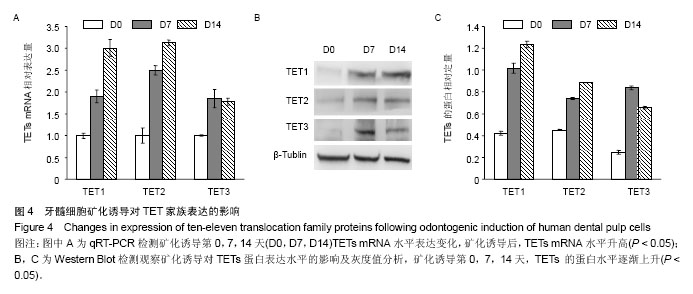
| [1] Jones PA, Liang G. Rethinking how DNA methylation patterns are maintained. Nat Rev Genet. 2009;10(11):805-811.
[2] Fuks F, Hurd P J, Wolf D, et al. The methyl-CpG-binding protein MeCP2 links DNA methylation to histone methylation. J Biol Chem. 2003;278(6):4035-4040.
[3] Wu SC, Zhang Y. Active DNA demethylation: many roads lead to Rome. Nat Rev Mol Cell Biol. 2010;11(9):607-620.
[4] Tahiliani M, Koh KP, Shen Y, et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science. 2009;324(5929):930-935.
[5] Ito S, D'Alessio AC, Taranova OV, et al. Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature. 2010;466(7310):1129-1133.
[6] Chen J, Gao Y, Huang H, et al. The combination of tet1 with oct4 generates high-quality mouse-induced pluripotent stem cells. Stem Cells. 2015;33(3):686-698.
[7] Lin LL, Wang W, Hu Z, et al. Erratum to: Negative feedback of miR-29 family TET1 involves in hepatocellular cancer. Med Oncol. 2015;32(3):39.
[8] Hsu CH, Peng KL, Kang ML, et al. TET1 suppresses cancer invasion by activating the tissue inhibitors of metalloproteinases. Cell Rep. 2012;2(3):568-579.
[9] Langemeijer SM, Kuiper RP, Berends M, et al. Acquired mutations in TET2 are common in myelodysplastic syndromes. Nat Genet. 2009;41(7):838-842.
[10] Kunimoto H, Fukuchi Y, Sakurai M, et al. Tet2 disruption leads to enhanced self-renewal and altered differentiation of fetal liver hematopoietic stem cells. Sci Rep. 2012;2:273.
[11] Ko M, Huang Y, Jankowska AM, et al. Impaired hydroxylation of 5-methylcytosine in myeloid cancers with mutant TET2. Nature. 2010;468(7325):839-843.
[12] Ko M, Bandukwala HS, An J, et al. Ten-Eleven-Translocation 2 (TET2) negatively regulates homeostasis and differentiation of hematopoietic stem cells in mice. Proc Natl Acad Sci U S A. 2011;108(35):14566-14571.
[13] Li T, Yang D, Li J, et al. Critical role of Tet3 in neural progenitor cell maintenance and terminal differentiation. Mol Neurobiol. 2015;51(1):142-154.
[14] Gronthos S, Mankani M, Brahim J, et al. Postnatal human dental pulp stem cells (DPSCs) in vitro and in vivo. Proc Natl Acad Sci U S A. 2000;97(25):13625-13630.
[15] Xu Y, Wu F, Tan L, et al. Genome-wide regulation of 5hmC, 5mC, and gene expression by Tet1 hydroxylase in mouse embryonic stem cells. Mol Cell. 2011;42(4):451-464.
[16] Yu J, He H, Tang C, et al. Differentiation potential of STRO-1+ dental pulp stem cells changes during cell passaging. BMC Cell Biol. 2010;11:32.
[17] Rawluszko-Wieczorek AA, Siera A, Horbacka K, et al. Clinical significance of DNA methylation mRNA levels of TET family members in colorectal cancer. J Cancer Res Clin Oncol. 2015.
