Chinese Journal of Tissue Engineering Research ›› 2023, Vol. 27 ›› Issue (29): 4696-4703.doi: 10.12307/2023.480
Previous Articles Next Articles
Mechanism underlying the mutual regulation between circadian rhythm and autophagy
Chen Chao, Wang Xuenan, Zhan Enyu, Lyu Zhengpin, Zhang Fan
- Department of Orthopedics, The First Affiliated Hospital of Kunming Medical University, Kunming 650032, Yunnan Province, China
-
Received:2022-05-23Accepted:2022-08-13Online:2023-10-18Published:2022-12-02 -
Contact:Zhang Fan, MD, Associate professor, Department of Orthopedics, The First Affiliated Hospital of Kunming Medical University, Kunming 650032, Yunnan Province, China -
About author:Chen Chao, Master candidate, Department of Orthopedics, The First Affiliated Hospital of Kunming Medical University, Kunming 650032, Yunnan Province, China -
Supported by:the National Natural Science Foundation of China, No. 82160428 (to ZF); the Major Science and Technology Special Project of Yunnan Provincial Science and Technology Department, No. 202102AA310068 (to ZF); a grant from Yunnan Provincial Clinical Center for Osteoarthritis, No. ZX2019-03-04 (to ZF)
CLC Number:
Cite this article
Chen Chao, Wang Xuenan, Zhan Enyu, Lyu Zhengpin, Zhang Fan. Mechanism underlying the mutual regulation between circadian rhythm and autophagy[J]. Chinese Journal of Tissue Engineering Research, 2023, 27(29): 4696-4703.
share this article
Add to citation manager EndNote|Reference Manager|ProCite|BibTeX|RefWorks
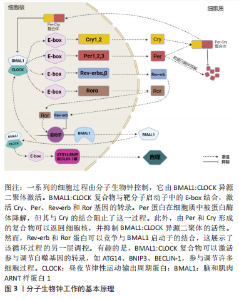
2.1 昼夜节律中的分子生物钟概述 分子生物钟分为2种:一种位于中枢神经系统中,由接受光刺激后的视交叉上核神经元调控;另一种存在于周围组织中,通过时钟基因的表达控制组织细胞,该机制也称细胞自主分子生物钟。中央时钟主要由视网膜感知,并通过与地球物理时间同步的电信号传递给视交叉上核神经元。而视交叉上核通过局部振荡器、神经元和体液信号的同步来影响昼夜节律[26]。在哺乳动物中,细胞自主分子生物钟能够产生稳定且健全的24 h基因表达节律,与两个互锁的转录/翻译反馈回路共同发挥作用是密不可分的。该核心回路——转录/翻译反馈回路与不同的环境和代谢刺激相结合,以产生细胞内部24 h昼夜节律。核心转录/翻译反馈回路由4种完整的时钟蛋白驱动,包括两种激活蛋白、两种阻遏蛋白(Per和Cry),以及调节上述整合时钟蛋白定位和稳定性的激酶和磷酸酶(激酶:CKIα、CKIδ和CKIε;磷酸酶:PP1、PP5)[27]。最近的研究发现,分子生物钟对于昼夜节律的产生至关重要[21],其基本原理依赖于转录激活因子诱导抑制因子的转录(图3),导致后者随着时间的推移积累,直到达到足以抑制自身激活的水平[28] 。基本的螺旋-环-螺旋PER-ARNT-SIM(bHLH-PAS)蛋白CLOCK和BMAL1是哺乳动物生物钟机制中的主要转录激活因子。CLOCK和BMAL1转录因子作为异二聚体,通过与E-box调控序列结合去激活Per(Per1、Per2和Per3)和Cry(Cry1和Cry2)基因家族[29]。有趣的是,Per蛋白在细胞质中被蛋白酶体降解,而Per:Cry异二聚体可以转移到细胞核,一方面防止Per蛋白降解,另一方面抑制和阻断BMAL1:CLOCK 复合物的活性[30]。在转录环的平行臂中,BMAL1:CLOCK异二聚体激活孤儿核受体REV-ERB(REV-ERBα和REV-ERBβ)和ROR(RORα、RORβ和RORγ)的转录[31]。REV-ERBs和RORs可以竞争结合BMAL1启动子,分别激活和抑制基因转录效应,从而导致BMAL1的昼夜节律振荡[19]。除此之外,BMAL1:CLOCK还激活了参与调节自噬的基因的转录,例如ATG14、BNIP3、BECLIN-1等[8]。"
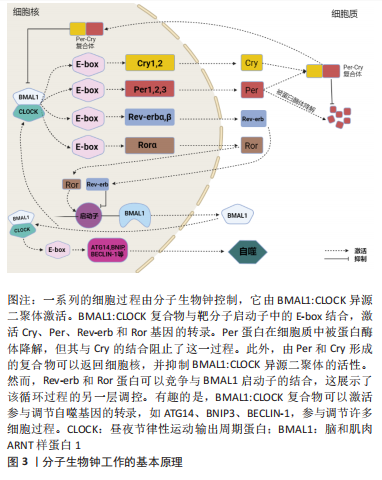
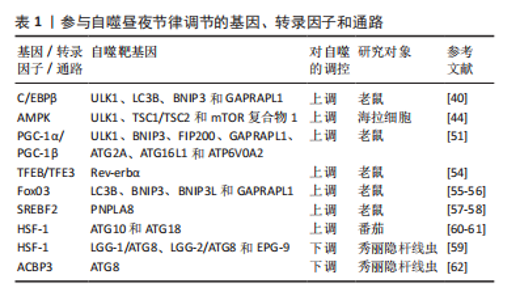
2.2 自噬的昼夜节律 1970年代初,PFEIFER[32]使用电子显微镜首次发现自噬与昼夜节律调节之间相关联的证据,与营养缺乏的对照组相比,膳食喂养大鼠的自噬泡和肝脏萎缩遵循昼夜规律。然而,随着研究的深入,有学者通过电子显微镜、自噬通量测量和荧光测量,进一步发现昼夜节律在肾脏、心脏、肝脏、大脑和视网膜中调节自噬的其他证据,其不仅存在于哺乳动物中,而且在果蝇和斑马鱼中也有所体现[30,33-35]。自噬节律因组织而异:首先,在肾小管中,单位面积自噬泡数量和分离物质总量显示出明显的昼夜节律,其最小值出现在夜间,而最大值出现在白天[36];其次,心脏中自噬泡的体积和数量密度在午夜达到峰值,随后在黑暗早期下降[30];最后,在小鼠肝脏中,自噬通量在下午达到峰值,在夜间迅速下降,并在整个光照阶段再次上升[37],因此肝脏自噬的日常节律可能并不直接来自分子昼夜节律振荡器,而是通过进食或其他正常的昼夜节律门控行为间接与分子时钟耦合[35]。总的来说,上述研究证实了外周调节的昼夜节律依赖性自噬在多个器官系统中被操纵,并且可能在组织和器官修复中发挥关键作用,同时为睡眠时间对日常生理过程和组织维持至关重要的理论提供了依据。 2.3 昼夜节律与自噬的相互调控机制 2.3.1 参与昼夜节律调控自噬的相关基因、转录因子和通路 在许多组织中,自噬活动和自噬泡的数量在白天发生变化,且一些参与自噬的基因、转录因子和通路在昼夜变化中具有节律性[38-78],见表1。"
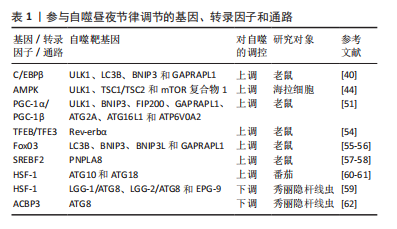
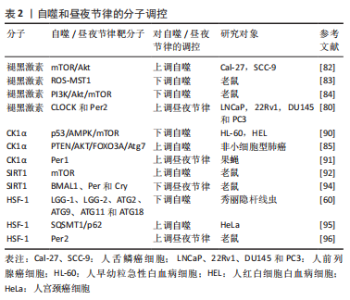
(1) CCAAT/增强子结合蛋白β:CCAAT/增强子结合蛋白β(CCAAT/enhancer binding proteinβ,C/EBPβ)是一种基本的亮氨酸拉链转录因子,在饥饿和光/暗周期中对自噬的激活至关重要[38],其可直接与自噬基因的启动子结合并诱导其转录[39]。目前,C/EBPβ正在成为参与节律性自噬基因表达的重要因子,如ULK1、LC3B、BNIP3和GABARAPL1,这些蛋白以振荡方式表达,并由昼夜节律和营养信号介导[40]。MA等[37]研究发现腺病毒介导的RNAi敲除C/EBPβ消除了昼夜节律调节的自噬,进一步的研究表明,在肝脏特异性BMAL1敲除小鼠中,C/EBPβ水平改变,同时自噬的节律性调节中断、自噬基因的表达降低。 (2)腺苷-磷酸活化蛋白激酶通路:腺苷单磷酸活化蛋白激酶(adenosine monophosphate-activated protein kinase,AMPK)通路是在功能上连接自噬和昼夜节律的桥梁。AMPK是一种众所周知的能量感应激酶,参与各种分解代谢和合成代谢过程,包括氧化代谢、葡萄糖摄取和营养生物合成[41]。此外,自噬受到mTOR信号通路和下游Atgs级联反应的负调控,如BECLIN-1、LC3、ATG1和ATG5[42]。最近的研究表明,AMPK可通过双重机制诱导自噬,包括通过激活结节性硬化基因1/结节性硬化基因2(TSC1/TSC2)复合物,直接激活和消除mTOR复合物1对ULK1的抑制作用[43-44]。有趣的是,AMPK和mTOR均表现出节律性调节。LAMIA等[45]发现AMPK的活性和核定位,呈现出周期性的节律,且与小鼠肝细胞中Cry1的核表达呈负相关。