Chinese Journal of Tissue Engineering Research ›› 2022, Vol. 26 ›› Issue (2): 276-282.doi: 10.12307/2022.045
Previous Articles Next Articles
The pivotal role of autophagy in bone cells: bone-related cell activity and bone metabolism
Sun Youqiang1, Ma Chao2, Liang Mengmeng3, Xin Pengfei2, Zhang Hua1, Xiang Xiaobing1, *
- 1Department of Sports Medicine, The First Affiliated Hospital of Guangzhou University of Chinese Medicine, Guangzhou 510405, Guangdong Province, China; 2The First Clinical College of Guangzhou University of Chinese Medicine, Guangzhou 510405, Guangdong Province, China; 3Department of Obstetrics, Guangdong Women and Children Hospital, Guangzhou 510010, Guangdong Province, China
-
Received:2020-10-20Revised:2020-10-28Accepted:2020-11-28Online:2022-01-18Published:2021-10-27 -
Contact:Xiang Xiaobing, MD, Chief physician, Department of Sports Medicine, The First Affiliated Hospital of Guangzhou University of Chinese Medicine, Guangzhou 510405, Guangdong Province, China -
About author:Sun Youqiang, MD, Attending physician, Department of Sports Medicine, The First Affiliated Hospital of Guangzhou University of Chinese Medicine, Guangzhou 510405, Guangdong Province, China -
Supported by:a grant from Guangdong Provincial Administration of Traditional Chinese Medicine, No. 20201112 (to SYQ); Youth Innovation Fund of the First Affiliated Hospital of Guangzhou University of Chinese Medicine, No. 2019QN12 (to SYQ)
CLC Number:
Cite this article
Sun Youqiang, Ma Chao, Liang Mengmeng, Xin Pengfei, Zhang Hua, Xiang Xiaobing. The pivotal role of autophagy in bone cells: bone-related cell activity and bone metabolism[J]. Chinese Journal of Tissue Engineering Research, 2022, 26(2): 276-282.
share this article
Add to citation manager EndNote|Reference Manager|ProCite|BibTeX|RefWorks
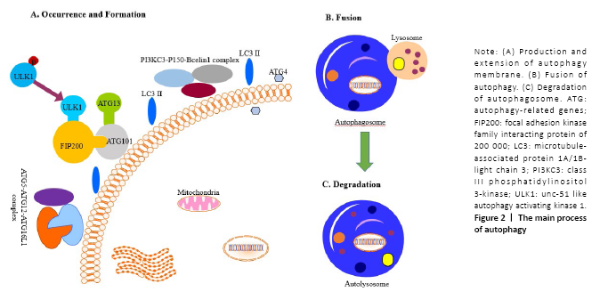
Autophagy and its formation Autophagy first observed by Ashford and Porter[8] under an electron microscope in 1962 is an essential cellular homeostatic mechanism that maintains homeostasis and stress sensitivity by removing dysfunctional organelles or macromolecules. There are various types of autophagy, including microautophagy, macroautophagy and chaperone-mediated autophagy. Each of them remains its unique mechanisms and functions[9,10]. Both microautophagy and macroautophagy have the capacity to engulf large structures through both selective and non-selective mechanisms, whereas chaperone-mediated autophagy degrades only soluble proteins in a selective manner. In microautophagy, lysosomes directly engulf the surrounding cytoplasm by invading or protruding their membranes. The maintenance of organelle cell, membrane homeostasis, and cell survival under nitrogen restriction are the main functions of macroautophagy[11], while macroautophagy will form double-layered membrane autophagosomes to wrap damaged cell components. Chaperone-mediated autophagy selectively transfers substrates to lysosomal membranes by binding to pentapeptides similar to the KFERQ sequence[12]. Macroautophagy is the main type of autophagy and has been identified to be regulated by more than 40 autophagy-related genes (ATG). Chaperone proteins will selectively bind to the target protein and directly enter the lysosome to complete the cycle. The autophagy can be divided into the following four stages[13,14] (Figure 2)."