[18] Li Q, Rao L, Zhang D, et al. Expression features of DNA methylcytosine dioxygenase ten-eleven translocation 1 in human dental pulp cells. J Endod. 2014;40(11):1791-1795.
[19] Guo X, Wang Y, Guo Z, et al. TET Proteins: A New Family of DNA Modifying Enzymes. Zhongguo Shengwu Huaxue yu Fenzi Shengwu Xuebao. 2011;27(12):1101-1106.
[20] Langemeijer SM, Kuiper RP, Berends M, et al. Acquired mutations in TET2 are common in myelodysplastic syndromes. Nat Genet. 2009;41(7):838-842.
[21] Song J, Rechkoblit O, Bestor T H, et al. Structure of DNMT1-DNA complex reveals a role for autoinhibition in maintenance DNA methylation. Science. 2011;331(6020): 1036-1040.
[22] Dunican DS, Pennings S, Meehan RR. The CXXC-TET bridge-mind the methylation gap! Cell Res. 2013;23(8): 973-974.
[23] Ko M, An J, Bandukwala HS, et al. Modulation of TET2 expression and 5-methylcytosine oxidation by the CXXC domain protein IDAX. Nature. 2013;497(7447):122-126.
[24] Arioka Y, Watanabe A, Saito K, et al. Activation-induced cytidine deaminase alters the subcellular localization of Tet family proteins. PLoS One. 2012;7(9):e45031.
[25] Wu H, D'Alessio AC, Ito S, et al. Dual functions of Tet1 in transcriptional regulation in mouse embryonic stem cells. Nature. 2011;473(7347):389-393.
[26] Williams K, Christensen J, Pedersen MT, et al. TET1 and hydroxymethylcytosine in transcription and DNA methylation fidelity. Nature. 2011;473(7347):343-348.
[27] Ficz G, Branco MR, Seisenberger S, et al. Dynamic regulation of 5-hydroxymethylcytosine in mouse ES cells and during differentiation. Nature. 2011;473(7347):398-402.
[28] Zhang R R, Cui Q Y, Murai K, et al. Tet1 regulates adult hippocampal neurogenesis and cognition. Cell Stem Cell. 2013;13(2):237-245.
[29] Ko M, Bandukwala H S, An J, et al. Tet2 Negatively Regulates Homeostasis and Differentiation of Hematopoietic Stem Cells in Mice. Blood. 2011;118(21):1017.
[30] Li T, Yang D, Li J, et al. Critical Role of Tet3 in Neural Progenitor Cell Maintenance and Terminal Differentiation. Mol Neurobiol. 2014.
[31] David L, Hirsch CL. How Tets and cytoskeleton dynamics MET in reprogramming. Cell Stem Cell. 2014;14(4):417-418.
[32] Hu X, Zhang L, Mao SQ, et al. Tet and TDG mediate DNA demethylation essential for mesenchymal-to-epithelial transition in somatic cell reprogramming. Cell Stem Cell. 2014;14(4):512-522.
[33] Zhang RR, Cui QY, Murai K, et al. Tet1 regulates adult hippocampal neurogenesis and cognition. Cell Stem Cell. 2013;13(2):237-245.
[34] Wang P, Qu J, Wu MZ, et al. "TET-on" pluripotency. Cell Res. 2013;23(7):863-865.
[35] Wang T, Wu H, Li Y, et al. Subtelomeric hotspots of aberrant 5-hydroxymethylcytosine-mediated epigenetic modifications during reprogramming to pluripotency. Nat Cell Biol. 2013; 15(6): 700-711.
[36] Spruijt CG, Gnerlich F, Smits AH, et al. Dynamic readers for 5-(hydroxy)methylcytosine and its oxidized derivatives. Cell. 2013;152(5):1146-1159.
[37] Huang Y, Chavez L, Chang X, et al. Distinct roles of the methylcytosine oxidases Tet1 and Tet2 in mouse embryonic stem cells. Proc Natl Acad Sci U S A. 2014;111(4):1361-1366. |