此外,AMPK的激活使Cry蛋白和正常的节律性振荡失去平衡,AMPK通路的破坏明显损害了小鼠的外周生物钟活性[46]。因此,AMPK基于其通过调节Cry1控制mTOR循环表达的能力,为自噬的节律性调控提供了方向。 (3)过氧化物酶体增殖物激活受体γ共激活因子1α(the peroxisome proliferator–activated receptor gamma coactivator 1α,PGC-1α)和PGC-1β:PGC-1转录共激活因子家族包括 PGC-1α和PGC-1β[47]。PGC-1α和PGC-1β可以激活许多细胞类型中由核基因组和线粒体基因组编码的线粒体基因程序的表达[48]。此外,研究证实PGC-1α和PGC-1β的表达在肝脏和骨骼肌中呈现出强烈的昼夜节律。对于PGC-1α,与昼夜节律基因表达的相关性表明,PGC-1α以昼夜节律方式调节线粒体的活动,同时缺乏PGC-1α的小鼠表现出异常的昼夜节律[49-50],因此,PGC-1α与昼夜节律之间的相互作用是昼夜节律与自噬之间的联系之一。然而,对于PGC-1β,在瞬时转染的293T细胞中发现其与C/EBPβ物理连接,表明其可能是C/EBPβ转录共激活因子[51]。进一步的研究发现,腺病毒介导的PGC-1β过表达可促进肝细胞中C/EBPβ调节的自噬基因的表达,包括ULK1、BNIP3、FIP200、GABARAPL1、ATG2A、ATG16L1和ATPase H+ 转运V0亚基a2(ATP6V0A2)[51]。上述结果表明,PGC-1β可能作为C/EBPβ的共激活因子,从而促进自噬的表达,增强自噬活性。因此,PGC-1β和C/EBPβ的节律性激活通过调节昼夜节律信号从而节律性地激活自噬。 (4)转录因子EB(transcription factor EB,TFEB)和TFE3:TFEB和TFE3通过激活参与这些过程的许多基因的表达,是自噬、溶酶体生物发生和溶酶体胞吐作用的主要调节因子[52-53]。研究发现,TFEB和TFE3以昼夜节律方式被激活并促进Rev-erbα(Nr1d1)的表达,Rev-erbα是控制自噬相关基因(BECLIN-1、BNIP3、ATG5、ATG7和ULK1)表达的主要时钟机制的转录抑制剂成分,然而,它们的消耗损害了Rev-erbα表达和节律性振荡[54]。 此外,TFEB/TFE3和Rev-erbα结合共同的启动子区域,表明它们可以诱导参与自噬的基因的节律性表达[54]。因此,上述结果为利用TFEB/TFE3和Rev-erbα之间的动态平衡来调控自噬的节律性振荡提供了一种新的机制。 (5) FoxO3、SREBF2、HSF-1及ACBP3:除此之外,其他一些转录因子包括FoxO3、SREBF2、HSF-1和ACBP3,也参与了自噬的昼夜节律调控。首先,FoxO3是一种自噬调节因子,已被证明可以诱导自噬基因的表达,包括LC3B、BNIP3、BCL2相互作用蛋白3like(BNIP3L)和GABARAPL1[55-56]。其次,SREBF2可以通过以昼夜节律方式直接激活与自噬体相关的磷脂酶patatin样域包含蛋白8(PNPLA8)基因的表达,从而增加高脂饮食喂养小鼠肝细胞的自噬[57-58]。接着,热激因子HSF-1表现出昼夜节律,且在一定的细胞条件下,通过作用于不同的自噬相关基因来诱导和抑制自噬,例如ATG10和ATG18[59-61]。最后,ACBP3的过表达增强了ATG8的降解,并阻碍了自噬小体的形成[62],同时ACBP3 的昼夜节律调节在黑暗中上调,但在光照下被抑制,是由顺式响应元件与单指盒(Dof)和GT-1的DNA结合介导的[63]。 2.3.2 昼夜节律生物钟核心成分调控自噬 昼夜节律与行为或环境之间的整合对正常的生理过程至关重要。目前的研究发现进食/禁食节律可调节肌肉、脂肪和肝脏组织中的自噬,并在新陈代谢和寿命中发挥保护作用[64]。研究发现,自噬相关基因和mTOR也显示出明显的昼夜节律[65],受多个昼夜节律生物钟核心成分的控制,特别是Per2、BMAL1和REV-ERB。 (1) Per2:研究发现肝脏中的Per2是一种支架蛋白,可将结节性硬化症复合物1、Raptor和mTOR相结合,从而特异性抑制mTORC1复合体的活性;因此Per2缺乏由于失去对mTORC1的抑制作用,会显著增强蛋白质合成和细胞增殖,但会导致自噬水平降低[66]。关于Per调控自噬的相关过程,一方面,KALFALAH等[67]指出小鼠成纤维细胞中Per2的敲除会下调自噬水平,而核心时钟振荡基本保持不变,同时Per2的敲除还降低了Ulk1蛋白的水平,但不影响Ulk1 mRNA水平,这与小鼠成纤维细胞中有节律的Ulk1蛋白水平和非节律的Ulk1 mRNA水平保持一致。另一方面,Per2的瞬时过表达导致体外PI3K/Akt通路的下调,并增加自噬通量[67-68]。同时在昼夜条件下,Tor、ATG5和ATG7在野生型苍蝇的大脑中表现出节律性的表达,而其在Per1时钟突变体中被消除[34]。进一步的研究表明,在Per1b突变体鱼中,其自噬相关基因LC3B、Gabarapa、ATPase H+转运溶酶体V1亚基D(Atp6v1d)、Atg4a、Atg4d、Beclin1、Bnip3、Ulk1a和Ulk1b的表达均上调,但无节律性[69]。上述研究表明,Per在维持自噬的节律性方面起着不可或缺的作用。 (2) BMAL1:研究表明,在心脏中BMAL1通过mTORC1信号下调诱导自噬,在高血糖条件下保护心肌细胞[70]。SCOTTON等[71] 研究发现BMAL1敲除小鼠肌肉中自噬标记基因LC3A水平升高,p62水平降低,而Beclin-1水平没有变化,且AMPK的激活已被证明参与自噬过程中BMAL1的调控。同时在心肌细胞特异性BMAL1敲除和心肌细胞特异性时钟突变小鼠表现出PI3K /Akt/mTOR 信号轴的过度激活,这可能有助于自噬的减弱和蛋白质合成的增强[72]。进一步的研究发现,C/EBPβ是一种有效的自噬激活剂,在肝脏中有节奏地表达,并受昼夜节律和营养信号的调节,可诱导自噬相关基因如Ulk1、Gabarapl1、LC3B、Bnip3的表达和蛋白质的降解[37],但在BMAL1缺陷小鼠肝脏中,C/EBPβ mRNA振荡的幅度显著降低,且Bnip3和Ulk1 mRNA表达的昼夜节律几乎完全消失,导致C/EBPβ和自噬的昼夜节律调节被破坏,小鼠肝脏中的峰值自噬通量显着降低[37]。因此,BMAL1的表达在自噬的激活过程中起着重要作用。 (3) REV-ERB:REV-ERB是一种与血红素结合的生物钟成分,可作为肿瘤发生过程的抑制因子,包括代谢、增殖和炎症过程。体外研究发现,REV-ERB激动剂阻断自噬并诱导细胞凋亡,同时在体内,REV-ERB激动剂被证明可以下调自噬相关基因Ulk3、Ulk1、Beclin1和Atg7的mRNA和蛋白水平,通过抑制自噬过程显著损害体内胶质母细胞瘤的生长并提高存活率[31]。此外,ZHANG等[73]指出在斑马鱼中,REV-ERBα被发现可直接与Ulk1启动子中的ROR反应元件(RORE)位点结合,从而在体内抑制Ulk1a,而另一种自噬相关基因Atp6v1d是REV-ERBα的直接靶点,进一步调控自噬。然而,有学者认为REV-ERBβ似乎并不直接参与自噬调控,而是可能作为自噬阻断下游的细胞保护因子[74],对于目前存在的争论还需进一步研究。 除此之外,其他生物钟基因也具有影响自噬的能力。研究发现,AMPK活性和核定位具有节律性,与小鼠肝脏中Cry1核蛋白丰度呈反比[45]。而昼夜节律调节器CLOCK可通过诱导自噬从而影响肿瘤细胞对顺铂的耐药性[75],同时自噬相关基因Atg14的表达受CLOCK/BMAL1与其E-box结合进行控制,也表现出明显的昼夜节律[76]。 2.3.3 自噬调控昼夜节律 昼夜节律可以调节自噬,并受自噬的调节。目前,与昼夜节律调节自噬相比,自噬调节昼夜节律的证据相对较少。KIJAK等[34]发现过表达细胞中Tor的沉默缩短了果蝇运动活动节律的周期,表明TOR信号通路和自噬参与调节成年果蝇大脑中神经元行为和可塑性的昼夜节律。同时AMPK的刺激使时钟蛋白Cry不稳定并改变昼夜节律,被基因破坏的AMPK通路小鼠表现为外周时钟发生改变[45]。有报道指出,昼夜节律蛋白BMAL1、CLOCK、REV-ERBα和Cry1是溶酶体靶标,在这之中自噬通过选择性降解Cry1来影响昼夜节律,当啮齿动物依赖糖异生时,Cry1作为一种糖异生抑制剂,其选择性自噬降解发生昼夜窗口中,这表明Cry1降解与维持血糖有关,是维持血糖的时间印记。同时,高脂喂养会加速自噬Cry1的降解,并导致肥胖相关的高血糖症[77]。此外在高脂喂养下,CLOCK突变体通过蛋白酶体和自噬途径减弱BMAL1的降解,这为核心时钟成分的自噬调节提供了证据[78]。 2.4 自噬和昼夜节律的分子调控 目前,研究发现,有几种分子可以调节自噬和昼夜节律[79-96],在节律性自噬的调节中起着至关重要的作用,见表2。"
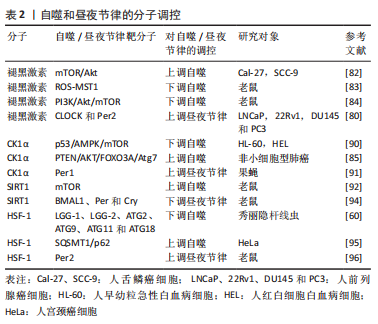

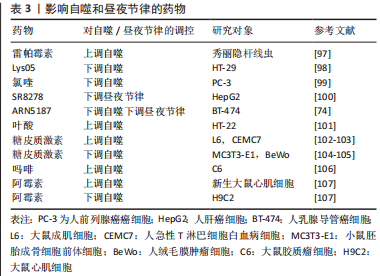