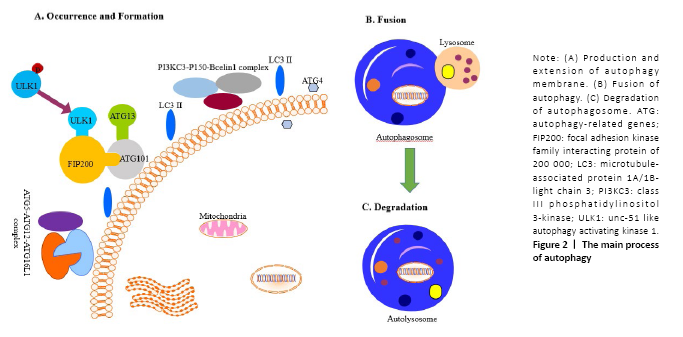
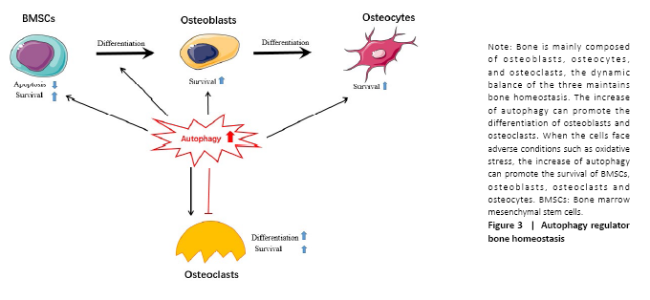
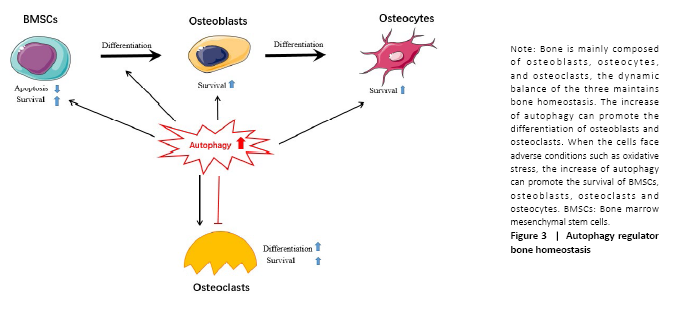
I. The occurrence of autophagy membrane: The cells accept the autophagy-induced signal and transfer it to the mammalian rapamycin target protein (mTOR); after inducing the dephosphorylation of the unc-51 like autophagy activating kinase 1 (ULK1), it promotes ATG13, ATG101, FIP200 and ULK1 to form the ATG13- ATG101-ULK1-FIP200 complex thus to induce the extension of the autophagy membrane[15]. II. The formation of autophagosomes: The extension of the autophagy membrane is mainly regulated by two ubiquitin-like systems, class III phosphatidylinositol 3-kinase (PI3KC3), p150, and Beclin1 combine to form a complex, and then induce the autophagy membrane continues to extend. At the same time, ATG7 activates ATG12, and is successively combined with ATG10 and ATG5, and then interacts with ATG16L1 and ATG5 to form the ATG5-ATG12-ATG16L1 poly-complex, together to promote the growth of autophagosomes[16,17]. Subsequently, microtubule-associated protein 1A/1B-light chain 3 (LC3) was processed into LC3-I by ATG4[18]; meanwhile, LC3-I combined with phosphatidylethanolamine to form LC3-II, thereby promoting the formation of autophagosomes. The latter is the second ubiquitin-like system. III. The transport of autophagosomes: After autophagy membrane extension and expansion is completed, autophagosome and lysosome fused to the formation of autolysosomes, This process may involve related SNARE proteins such as syntaxin 17 on the outer membrane of autophagy[19,20]. IV. The degradation of autophagosomes: The mTOR pathway is the key to the process of autophagy. After the lysosome degrades the substance in the cytoplasm, the degraded product is transported out of the lysosome for reuse by the cells. The release of lysosomal nutrients will activate the mTOR signaling pathway, forming a negative feedback mechanism to avoid excessive cytoplasmic degradation. Activation will cause the lysosomes to extend out of the tubules. These tubules are often called “protolysosomes” and can continue to develop into mature lysosomes[21]. On one hand, the basal levels of autophagy are important for maintaining normal cellular homeostasis. On the other hand, even though the capacity for large-scale degradation is important for the function of autophagy, but it carries a certain risk because unregulated degradation of the cytoplasm is likely to be lethal[22]. Autophagy regulates bone homeostasis Autophagy is of great significance to bone homeostasis. Bone is a dynamic organ, the activities and functions of osteocytes, osteoclasts and osteoblasts will directly affect the changes in bone mass and bone strength. Osteoblasts have mainly synthesized bone matrix while osteoclasts have a bone resorption effect, under normal circumstances, bone resorption and bone formation maintain a relative balance. Once the balance is broken, it will lead to osteoporosis, osteonecrosis or Paget’s disease, etc.[23] (Figure 3). When the bone cells were in the condition of nutritional deficiency, hypoxia or infection and other adverse stimuli, a higher level of autophagy would be induced, to maintain the homeostasis of the bone system and prolong life. Therefore, a comprehensive understanding of the role of autophagy in bone and the mechanisms of autophagy in bone-related diseases will benefit for the exploration of the treatment therapies. Autophagy and BMSCs BMSCs have the potential for self-renewal and multidirectional differentiation including adipose, bone, cartilage, and muscle[24], The osteoblast precursor cells required for bone tissue development, bone metabolism, and bone repair are all differentiated from BMSCs. Current data demonstrate that the autophagy activity of BMSCs affects their differentiation process, which in turn affects bone mass and even secondary bone. Autophagy regulates BMSC regeneration and controls osteoporosis development[25]. Older BMSCs are more likely to differentiate into adipocytes than to osteoblasts, and their autophagy activity is reduced, the use of autophagy inhibitor 3-methyladenine can reduce the differentiation and proliferation of BMSCs into osteoblasts while activator rapamycin reverses this process[26, 27]. Recently, it was shown that primary human BMSCs exhibit a high level of constitutive autophagy, with decreased BMSCs’ differentiation into osteoblasts[28]. In agreement with this observation, scientists have found that there is a high level of green fluorescent protein LC3 (GFP-LC3) puncta in primary BMSCs isolated from GFP-LC3 transgenic mice, to the authors’ surprise; these GFP-LC3 puncta