissue in a model of liver fibrosis and functional analysis of their target genes. Hepatol Res. 2019;49(3):324-334. [26] OU Q, ZHAO Y, ZHOU J, et al. Comprehensive circular RNA expression profiles in a mouse model of nonalcoholic steatohepatitis. Mol Med Rep. 2019;19(4):2636-2648. [27] ZHU L, REN T, ZHU Z, et al. Thymosin-β4 Mediates Hepatic Stellate Cell Activation by Interfering with CircRNA-0067835/miR-155/FoxO3 Signaling Pathway. Cell Physiol Biochem. 2018;51(3):1389-1398. [28] FANG M, LIU S, ZHOU Y, et al. Circular RNA involved in the protective effect of losartan on ischemia and reperfusion induced acute kidney injury in rat model. Am J Transl Res. 2019;11(2):1129-1144. [29] KÖLLING M, SEEGER H, HADDAD G, et al. The Circular RNA ciRs-126 Predicts Survival in Critically Ill Patients With Acute Kidney Injury. Kidney Int Rep. 2018;3(5):1144-1152. [30] ZHOU LY, ZHAI M, HUANG Y, et al. The circular RNA ACR attenuates myocardial ischemia/reperfusion injury by suppressing autophagy via modulation of the Pink1/ FAM65B pathway. Cell Death Differ. 2019; 26(7):1299-1315. [31] TANG CM, ZHANG M, HUANG L, et al. CircRNA_000203 enhances the expression of fibrosis-associated genes by derepressing targets of miR-26b-5p, Col1a2 and CTGF, in cardiac fibroblasts. Sci Rep. 2017;7: 40342. [32] WANG K, LONG B, LIU F, et al. A circular RNA protects the heart from pathological hypertrophy and heart failure by targeting miR-223. Eur Heart J. 2016;37(33):2602-2611. [33] ZHOU ZB, NIU YL, HUANG GX, et al. Silencing of circRNA.2837 Plays a Protective Role in Sciatic Nerve Injury by Sponging the miR-34 Family via Regulating Neuronal Autophagy. Mol Ther Nucleic Acids. 2018;12:718-729. [34] CAO S, DENG W, LI Y, et al. Chronic constriction injury of sciatic nerve changes circular RNA expression in rat spinal dorsal horn. J Pain Res. 2017;10:1687-1696. [35] GU X, LI M, JIN Y, et al. Identification and integrated analysis of differentially expressed lncRNAs and circRNAs reveal the potential ceRNA networks during PDLSC osteogenic differentiation. BMC Genet. 2017;18(1):100. [36] WANG H, FENG C, JIN Y, et al. Identification and characterization of circular RNAs involved in mechanical force-induced periodontal ligament stem cells. J Cell Physiol. 2019;234(7):10166-10177. [37] HUANG JL, QIN MC, ZHOU Y, et al. Comprehensive analysis of differentially expressed profiles of Alzheimer's disease associated circular RNAs in an Alzheimer's disease mouse model. Aging (Albany NY). 2018;10(2):253-265. [38] SONG J, WANG HL, SONG KH, et al. CircularRNA_104670 plays a critical role in intervertebral disc degeneration by functioning as a ceRNA. Exp Mol Med. 2018;50(8):94. [39] SUN B, SHI L, SHI Q, et al. Circular RNAs are abundantly expressed and upregulated during repair of the damaged endometrium by Wharton's jelly-derived mesenchymal stem cells. Stem Cell Res Ther. 2018;9(1):314. |
| [1] | Geng Yao, Yin Zhiliang, Li Xingping, Xiao Dongqin, Hou Weiguang. Role of hsa-miRNA-223-3p in regulating osteogenic differentiation of human bone marrow mesenchymal stem cells [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(7): 1008-1013. |
| [2] | Liu Cong, Liu Su. Molecular mechanism of miR-17-5p regulation of hypoxia inducible factor-1α mediated adipocyte differentiation and angiogenesis [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(7): 1069-1074. |
| [3] | Ye Haimin, Ding Linghua, Kong Weihao, Huang Zutai, Xiong Long. Role and mechanism of hierarchical microchanneled bone scaffolds in promoting osteogenesis and angiogenesis [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(4): 621-625. |
| [4] | Chen Ziyang, Pu Rui, Deng Shuang, Yuan Lingyan. Regulatory effect of exosomes on exercise-mediated insulin resistance diseases [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(25): 4089-4094. |
| [5] | Yuan Changshen, Rong Weiming, Lu Zhixian, Duan Kan, Guo Jinrong, Mei Qijie. Construction of osteosarcoma miRNA-mRNA regulatory network based on bioinformatics [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(17): 2740-2746. |
| [6] | Qian Nannan, Zhang Qian, Yang Rui, Ao Jun, Zhang Tao. Mesenchymal stem cells in the treatment of spinal cord injury: cell therapy and combination of new drugs and biomaterials [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(13): 2114-2120. |
| [7] | Liu Tao, Zhang Nini, Huang Guilin . Relationship between extracellular vesicles and radiation-induced tissue injury [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(13): 2121-2126. |
| [8] | Deng Junhao, Li Miao, Zhang Licheng, Tang Peifu. Three-dimensional hanging-drop culture of mesenchymal stem cells in the treatment of tissue injury [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(7): 1101-1106. |
| [9] | Yuan Yiming, Wang Yan, Chen Chengcheng, Zhao Mingyue, Pei Fei. Efficacy of exosomes in peripheral nerve injury [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(31): 5079-5084. |
| [10] | Lu Shuqing, Li Xin, Guo Jin, Liu Shiyu, Yang Shunbo, Feng Yuxia, Pang Wei. Advantages in instrument-assisted soft tissue mobilization for chronic soft tissue injury [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(29): 4709-4716. |
| [11] | Lai Yu, Han Jie. Mechanism and progress of exosomes in the treatment of nonunion [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(27): 4349-4355. |
| [12] | Hu Zhongling, Li Binbin, Cui Yishuang, Wang Qian, Zhang Hui, Li Qijia, Wang Zhiqiang. Concept, mechanism and problem of gene therapy for osteoarthritis [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(27): 4356-4363. |
| [13] |
Wang Zhenggang, Liu Mingjiang, Huang Xinfeng, Xiao Xiangjun.
Prefabricated composite tissue flaps for repairing bone and soft tissue defects of the extremities [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(26): 4208-4212. |
| [14] |
Zeng Zhaomu, Wen Xichao, Zhang Yuhao, Geng Lianting, Shen Yang, Zheng Kebin.
Role and application of exosomes-mediated miRNAs in the treatment of glioma [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(25): 4073-4080. |
| [15] | Hu Baoyang, Yang Xuejun. Research progress and development space of microRNA in the process of intervertebral disc degeneration [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(21): 3372-3378. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||