2.4.1 褪黑激素 褪黑激素是一种受生物钟调节的神经激素,由大脑松果体合成和分泌,在自噬调节、控制生物钟相关基因和睡眠调节中起着重要作用。褪黑激素有助于维持昼夜节律,通过增强或抑制自噬过程去增强不同系统的保护作用,包括中枢神经、心血管、胃肠道和内分泌系统等[79]。研究发现,褪黑激素可以上调人前列腺癌细胞中CLOCK和Per2蛋白的表达,从而影响昼夜节律[80]。此外,褪黑激素作为促自噬指示剂或抗自噬调节剂,在自噬中发挥关键作用。有学者指出,自噬可被氧化应激激活,同时褪黑激素具有抗氧化特性,可以抵消线粒体损伤、氧化应激和细胞凋亡,因此褪黑激素对自噬的抑制作用可通过抗氧化活性解释[81],可推论褪黑激素可能影响了自噬的发生机制,而不是影响其自噬过程本身。同时在某些疾病中,例如癌症、神经退行性变和肥胖症,褪黑激素可通过调节ROS/MST1和PI3K/Akt/mTOR信号通路在自噬中发挥双重作用[82-84]。 2.4.2 CK1蛋白家族的酪蛋白激酶1α CK1蛋白家族的酪蛋白激酶1α(CK1α)调节与膜运输、细胞周期进程、染色体分离、细胞凋亡、自噬、细胞代谢和发育分化、昼夜节律和免疫反应以及神经变性和癌症相关的信号通路[85],在自噬调节中表现出双重功能[86-87]。一方面,通过D4476(一种CK1抑制剂)抑制CK1α,或siRNA介导CK1α敲低导致mTOR信号通路的抑制和自噬的激活[88-89],同时CK1α可抑制急性髓系白血病介导MDM2介导自噬下游的p53[90]。另一方面,CK1α的过表达可通过PTEN/AKT/FOXO3A/ATG7轴显著激活非小细胞肺癌(NSCLC)中的自噬通量[85]。此外,CK1α介导的磷酸化促进Per1的降解,表明其是维持24 h昼夜节律所必需的,但目前其调控机制尚不清楚[91]。 2.4.3 沉默信息调节因子2相关酶1(silent mating type information regulation 2 homolog 1,SIRT1)和热休克转录因子1(heat shock transcription 1,HSF1) SIRT1是一种核蛋白,在细胞中占据特殊位置并控制关键的代谢和生理过程,属于sirtuin家族,可以通过阻断mTOR或激活AMPK来促进自噬[92]。此外,SIRT1作为中央时钟组件的去乙酰化酶[93],可以使组蛋白BMAL1、Per和Cry去乙酰化,通过调节昼夜节律蛋白的转录,有助于体内的昼夜节律控制[94]。综上,这些研究结果表明,SIRT1在自噬和昼夜节律之间、氧化还原状态和生物钟之间的联系中起着重要作用。HSF1在自噬中表现出双重功能,一方面,HSF1应激反应通路通过调节自噬受体SQSTM1/p62 相关蛋白稳态,从而下调自噬过程[95];另一方面,通过作用于不同自噬相关基因如LGG-1、LGG-2、ATG2、ATG9、ATG11和ATG18等来抑制自噬[60]。此外,HSF1还可以通过直接调节Per2蛋白来诱导生物钟同步[96]。 2.5 影响自噬和昼夜节律的药物 研究发现,除了某些影响节律性自噬的分子外,一些药物也能发挥类似的作用。例如,已在临床实践中使用的雷帕霉素是一种有效的自噬激活剂[97],而氯喹和Lys05是抑制自噬的溶酶体药物[98-99]。KOJETIN等[100]发现了第一个REV-ERB拮抗剂化合物SR8278,但其药代动力学特性限制了其药理用途。与此同时,最近的研究发现,化合物ARN5187对BT-474乳腺癌细胞中的自噬和REV-ERB具有双重抑制作用[74],一方面,ARN5187被证明是一种自噬抑制因子,其具有阻断溶酶体功能、破坏晚期自噬过程和降低乳腺癌细胞活力的能力;另一方面,ARN5187还显示出抑制REV-ERB介导的转录调控。此外,有学者表明叶酸缺乏可以增强HT-22海马神经元细胞体外自噬的活性,同时通过糖皮质激素受体介导的途径诱导自噬和昼夜节律相关基因的表达,如ATG12、ATG13和Per2等[101],该研究表明叶酸可能参与节律性自噬的调节。然而,糖皮质激素在不同组织或细胞中对自噬的调控存在差异。首先,糖皮质激素通过抑制IP(3)依赖性钙信号来促进自噬[102],同时诱导的自噬可通过线粒体清除介导肌肉萎缩[103]。其次,在胎盘和骨骼中,糖皮质激素可降低其自噬活性[104-105]。然而,吗啡可增加大鼠胶质瘤C6细胞中自噬相关基因(包括ATG3、ATG5、ATG7 和 ATG12)的mRNA表达水平[106],同时在阿霉素诱导的心脏毒性过程中可以调节自噬[107]。见表3。"
| [1] YAN Y, CHEN X, WANG X, et al. The effects and the mechanisms of autophagy on the cancer-associated fibroblasts in cancer. J Exp Clin Cancer Res. 2019;38(1):171. [2] MARTEL J, OJCIUS DM, KO YF, et al. Phytochemicals as Prebiotics and Biological Stress Inducers. Trends Biochem Sci. 2020;45(6):462-471. [3] GOU Q, JIANG Y, ZHANG R, et al. PPARδ is a regulator of autophagy by its phosphorylation. Oncogene. 2020;39(25):4844-4853. [4] LI F, ZHANG C, TANG Z, et al. A plant RNA virus activates selective autophagy in a UPR-dependent manner to promote virus infection. New Phytol. 2020;228(2):622-639. [5] PATRA S, MISHRA SR, BEHERA BP, et al. Autophagy-modulating phytochemicals in cancer therapeutics: Current evidences and future perspectives. Semin Cancer Biol. 2022;80:205-217. [6] BONAM SR, BAYRY J, TSCHAN MP, et al. Progress and Challenges in The Use of MAP1LC3 as a Legitimate Marker for Measuring Dynamic Autophagy In Vivo. Cells. 2020;9(5):1321. [7] WU X, NAGY LE. MLKL contributes to Western diet-induced liver injury through inhibiting autophagy. Autophagy. 2020;16(7):1351-1352. [8] WANG X, XU Z, CAI Y, et al. Rheostatic Balance of Circadian Rhythm and Autophagy in Metabolism and Disease. Front Cell Dev Biol. 2020;8: 616434. [9] WONG SQ, KUMAR AV, MILLS J, et al. Autophagy in aging and longevity. Hum Genet. 2020;139(3):277-290. [10] LEVY J MM, TOWERS CG, THORBURN A. Targeting autophagy in cancer. Nat Rev Cancer. 2017;17(9):528-542. [11] PAVLINOV I, SALKOVSKI M, ALDRICH LN. Beclin 1-ATG14L Protein-Protein Interaction Inhibitor Selectively Inhibits Autophagy through Disruption of VPS34 Complex I. J Am Chem Soc. 2020;142(18):8174-8182. [12] VAHSEN BF, RIBAS VT, SUNDERMEYER J, et al. Inhibition of the autophagic protein ULK1 attenuates axonal degeneration in vitro and in vivo, enhances translation, and modulates splicing. Cell Death Differ. 