disappear after these cells are differentiated into osteoblast-like cells[29]. These suggest that the level of autophagy in BMSCs is higher in osteoblasts. However, the role of the constitutive autophagy in BMSCs remains unclear. Researchers have also found that when cells are hypoxic, peripheral reactive oxygen species is produced, high glucose cause apoptosis, and MSCs can produce exosomes through multiple pathways such as AMPK/mTOR, Akt/mTOR to enhance and enhance autophagy, and protect BMSC survival[30-32]. Induced mitochondria[33], vitamin C transporter 2[34] and insulin-like growth factor 1[35] may play key roles in BMSCs’ antioxidative stress-related autophagy and apoptosis. Another discovery is that there are massive undegraded autophagic vesicles in undifferentiated mesenchymal stem cells, and the number of aggregated autophagic vesicles is reduced during the differentiation process of mesenchymal stem cells into osteoblasts[36]. This reminds us that the autophagic vesicles can supply the energy for the promotion of the differentiation of BMSCs. Autophagy is inseparable from BMSC; however, the detailed biological functions of autophagy in BMSC such as maintenance, self-renewal, and differentiation are largely unknown. The detailed cellular and molecular mechanisms for the regulation of BMSCs during differentiation process demand a future investigation. Autophagy and osteoblasts Osteoblasts are mesenchymal-derived cells responsible for synthesis and secretion of bone matrix, and its subsequent mineralization[6]. The development and growth of bone are closely related to osteoblasts. Osteoblasts activate osteogenesis by secreting a large number of bone collagen matrix and some important cytokines, and through this series of factors coupled with the regulation of osteoclast to control the osteoclastogenesis maturation and activation[6]. In the process of bone formation, a portion of osteoblasts are differentiated into osteocytes and the other part is apoptotic[37]. Existing research supports the important role of autophagy in the differentiation and mineralization of osteoblasts. Studies have reported that there is a significantly increased autophagy in the early differentiation and the mineralization of the skull osteoblast, indicating that the autophagy and the differentiation and mineralization of osteoblast are closely related[38]. Liu et al.[39] found autophagic vesicles contain apatite-like structure in BMSCs after he observed the transmission process under electron microscopy, indicating that autophagic vesicles may be a mean of transportation during bone mineralization, autophagic vacuoles could be used as vehicles in osteoblasts to secrete apatite crystals. The deletion of FIP200 led to an abnormal increase in p62 expression and insufficient conversion of LC3-II, resulting in a defect in osteoblast function[39]. Besides, AMPK activation induced autophagy, as determined by the upregulation of LC3, increased autophagosome density and downregulation of p62[40]. Mice lacking the stress-inducing nuclear protein 1 showed an increase in bone mass. In nuclear protein 1-deficient osteoblasts, the expression of ATG, autophagosome formation, and cell survival was up-regulated[41]. Properly increased autophagy levels cannot only protect BMSCs from oxidative stress damage but also promote the survival of osteoblasts. Oxidative stress destroys many cellular components of osteoblasts and is considered to be a key pathogenic factor for bone loss[42,43]. Silencing ATG5 can inhibit the proliferation and differentiation of osteoblasts, making them more vulnerable to oxidative stress[44]. Autophagy also can regulate the mechanical stimulation required for osteoblast differentiation[45] and reduce the inhibitory and apoptotic effects of glucocorticoids on osteoblasts[46]. The presence of estrogen can promote the differentiation and autophagy of osteoblasts by up-regulating the expression of autophagy-related protein RAB3GAP1, thereby improving its survival rate and mineralization[47], probably through the ER-ERK-mTOR signaling pathway[48]. Besides, autophagy-deficient osteoblasts can cause increased levels of oxidative stress and are potent in the production of osteoclasts[49]. It has been reported that the knockout of the UBA domain and the LIR site in the selective autophagy receptor (NBR1) gene of the ubiquitination substrate could inhibit the MAPK-p38 pathway, promoting the differentiation of BMSCs to osteoblast and significantly increasing bone mass[50]. It has been also reported that the expression of autophagy and bone mineral density were significantly increased when the receptor NBR1 gene was knocked out