2020;27(10):2810-2827. [13] PETHERICK KJ, CONWAY OJ, MPAMHANGA C, et al. Pharmacological inhibition of ULK1 kinase blocks mammalian target of rapamycin (mTOR)- dependent autophagy. J Biol Chem. 2015;290(18):11376-11383. [14] FRACCHIOLLA D, CHANG C, HURLEY JH, et al. A PI3K-WIPI2 positive feedback loop allosterically activates LC3 lipidation in autophagy. J Cell Biol. 2020;219(7):e201912098. [15] BOUKHALFA A, NASCIMBENI AC, RAMEL D, et al. PI3KC2α-dependent and VPS34-independent generation of PI3P controls primary cilium-mediated autophagy in response to shear stress. Nat Commun. 2020; 11(1):294. [16] WU MY, LIU L, WANG EJ, et al. PI3KC3 complex subunit NRBF2 is required for apoptotic cell clearance to restrict intestinal inflammation. Autophagy. 2021;17(5):1096-1111. [17] LYSTAD AH, CARLSSON SR, SIMONSEN A. Toward the function of mammalian ATG12-ATG5-ATG16L1 complex in autophagy and related processes. Autophagy. 2019;15(8):1485-1486. [18] GALLUZZI L, BRAVO-SAN PEDRO JM, LEVINE B, et al. Pharmacological modulation of autophagy: therapeutic potential and persisting obstacles. Nat Rev Drug Discov. 2017;16(7):487-511. [19] MAIESE K. Moving to the Rhythm with Clock (Circadian) Genes, Autophagy, mTOR, and SIRT1 in Degenerative Disease and Cancer. Curr Neurovasc Res. 2017;14(3):299-304. [20] KIM H, WILLIAMS D, QIU Y, et al. Regulation of hepatic autophagy by stress-sensing transcription factor CREBH. FASEB J. 2019;33(7): 7896-7914. [21] KIM YH, LAZAR MA. Transcriptional Control of Circadian Rhythms and Metabolism: A Matter of Time and Space. Endocr Rev. 2020;41(5): 707-732. [22] RYZHIKOV M, EUBANKS A, HASPEL JA. Measuring Diurnal Rhythms in Autophagic and Proteasomal Flux. J Vis Exp. 2019;(151):60133. [23] TAN X, YE J, LIU W, et al. Acrylamide aggravates cognitive deficits at night period via the gut-brain axis by reprogramming the brain circadian clock. Arch Toxicol. 2019;93(2):467-486. [24] PAUL S, HANNA L, HARDING C, et al. Output from VIP cells of the mammalian central clock regulates daily physiological rhythms. Nat Commun. 2020;11(1):1453. [25] DREYER AP, MARTIN MM, FULGHAM CV, et al. A circadian output center controlling feeding:fasting rhythms in Drosophila. PLoS Genet. 2019;15(11):e1008478. [26] DIBNER C, SCHIBLER U, ALBRECHT U. The mammalian circadian timing system: organization and coordination of central and peripheral clocks. Annu Rev Physiol. 2010;72:517-549. [27] PARTCH CL, GREEN CB, TAKAHASHI JS. Molecular architecture of the mammalian circadian clock. Trends Cell Biol. 2014;24(2):90-99. [28] REDDY AB, REY G. Metabolic and nontranscriptional circadian clocks: eukaryotes. Annu Rev Biochem. 2014;83:165-189. [29] GABRYELSKA A, SOCHAL M, TURKIEWICZ S, et al. Relationship between HIF-1 and Circadian Clock Proteins in Obstructive Sleep Apnea Patients-Preliminary Study. J Clin Med. 2020;9(5):1599. [30] RABINOVICH-NIKITIN I, LIEBERMAN B, MARTINO TA, et al. Circadian-Regulated Cell Death in Cardiovascular Diseases. Circulation. 2019; 139(7):965-980. [31] SULLI G, ROMMEL A, WANG X, et al. Pharmacological activation of REV-ERBs is lethal in cancer and oncogene-induced senescence. Nature. 2018;553(7688):351-355. [32] PFEIFER U. Cellular autophagy and cell atrophy in the rat liver during long-term starvation. A quantitative morphological study with regard to diurnal variations. Virchows Arch B Cell Pathol. 1973;12(3):195-211. [33] YAO J, JIA L, SHELBY SJ, et al. Circadian and noncircadian modulation of autophagy in photoreceptors and retinal pigment epithelium. Invest Ophthalmol Vis Sci. 2014;55(5):3237-3246. [34] KIJAK E, PYZA E. TOR signaling pathway and autophagy are involved in the regulation of circadian rhythms in behavior and plasticity of L2 interneurons in the brain of Drosophila melanogaster. PLoS One. 2017; 12(2):e0171848. [35] RYZHIKOV M, EHLERS A, STEINBERG D, et al. Diurnal Rhythms Spatially and Temporally Organize Autophagy. Cell Rep. 2019;26(7):1880-1892.e1886. [36] PFEIFER U, SCHELLER H. A morphometric study of cellular autophagy including diurnal variations in kidney tubules of normal rats. J Cell Biol. 1975;64(3):608-621. [37] MA D, PANDA S, LIN JD. Temporal orchestration of circadian autophagy rhythm by C/EBPβ. EMBO J. 2011;30(22):4642-4651. [38] XU X, HUANG E, LUO B, et al. Methamphetamine exposure triggers apoptosis and autophagy in neuronal cells by activating the C/EBPβ-related signaling pathway. FASEB J. 2018:fj201701460RRR. [39] BARAKAT DJ, MENDONCA J, BARBERI T, et al. C/EBPβ regulates sensitivity to bortezomib in prostate cancer cells by inducing REDD1 and autophagosome-lysosome fusion. Cancer Lett. 2016;375(1): 152-161. [40] MA D, LIN JD. Circadian regulation of autophagy rhythm through transcription factor C/EBPβ. Autophagy. 2012;8(1):124-125. [41] GONZáLEZ A, HALL MN, LIN SC, et al. AMPK and TOR: The Yin and Yang of Cellular Nutrient Sensing and Growth Control. Cell Metab. 2020;31(3):472-492. [42] MIZUSHIMA N, LEVINE B, CUERVO AM, et al. Autophagy fights disease through cellular self-digestion. Nature. 2008;451(7182):1069-1075. [43] KIM J, KUNDU M, VIOLLET B, et al. AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat Cell Biol. 2011;13(2): 132-141. [44] SHANG L, CHEN S, DU F, et al. Nutrient starvation elicits an acute autophagic response mediated by Ulk1 dephosphorylation and its subsequent dissociation from AMPK. Proc Natl Acad Sci U S A. 2011; 108(12):4788-4793. [45] LAMIA KA, SACHDEVA UM, DITACCHIO L, et al. AMPK regulates the circadian clock by cryptochrome phosphorylation and degradation. Science. 2009;326(5951):437-440. [46] LEE Y, KIM EK. AMP-activated protein kinase as a key molecular link between metabolism and clockwork. Exp Mol Med. 2013;45(7):e33. [47] PICCININ E, PERES C, BELLAFANTE E, et al. Hepatic peroxisome proliferator-activated receptor γ coactivator 1β drives mitochondrial and anabolic signatures that contribute to hepatocellular carcinoma progression in mice. Hepatology. 2018;67(3):884-898. [48] NIERENBERG AA, GHAZNAVI SA, SANDE MATHIAS I, et al. Peroxisome Proliferator-Activated Receptor Gamma Coactivator-1 Alpha as a Novel Target for Bipolar Disorder and Other Neuropsychiatric Disorders. Biol Psychiatry. 2018;83(9):761-769. [49] SONODA J, MEHL IR, CHONG LW, et al. PGC-1beta controls mitochondrial metabolism to modulate circadian activity, adaptive thermogenesis, and hepatic steatosis. Proc Natl Acad Sci U S A. 2007; 104(12):5223-5228. [50] LIU C, LI S, LIU T, et al. Transcriptional coactivator PGC-1alpha integrates the mammalian clock and energy metabolism. Nature. 2007; 447(7143):477-481. [51] LI S, LIN JD. Transcriptional control of circadian metabolic rhythms in the liver. Diabetes Obes Metab. 2015;17 Suppl 1(1):33-38. [52] ANNUNZIATA I, VAN DE VLEKKERT D, WOLF E, et al. MYC competes with MiT/TFE in regulating lysosomal biogenesis and autophagy through an epigenetic rheostat. Nat Commun. 2019;10(1):3623. [53] YANG C, ZHU Z, TONG BC, et al. A stress response p38 MAP kinase inhibitor SB202190 promoted TFEB/TFE3-dependent autophagy and lysosomal biogenesis independent of p38. Redox Biol. 2020;32:101445. [54] PASTORE N, VAINSHTEIN A, HERZ NJ, et al. Nutrient-sensitive transcription factors TFEB and TFE3 couple autophagy and metabolism to the peripheral clock. EMBO J. 2019;38(12):e101347. [55] RUAN C, WANG C, GONG X, et al. An integrative multi-omics approach uncovers the regulatory role of CDK7 and CDK4 in autophagy activation induced by silica nanoparticles. Autophagy. 2021;17(6):1426-1447. [56] MAMMUCARI C, MILAN G, ROMANELLO V, et al. FoxO3 controls autophagy in skeletal muscle in vivo. Cell Metab. 2007;6(6):458-471. [57] OISHI K, YAMAMOTO S, ITOH N, et al. Wheat alkylresorcinols suppress high-fat, high-sucrose diet-induced obesity and glucose intolerance by increasing insulin sensitivity and cholesterol excretion in male mice. J Nutr. 2015;145(2):199-206. [58] KIM KY, JANG HJ, YANG YR, et al. SREBP-2/PNPLA8 axis improves non-alcoholic fatty liver disease through activation of autophagy. Sci Rep. 2016;6:35732. [59] BARNA J, CSERMELY P, VELLAI T. Roles of heat shock factor 1 beyond the heat shock response. Cell Mol Life Sci. 2018;75(16):2897-2916. [60] LI J, CHAUVE L, PHELPS G, et al. E2F coregulates an essential HSF developmental program that is distinct from the heat-shock response. Genes Dev. 2016;30(18):2062-2075. [61] WANG Y, CAI S, YIN L, et al. Tomato HsfA1a plays a critical role in plant drought tolerance by activating ATG genes and inducing autophagy. Autophagy. 