in an animal model[51]. In an in vitro study, scientists further found the osteoblast activity and the differentiation ability were significantly enhanced after the NBR1 gene was deleted. Furthermore, studies have found that P62 activity has increased when the NBR1 gene was knocked out. All of these studies demonstrated the NBR1 gene has a positive effect on the autophagy process and also regulates p38 mitogen-activated protein kinase MAPK in the osteoblast in a negative way[51]. Therefore, knocking out the NBR1 gene will inhibit autophagy and change the mechanism of osteoblast[52]. Similarly, ATG7 and BECN1 genes of a knockout rat can also significantly reduce osteogenic efficiency[49]. Knockout of ATG7 can cause bone loss in mice and cause stress in the endoplasmic reticulum, and phenylbutyrate acid released through the body can alleviate the decrease in bone formation capacity caused by endoplasmic reticulum stress[53,54]. Indeed, the specific mechanism of autophagy in the regulation differentiation of osteoblasts and its effect remain to be further confirmed. Autophagy and osteocytes Osteocytes are the main cells in bone tissue, which are the terminally differentiated cells buried in the bone matrix[55]. The upregulation of autophagy may accompany the transition from osteoblast to osteocyte to recycle organelles and preserve nutrients. Osteocytes make up over 95% of the bone cells in the adult skeleton[56]. Hocking found that autophagy totally exists in the differentiation of osteoblasts to osteocytes. Hocking et al.[57] found that the higher the differentiation degree of the osteocytes, the higher the level of autophagy. Osteocytes are mechanically sensitive cells that can make adaptive changes to external forces. Appropriate mechanical loading can improve bone strength and inhibit bone loss. King et al.[58] found that mammalian cells are highly sensitive to mechanical stress and can induce non-dependent mTOR pathways that cause temporary autophagy. Autophagic vesicles are observed in osteoblast-like MLO-Y4 cells after fluid shear stress, which causes an increase in LC3-II and degradation of p62, indicating that fluid shear stress can induce protective autophagy of osteocytes, while mechanically induced autophagy is related to ATP metabolism and bone cell survival, suggesting that the use of drugs that regulate autophagy status in the clinic may enhance the survival of skeletal cells, providing new ideas for clinical prevention of bone aging-related diseases[59]. Autophagy of osteocytes is inversely related to oxidative stress and bone loss. Ovariectomy results in increased expression of Atg5, LC3, and Beclin1 in rats, while decreased expression of p62, and also reduces total antioxidant capacity, superoxide dismutase activity[60]. Autophagy can protect cells from the effects of oxidative stress, and give cells a certain ability to resist oxidative stress. Increased oxidative stress can also reduce the basic autophagy activity of bone cells, and vice versa. After moderate-level (1.4 mg/kg) pretreatment with glucocorticoids, osteocyte autophagy levels increased, and the ability to resist oxidative stress injury also increased, possibly by activating the MAPK/ERK signaling pathway[61]. Besides, EphrinB2 in bone cells will limit the accumulation of minerals. Bone cells lacking EphrinB2 can see more autophagosomes in vitro and in vivo. EphrinB2 may inhibit autophagy through the RhoA-ROCK signal[62]. EphrinB2 can also affect the mineralization function of osteoblasts[63]. Interestingly, ATG7 seems to change the appearance of bone cells. The loss of Atg7 disrupts the formation or maintenance of mouse bone cell networks. The bone cells of knockout mice are large, but the nucleus is small. The shape of the bone cell nucleus looks more round and eccentric. This may be the delay or prevention of Atg7 deletion. The number of cytoplasmic components associated with bone cell maturation[54]. Another study pointed out that periodic mechanical stretching will reduce the size and ovality of bone cells and increase the expression of LC3b and ATG7, indicating that autophagy upregulation will affect the spherical changes of bone cells[64]. Autophagy and osteoclasts Osteoclasts lie in small cavities called Howship’s lacunae. By the way of secret a variety of proteolytic enzymes to form absorption lacunae, the osteoclast finally functions as bone resorption[65]. Osteoclasts secrete β3 integrin and produce actin rings, which are adsorbed on the surface of the bone, causing the surface of the bone to form a wrinkled edge, and secrete proteins such as tartrate-resistant acid phosphatase and cathepsin K to acidify and absorb bone. The complex folding of this edge is through the lysosome and periosteal fusion. The fusion process of secreted lysosomes and wrinkle edges is closely related to autophagy. Atg5, Atg7, Atg4B, and LC3 are important for the formation of wrinkle edges and osteoclasts for osteoclasts, and for bone resorption in vivo and in vitro important. kindlin3, which is an necessary adaptor protein in the podosome, will interact with LC3B and undergo autophagy-mediated protein degradation to regulating osteoclast migration[66]. Besides, Rab7, which is required for osteoclast function, is localized to the edges of the folds in a manner dependent on Atg5[67,68]. When osteoclasts are exposed to hypoxic stress, it will cause an increase in autophagy flux and increase the expression of ATG to reduce cellular stress[69]. Beclin-1 is required for RANKL-induced osteoclast differentiation[70], RANKL can induce Bcl-2 phosphorylation and dissociate Beclin1 from the Bcl-2-Beclin1 complex, JNK1 can block this process[71]. During in vitro osteoclast differentiation, autophagy is activated and Beclin1 is enhanced. In vivo experiments found that mice lacking Beclin1 showed impaired osteoclast function and increased cortical bone thickness[72]. It may be related to KLF2 (kruppel-like factor 2) regulating Beclin1-mediated autophagy during osteoclast formation[73]. Under starvation conditions, GPCR kinase 2-interacting protein 1 induces disruption of Beclin1 and Bcl2 binding and contributes to osteoclasts autophagy[74]. It was found that the transport receptor p62, which had an important regulatory effect on the formation of autophagosomes, mutated in Paget’s bone disease, promoting osteoclast proliferation as well as cell activity[75]. The p62 protein is involved in the mTOR nutrient-sensing signal transduction pathway in influencing the differentiation and function of OCs[76]. During RANKL-induced osteoclast differentiation, p62 is significantly down-regulated in the initial stage of osteoclast formation, and then gradually increases over time. The expression of p62 is related to the ratio of LC3-II/LC3-I, This phenomenon may be caused by autophagy activation and plays an important role in F-actin loop formation[77]. In addition to Beclin1 and p62 being closely related to osteoclasts, IL-17A stimulates osteoclast differentiation and bone resorption and may promote autophagy activity by activating the RANKL-JNK pathway during osteoclast formation[78,79]. JNK1-mediated autophagy could promote RANKL-induced osteoclastogenesis via enhancing TRAF3 degradation, and JNK1 also could prevent osteoclast precursors apoptosis through autophagy-TRAF3 signaling[80]. Pro-inflammatory cytokine tumor necrosis factor activates osteoclasts in rheumatoid arthritis to initiate autophagy, and then promotes osteoclasts to perform bone resorption in the body. Chloroquine and hydroxychloroquine can increase The pH value that can inhibit autophagy and damage protein degradation in autolysosomes, which may explain the mechanism of chloroquine and hydroxychloroquine in preventing bone erosion[81]. It may be related to lysosomal pH regulating mTOR activity in osteoclasts[82]. However, some studies have reached the opposite conclusion. For example, OPG can enhance autophagy and inhibit osteoclast differentiation and bone resorption through the AMPK/mTOR/p70S6K signaling pathway in vitro[83]. In vitro culture of osteoclasts has found that the use of rapamycin can inhibit the reduction of cells and inhibit bone loss[84]. "
| [1] MIZUSHIMA N, KOMATSU M. Autophagy: renovation of cells and tissues. Cell. 2011;147(4):728-741. [2] MIZUSHIMA N, LEVINE B, CUERVO AM, et al. Autophagy fights disease through cellular self-digestion. Nature. 2008;451(7182):1069-1075. [3] CAMUZARD O, SANTUCCI-DARMANIN S, CARLE GF, et al. Role of autophagy in osteosarcoma. J Bone Oncol. 2019;16:100235. [4] CHADHA S, BEHL T, BUNGAU S, et al. Focus on the Multimodal Role of Autophagy in Rheumatoid Arthritis. Inflammation. 2021;44(1):1-12. [5] WANG S, DENG Z, MA Y, et al. The Role of Autophagy and Mitophagy in Bone Metabolic Disorders. Int J Biol Sci. 2020;16(14):2675-2691. [6] FLORENCIO-SILVA R, SASSO GR, SASSO-CERRI E, et al. Biology of Bone Tissue: Structure, Function, and Factors That Influence Bone Cells. Biomed Res Int. 2015;2015:421746. [7] GALLAGHER JC, SAI AJ. Molecular biology of bone remodeling: implications for new therapeutic targets for osteoporosis. Maturitas. 