2015;11(11):2033-2047. [62] XIAO S, CHYE ML. The Arabidopsis thaliana ACBP3 regulates leaf senescence by modulating phospholipid metabolism and ATG8 stability. Autophagy. 2010;6(6):802-804. [63] ZHENG SX, XIAO S, CHYE ML. The gene encoding Arabidopsis acyl-CoA-binding protein 3 is pathogen inducible and subject to circadian regulation. J Exp Bot. 2012;63(8):2985-3000. [64] MARTINEZ-LOPEZ N, TARABRA E, TOLEDO M, et al. System-wide Benefits of Intermeal Fasting by Autophagy. Cell Metab. 2017;26(6): 856-871.e855. [65] ROTTER D, ROTHERMEL BA. Targets, trafficking, and timing of cardiac autophagy. Pharmacol Res. 2012;66(6):494-504. [66] WU R, DANG F, LI P, et al. The Circadian Protein Period2 Suppresses mTORC1 Activity via Recruiting Tsc1 to mTORC1 Complex. Cell Metab. 2019;29(3):653-667.e656. [67] KALFALAH F, JANKE L, SCHIAVI A, et al. Crosstalk of clock gene expression and autophagy in aging. Aging (Albany NY). 2016;8(9):1876-1895. [68] WANG Z, LI L, WANG Y. Effects of Per2 overexpression on growth inhibition and metastasis, and on MTA1, nm23-H1 and the autophagy-associated PI3K/PKB signaling pathway in nude mice xenograft models of ovarian cancer. Mol Med Rep. 2016;13(6):4561-4568. [69] HUANG G, ZHANG F, YE Q, et al. The circadian clock regulates autophagy directly through the nuclear hormone receptor Nr1d1/Rev-erbα and indirectly via Cebpb/(C/ebpβ) in zebrafish. Autophagy. 2016; 12(8):1292-1309. [70] QIAO L, GUO B, ZHANG H, et al. The clock gene, brain and muscle Arnt-like 1, regulates autophagy in high glucose-induced cardiomyocyte injury. Oncotarget. 2017;8(46):80612-80624. [71] SCOTTON C, BOVOLENTA M, SCHWARTZ E, et al. Deep RNA profiling identified CLOCK and molecular clock genes as pathophysiological signatures in collagen VI myopathy. J Cell Sci. 2016;129(8):1671-1684. [72] MCGINNIS GR, TANG Y, BREWER RA, et al. Genetic disruption of the cardiomyocyte circadian clock differentially influences insulin-mediated processes in the heart. J Mol Cell Cardiol. 2017;110:80-95. [73] ZHANG TW, LI ZF, DONG J, et al. The circadian rhythm in intervertebral disc degeneration: an autophagy connection. Exp Mol Med. 2020; 52(1):31-40. [74] DE MEI C, ERCOLANI L, PARODI C, et al. Dual inhibition of REV-ERBβ and autophagy as a novel pharmacological approach to induce cytotoxicity in cancer cells. Oncogene. 2015;34(20):2597-2608. [75] SUN Y, JIN L, SUI YX, et al. Circadian Gene CLOCK Affects Drug-Resistant Gene Expression and Cell Proliferation in Ovarian Cancer SKOV3/DDP Cell Lines Through Autophagy. Cancer Biother Radiopharm. 2017; 32(4):139-146. [76] XIONG X, TAO R, DEPINHO RA, et al. The autophagy-related gene 14 (Atg14) is regulated by forkhead box O transcription factors and circadian rhythms and plays a critical role in hepatic autophagy and lipid metabolism. J Biol Chem. 2012;287(46):39107-39114. [77] TOLEDO M, BATISTA-GONZALEZ A, MERHEB E, et al. Autophagy Regulates the Liver Clock and Glucose Metabolism by Degrading CRY1. Cell Metab. 2018;28(2):268-281.e264. [78] JEONG K, HE B, NOHARA K, et al. Dual attenuation of proteasomal and autophagic BMAL1 degradation in Clock Δ19/+ mice contributes to improved glucose homeostasis. Sci Rep. 2015;5:12801. [79] ROOHBAKHSH A, SHAMSIZADEH A, HAYES AW, et al. Melatonin as an endogenous regulator of diseases: The role of autophagy. Pharmacol Res. 2018;133:265-276. [80] JUNG-HYNES B, HUANG W, REITER RJ, et al. Melatonin resynchronizes dysregulated circadian rhythm circuitry in human prostate cancer cells. J Pineal Res. 2010;49(1):60-68. [81] HOSSAIN MF, WANG N, CHEN R, et al. Exploring the multifunctional role of melatonin in regulating autophagy and sleep to mitigate Alzheimer’s disease neuropathology. Ageing Res Rev. 2021;67:101304. [82] SHEN YQ, GUERRA-LIBRERO A, FERNANDEZ-GIL BI, et al. Combination of melatonin and rapamycin for head and neck cancer therapy: Suppression of AKT/mTOR pathway activation, and activation of mitophagy and apoptosis via mitochondrial function regulation. J Pineal Res. 2018;64(3):12461. [83] SHI L, LIANG F, ZHENG J, et al. Melatonin Regulates Apoptosis and Autophagy Via ROS-MST1 Pathway in Subarachnoid Hemorrhage. Front Mol Neurosci. 