2010;65(4):301-307. [8] ASHFORD TP, PORTER KR. Cytoplasmic components in hepatic cell lysosomes. J Cell Biol. 1962;12(1):198-202. [9] CUERVO AM, WONG E. Chaperone-mediated autophagy: roles in disease and aging. Cell Res. 2014;24(1):92-104. [10] KLIONSKY DJ. The molecular machinery of autophagy: unanswered questions. J Cell Sci. 2005;118(Pt 1):7-18. [11] LI WW, LI J, BAO JK. Microautophagy: lesser-known self-eating. Cell Mol Life Sci. 2012;69(7):1125-1136. [12] RAVIKUMAR B, SARKAR S, DAVIES JE, et al. Regulation of mammalian autophagy in physiology and pathophysiology. Physiol Rev. 2010;90(4): 1383-1435. [13] WANG J, DAVIS S, ZHU M, MILLER EA, et al. Autophagosome formation: Where the secretory and autophagy pathways meet. Autophagy. 2017;13(5):973-974. [14] BAMPTON ET, GOEMANS CG, NIRANJAN D, et al. The dynamics of autophagy visualized in live cells: from autophagosome formation to fusion with endo/lysosomes. Autophagy. 2005;1(1):23-36. [15] KAMADA Y, FUNAKOSHI T, SHINTANI T, et al. Tor-mediated induction of autophagy via an Apg1 protein kinase complex. J Cell Biol. 2000; 150(6):1507-1513. [16] TANIDA I, MIZUSHIMA N, KIYOOKA M, et al. Apg7p/Cvt2p: A novel protein-activating enzyme essential for autophagy. Mol Biol Cell. 1999; 10(5):1367-1379. [17] SHINTANI T, MIZUSHIMA N, OGAWA Y, et al. Apg10p, a novel protein-conjugating enzyme essential for autophagy in yeast. EMBO J. 1999; 18(19):5234-5241. [18] POOLE AC, THOMAS RE, ANDREWS LA, et al. The PINK1/Parkin pathway regulates mitochondrial morphology. Proc Natl Acad Sci U S A. 2008;105(5):1638-1643. [19] ITAKURA E, KISHI-ITAKURA C, MIZUSHIMA N. The hairpin-type tail-anchored SNARE syntaxin 17 targets to autophagosomes for fusion with endosomes/lysosomes. Cell. 2012;151(6):1256-1269. [20] ANDING AL, BAEHRECKE EH. Cleaning House: Selective Autophagy of Organelles. Dev Cell. 2017;41(1):10-22. [21] MUNSON MJ, GANLEY IG. MTOR, PIK3C3, and autophagy: Signaling the beginning from the end. Autophagy. 2015;11(12):2375-2376. [22] WU DJ, ADAMOPOULOS IE. Autophagy and autoimmunity. Clin Immunol. 2017;176:55-62. [23] JABER FA, KHAN NM, ANSARI MY, et al. Autophagy plays an essential role in bone homeostasis. J Cell Physiol. 2019;234(8):12105-12115. [24] NOMBELA-ARRIETA C, RITZ J, SILBERSTEIN LE. The elusive nature and function of mesenchymal stem cells. Nat Rev Mol Cell Biol. 2011;12(2): 126-131. [25] QI M, ZHANG L, MA Y, et al. Autophagy Maintains the Function of Bone Marrow Mesenchymal Stem Cells to Prevent Estrogen Deficiency-Induced Osteoporosis. Theranostics. 2017;7(18):4498-4516. [26] MA Y, QI M, AN Y, et al. Autophagy controls mesenchymal stem cell properties and senescence during bone aging. Aging Cell. 2018; 17(1):e12709. [27] WAN Y, ZHUO N, LI Y, et al. Autophagy promotes osteogenic differentiation of human bone marrow mesenchymal stem cell derived from osteoporotic vertebrae. Biochem Biophys Res Commun. 2017;488(1):46-52. [28] OLIVER L, HUE E, PRIAULT M, et al. Basal autophagy decreased during the differentiation of human adult mesenchymal stem cells. Stem Cells Dev. 2012;21(15):2779-2788. [29] MIZUSHIMA N, YAMAMOTO A, MATSUI M, et al. In vivo analysis of autophagy in response to nutrient starvation using transgenic mice expressing a fluorescent autophagosome marker. Mol Biol Cell. 2004; 15(3):1101-1111. [30] LIU L, JIN X, HU CF, et al. Exosomes Derived from Mesenchymal Stem Cells Rescue Myocardial Ischaemia/Reperfusion Injury by Inducing Cardiomyocyte Autophagy Via AMPK and Akt Pathways. Cell Physiol Biochem. 2017;43(1):52-68. [31] HE H, ZENG Q, HUANG G, et al. Bone marrow mesenchymal stem cell transplantation exerts neuroprotective effects following cerebral ischemia/reperfusion injury by inhibiting autophagy via the PI3K/Akt pathway. Brain Res. 2019;1707:124-132. [32] LI J, ZHOU J, ZHANG D, et al. Bone marrow-derived mesenchymal stem cells enhance autophagy via PI3K/AKT signalling to reduce the severity of ischaemia/reperfusion-induced lung injury. J Cell Mol Med. 2015;19(10):2341-2351. [33] FAN P, YU XY, XIE XH, et al. Mitophagy is a protective response against oxidative damage in bone marrow mesenchymal stem cells. Life Sci. 2019;229:36-45. [34] SANGANI R, PERIYASAMY-THANDAVAN S, PATHANIA R, et al. The crucial role of vitamin C and its transporter (SVCT2) in bone marrow stromal cell autophagy and apoptosis. Stem Cell Res. 2015;15(2):312-321. [35] YANG M, WEN T, CHEN H, et al. Knockdown of insulin-like growth factor 1 exerts a protective effect on hypoxic injury of aged BM-MSCs: role of autophagy. Stem Cell Res Ther. 2018;9(1):284. [36] NUSCHKE A, RODRIGUES M, STOLZ DB, et al. Human mesenchymal stem cells/multipotent stromal cells consume accumulated autophagosomes early in differentiation. Stem Cell Res Ther. 2014;5(6):140. [37] NOBLE BS. The osteocyte lineage. Arch Biochem Biophys. 2008;473(2): 106-111. [38] KARSENTY G. Minireview: transcriptional control of osteoblast differentiation. Endocrinology. 2001;142(7):2731-2733. [39] LIU F, FANG F, YUAN H, et al. Suppression of autophagy by FIP200 deletion leads to osteopenia in mice through the inhibition of osteoblast terminal differentiation. J Bone Miner Res. 2013;28(11):2414-2430. [40] LI Y, SU J, SUN W, et al. AMP-activated protein kinase stimulates osteoblast differentiation and mineralization through autophagy induction. Int J Mol Med. 2018;41(5):2535-2544. [41] SHIRAKI M, XU X, IOVANNA JL, et al. Deficiency of stress-associated gene Nupr1 increases bone volume by attenuating differentiation of osteoclasts and enhancing differentiation of osteoblasts. FASEB J. 2019;33(8):8836-8852. [42] YANG YH, LI B, ZHENG XF, et al. Oxidative damage to osteoblasts can be alleviated by early autophagy through the endoplasmic reticulum stress pathway--implications for the treatment of osteoporosis. Free Radic Biol Med. 2014;77:10-20. [43] CAMUZARD O, SANTUCCI-DARMANIN S, BREUIL V, et al. Sex-specific autophagy modulation in osteoblastic lineage: a critical function to counteract bone loss in female. Oncotarget. 2016;7(41): 66416-66428. [44] WENG YM, KE CR, KONG JZ, et al. The significant role of ATG5 in the maintenance of normal functions of Mc3T3-E1 osteoblast. Eur Rev Med Pharmacol Sci. 2018;22(5):1224-1232. [45] ZHOU Z, SHI G, ZHENG X, et al. Autophagy activation facilitates mechanical stimulation-promoted osteoblast differentiation and ameliorates hindlimb unloading-induced bone loss. Biochem Biophys Res Commun. 2018;498(3):667-673. [46] HAN Y, ZHANG L, XING Y, et al. Autophagy relieves the function inhibition and apoptosis‑promoting effects on osteoblast induced by glucocorticoid. Int J Mol Med. 2018;41(2):800-808. [47] GAVALI S, GUPTA MK, DASWANI B, et al. Estrogen enhances human osteoblast survival and function via promotion of autophagy. Biochim Biophys Acta Mol Cell Res. 2019;1866(9):1498-1507. [48] YANG YH, CHEN K, LI B, et al. Estradiol inhibits osteoblast apoptosis via promotion of autophagy through the ER-ERK-mTOR pathway. Apoptosis. 2013;18(11):1363-1375. [49] NOLLET M, SANTUCCI-DARMANIN S, BREUIL V, et al. Autophagy in osteoblasts is involved in mineralization and bone homeostasis. Autophagy. 2014;10(11):1965-1977. [50] KIRKIN V, LAMARK T, SOU YS, et al. A role for NBR1 in autophagosomal degradation of ubiquitinated substrates. Mol Cell. 2009;33(4):505-516. [51] WHITEHOUSE CA, WATERS S, MARCHBANK K, et al. Neighbor of Brca1 gene (Nbr1) functions as a negative regulator of postnatal osteoblastic bone formation and p38 MAPK activity. Proc Natl Acad Sci U S A. 2010; 107(29):12913-12918. [52] DARCY A, MELTZER M, MILLER J, et al. A novel library screen identifies immunosuppressors that promote osteoblast differentiation. Bone. 2012;50(6):1294-1303. [53] LI H, LI D, MA Z, et al. Defective autophagy in osteoblasts induces endoplasmic reticulum stress and causes remarkable bone loss. Autophagy. 2018;14(10):1726-1741. [54] PIEMONTESE M, ONAL M, XIONG J, et al. Low bone mass and changes in the osteocyte network in mice lacking autophagy in the osteoblast lineage. Sci Rep. 2016;6:24262. [55] MAROTTI G, ZALLONE AZ, LEDDA M. Number, size and arrangement of osteoblasts in osteons at different stages of formation. Calcif Tissue Res. 1976;21 Suppl:96-101. [56] DALLAS SL, PRIDEAUX M, BONEWALD LF. The osteocyte: an endocrine cell ... and more. Endocr Rev. 2013;34(5):658-690. [57] HOCKING LJ, WHITEHOUSE C, HELFRICH MH. Autophagy: a new player in skeletal maintenance?. J Bone Miner Res. 2012;27(7):1439-1447. [58] KING JS, VELTMAN DM, INSALL RH. The induction of autophagy by mechanical stress. Autophagy. 2011;7(12):1490-1499. [59] ZHANG B, HOU R, ZOU Z, et al. Mechanically induced autophagy is associated with ATP metabolism and cellular viability in osteocytes in vitro. Redox Biol. 2018;14:492-498. [60] YANG Y, ZHENG X, LI B, et al. Increased activity of osteocyte autophagy in ovariectomized rats and its correlation with oxidative stress status and bone loss. Biochem Biophys Res Commun. 2014;451(1):86-92. [61] KAR R, RIQUELME MA, HUA R, et al. Glucocorticoid-Induced Autophagy Protects Osteocytes Against Oxidative Stress Through Activation of MAPK/ERK Signaling. JBMR Plus. 2018;3(4):e10077. [62] VRAHNAS C, BLANK M, DITE TA, et al. Increased autophagy in EphrinB2-deficient osteocytes is associated with elevated secondary mineralization and brittle bone. Nat Commun. 