2018;11:93. [84] ZHENG Y, HOU J, LIU J, et al. Inhibition of autophagy contributes to melatonin-mediated neuroprotection against transient focal cerebral ischemia in rats. J Pharmacol Sci. 2014;124(3):354-364. [85] JIANG S, ZHANG M, SUN J, et al. Casein kinase 1α: biological mechanisms and theranostic potential. Cell Commun Signal. 2018; 16(1):23. [86] CAI J, LI R, XU X, et al. CK1α suppresses lung tumour growth by stabilizing PTEN and inducing autophagy. Nat Cell Biol. 2018;20(4): 465-478. [87] HERMANOVA I, CARRACEDO A. CK1α promotes tumour suppressive autophagy. Nat Cell Biol. 2018;20(4):369-371. [88] GAO D, INUZUKA H, TAN MK, et al. mTOR drives its own activation via SCF(βTrCP)-dependent degradation of the mTOR inhibitor DEPTOR. Mol Cell. 2011;44(2):290-303. [89] ZHAO Y, XIONG X, SUN Y. DEPTOR, an mTOR inhibitor, is a physiological substrate of SCF(βTrCP) E3 ubiquitin ligase and regulates survival and autophagy. Mol Cell. 2011;44(2):304-316. [90] XU W, HUANG Z, GAN Y, et al. Casein kinase 1α inhibits p53 downstream of MDM2‑mediated autophagy and apoptosis in acute myeloid leukemia. Oncol Rep. 2020;44(5):1895-1904. [91] LAM VH, LI YH, LIU X, et al. CK1α Collaborates with DOUBLETIME to Regulate PERIOD Function in the Drosophila Circadian Clock. J Neurosci. 2018;38(50):10631-10643. [92] WU Y, LI W, HU Y, et al. Suppression of sirtuin 1 alleviates airway inflammation through mTOR‑mediated autophagy. Mol Med Rep. 2020; 22(3):2219-2226. [93] BELDEN WJ, DUNLAP JC. SIRT1 is a circadian deacetylase for core clock components. Cell. 2008;134(2):212-214. [94] ASHER G, GATFIELD D, STRATMANN M, et al. SIRT1 regulates circadian clock gene expression through PER2 deacetylation. Cell. 2008;134(2): 317-328. [95] WATANABE Y, TSUJIMURA A, TAGUCHI K, et al. HSF1 stress response pathway regulates autophagy receptor SQSTM1/p62-associated proteostasis. Autophagy. 2017;13(1):133-148. [96] TAMARU T, HATTORI M, HONDA K, et al. Synchronization of circadian Per2 rhythms and HSF1-BMAL1:CLOCK interaction in mouse fibroblasts after short-term heat shock pulse. PLoS One. 2011;6(9):e24521. [97] PULAKAT L, CHEN HH. Pro-Senescence and Anti-Senescence Mechanisms of Cardiovascular Aging: Cardiac MicroRNA Regulation of Longevity Drug-Induced Autophagy. Front Pharmacol. 2020;11:774. [98] BAQUERO P, DAWSON A, MUKHOPADHYAY A, et al. Targeting quiescent leukemic stem cells using second generation autophagy inhibitors. Leukemia. 2019;33(4):981-994. [99] ERKISA M, AYDINLIK S, CEVATEMRE B, et al. A promising therapeutic combination for metastatic prostate cancer: Chloroquine as autophagy inhibitor and palladium(II) barbiturate complex. Biochimie. 2020;175: 159-172. [100] KOJETIN D, WANG Y, KAMENECKA TM, et al. Identification of SR8278, a synthetic antagonist of the nuclear heme receptor REV-ERB. ACS Chem Biol. 2011;6(2):131-134. [101] SUN Q, YANG Y, LI X, et al. Folate deprivation modulates the expression of autophagy- and circadian-related genes in HT-22 hippocampal neuron cells through GR-mediated pathway. Steroids. 2016;112:12-19. [102] HARR MW, MCCOLL KS, ZHONG F, et al. Glucocorticoids downregulate Fyn and inhibit IP(3)-mediated calcium signaling to promote autophagy in T lymphocytes. Autophagy. 2010;6(7):912-921. [103] TRONCOSO R, PAREDES F, PARRA V, et al. Dexamethasone-induced autophagy mediates muscle atrophy through mitochondrial clearance. Cell Cycle. 2014;13(14):2281-2295. [104] HE B, ZHANG N, ZHAO R. Dexamethasone Downregulates SLC7A5 Expression and Promotes Cell Cycle Arrest, Autophagy and Apoptosis in BeWo Cells. J Cell Physiol. 2016;231(1):233-242. [105] LIN NY, CHEN CW, KAGWIRIA R, et al. Inactivation of autophagy ameliorates glucocorticoid-induced and ovariectomy-induced bone loss. Ann Rheum Dis. 2016;75(6):1203-1210. [106] FENG YM, JIA YF, SU LY, et al. Decreased mitochondrial DNA copy number in the hippocampus and peripheral blood during opiate addiction is mediated by autophagy and can be salvaged by melatonin. Autophagy. 2013;9(9):1395-1406. [107] SISHI BJ, LOOS B, VAN ROOYEN J, et al. Autophagy upregulation promotes survival and attenuates doxorubicin-induced cardiotoxicity. Biochem Pharmacol. 2013;85(1):124-134. |
| [1] | Guo Shuhui, Yang Ye, Jiang Yangyang, Xu Jianwen. Screening and validation of neurogenic bladder miRNA-mRNA regulatory network [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(在线): 1-8. |
| [2] | Cao Sheng, Kong Lingwei, Xu Kun, Sun Zhijie. Correlation of cervical sagittal force line parameters with degenerative segment and Pfirrmann classification in patients with cervical intervertebral disc degeneration [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(9): 1319-1324. |
| [3] | Tang Liang, Li Xiheng, Niu Ruijuan, Li Xinyue, Zou Xinying, Mao Tianjiao, Li Jiang. Naringin regulates the function of RAW264.7 macrophages to affect the osteogenic differentiation of MC-3T3-E1 cells [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(8): 1205-1210. |
| [4] | Zhang Yongqiang, Pan Feng, Sun Peng, Tan Minghui, Xuan Liuming, Wang Qinzhang. Effect of Gja1 gene recombinant lentivirus on Cx43 protein and mRNA expression in a diabetic guinea pig bladder model [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(8): 1161-1165. |
| [5] | Lian Shilin, Zhang Yan, Jiang Qiang, Zhang Hanshuo, Li Tusheng, Ding Yu. Interventional effects of whole blood and platelet-rich plasma with different preparation methods on nucleus pulposus cells [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(8): 1199-1204. |
| [6] | Yang Zhishan, Tang Zhenglong. YAP/TAZ, a core factor of the Hippo signaling pathway, is involved in bone formation [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(8): 1264-1271. |
| [7] | Song Hehua, Wei Zairong. Diabetic peripheral neuropathy: research and therapy [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(8): 1278-1285. |
| [8] | Nie Chenchen, Su Kaiqi, Gao Jing, Fan Yongfu, Ruan Xiaodi, Yuan Jie, Duan Zhaoyuan, Feng Xiaodong. The regulatory role of circular RNAs in cerebral ischemia-reperfusion injury [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(8): 1286-1291. |
| [9] | Xu Xingxing, Wen Chaoju, Meng Maohua, Wang Qinying, Chen Jingqiao, Dong Qiang. Carbon nanomaterials in oral implant [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(7): 1062-1070. |
| [10] | Tang Haotian, Liao Rongdong, Tian Jing. Application and design of piezoelectric materials for bone defect repair [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(7): 1117-1125. |
| [11] | Xu Yan, Li Ping, Lai Chunhua, Zhu Peijun, Yang Shuo, Xu Shulan. Piezoelectric materials for vascularized bone regeneration [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(7): 1126-1132. |
| [12] | Zhang Qijian, Xu Ximing. Acquisition and application of ectodermal mesenchymal stem cells [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(6): 928-934. |
| [13] | Yuan Bo, Xie Lide, Fu Xiumei. Schwann cell-derived exosomes promote the repair and regeneration of injured peripheral nerves [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(6): 935-940. |
| [14] | Zhou Yinan, Yang Kun, Jiang Jiayi, Zhang Sizhuo, Zhu Yingwen. The relationship between glucose transporter 1/4 gene polymorphisms and athletic ability [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(5): 745-750. |
| [15] | Li Long, Li Guangdi, Shi Hao, Deng Keqi. Circular RNA as a competing endogenous RNA is involved in the regulation of osteoarthritis [J]. Chinese Journal of Tissue Engineering Research, 2023, 27(5): 751-757. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||