2019;10(1):3436. [63] BLANK M, SIMS NA. Cellular Processes by Which Osteoblasts and Osteocytes Control Bone Mineral Deposition and Maturation Revealed by Stage-Specific EphrinB2 Knockdown. Curr Osteoporos Rep. 2019; 17(5):270-280. [64] INABA N, KUROSHIMA S, UTO Y, et al. Cyclic mechanical stretch contributes to network development of osteocyte-like cells with morphological change and autophagy promotion but without preferential cell alignment in rat. Biochem Biophys Rep. 2017;11:191-197. [65] EVERTS V, KORPER W, HOEBEN KA, et al. Osteoclastic bone degradation and the role of different cysteine proteinases and matrix metalloproteinases: differences between calvaria and long bone. J Bone Miner Res. 2006;21(9):1399-1408. [66] ZHANG Y, CUI Y, WANG L, et al. Autophagy promotes osteoclast podosome disassembly and cell motility athrough the interaction of kindlin3 with LC3. Cell Signal. 2020;67:109505. [67] DESELM CJ, MILLER BC, ZOU W, et al. Autophagy proteins regulate the secretory component of osteoclastic bone resorption. Dev Cell. 2011;21(5):966-974. [68] GELMAN A, ELAZAR Z. Autophagic factors cut to the bone. Dev Cell. 2011;21(5):808-810. [69] ZHAO Y, CHEN G, ZHANG W, et al. Autophagy regulates hypoxia-induced osteoclastogenesis through the HIF-1α/BNIP3 signaling pathway. J Cell Physiol. 2012;227(2):639-648. [70] CHUNG YH, JANG Y, CHOI B, et al. Beclin-1 is required for RANKL-induced osteoclast differentiation. J Cell Physiol. 2014;229(12): 1963-1971. [71] KE D, JI L, WANG Y, et al. JNK1 regulates RANKL-induced osteoclastogenesis via activation of a novel Bcl-2-Beclin1-autophagy pathway. FASEB J. 2019;33(10):11082-11095. [72] ARAI A, KIM S, GOLDSHTEYN V, et al. Beclin1 Modulates Bone Homeostasis by Regulating Osteoclast and Chondrocyte Differentiation. J Bone Miner Res. 2019;34(9):1753-1766. [73] LAHA D, DEB M, DAS H. KLF2 (kruppel-like factor 2 [lung]) regulates osteoclastogenesis by modulating autophagy. Autophagy. 2019;15(12): 2063-2075. [74] ZHAO SJ, KONG FQ, CAI W, et al. GIT1 contributes to autophagy in osteoclast through disruption of the binding of Beclin1 and Bcl2 under starvation condition. Cell Death Dis. 2018;9(12):1195. [75] FRASER PD, ENFISSI EM, BRAMLEY PM. Genetic engineering of carotenoid formation in tomato fruit and the potential application of systems and synthetic biology approaches. Arch Biochem Biophys. 2009;483(2):196-204. [76] DURAN A, AMANCHY R, LINARES JF, et al. p62 is a key regulator of nutrient sensing in the mTORC1 pathway. Mol Cell. 2011;44(1):134-146. [77] LI RF, CHEN G, REN JG, et al. The adaptor protein p62 is involved in RANKL-induced autophagy and osteoclastogenesis. J Histochem Cytochem. 2014;62(12):879-888. [78] SONG L, TAN J, WANG Z, et al. Interleukin‑17A facilitates osteoclast differentiation and bone resorption via activation of autophagy in mouse bone marrow macrophages. Mol Med Rep. 2019;19(6):4743-4752. [79] KE D, FU X, XUE Y, et al. IL-17A regulates the autophagic activity of osteoclast precursors through RANKL-JNK1 signaling during osteoclastogenesis in vitro. Biochem Biophys Res Commun. 2018; 497(3):890-896. [80] KE D, ZHU Y, ZHENG W, et al. Autophagy mediated by JNK1 resists apoptosis through TRAF3 degradation in osteoclastogenesis. Biochimie. 2019;167:217-227. [81] LIN NY, STEFANICA A, DISTLER JH. Autophagy: a key pathway of TNF-induced inflammatory bone loss. Autophagy. 2013;9(8):1253-1255. [82] HU Y, CARRARO-LACROIX LR, WANG A, et al. Lysosomal pH Plays a Key Role in Regulation of mTOR Activity in Osteoclasts. J Cell Biochem. 2016;117(2):413-425. [83] TONG X, GU J, SONG R, et al. Osteoprotegerin inhibit osteoclast differentiation and bone resorption by enhancing autophagy via AMPK/mTOR/p70S6K signaling pathway in vitro. J Cell Biochem. 2018. doi:10.1002/jcb.27468 [84] OWEN HC, VANHEES I, GUNST J, et al. Critical illness-induced bone loss is related to deficient autophagy and histone hypomethylation. Intensive Care Med Exp. 2015;3(1):52. |
| [1] | Wang Jing, Xiong Shan, Cao Jin, Feng Linwei, Wang Xin. Role and mechanism of interleukin-3 in bone metabolism [J]. Chinese Journal of Tissue Engineering Research, 2022, 26(8): 1260-1265. |
| [2] | Xiao Hao, Liu Jing, Zhou Jun. Research progress of pulsed electromagnetic field in the treatment of postmenopausal osteoporosis [J]. Chinese Journal of Tissue Engineering Research, 2022, 26(8): 1266-1271. |
| [3] | Tian Chuan, Zhu Xiangqing, Yang Zailing, Yan Donghai, Li Ye, Wang Yanying, Yang Yukun, He Jie, Lü Guanke, Cai Xuemin, Shu Liping, He Zhixu, Pan Xinghua. Bone marrow mesenchymal stem cells regulate ovarian aging in macaques [J]. Chinese Journal of Tissue Engineering Research, 2022, 26(7): 985-991. |
| [4] | Hu Wei, Xie Xingqi, Tu Guanjun. Exosomes derived from bone marrow mesenchymal stem cells improve the integrity of the blood-spinal cord barrier after spinal cord injury [J]. Chinese Journal of Tissue Engineering Research, 2022, 26(7): 992-998. |
| [5] | Hou Jingying, Guo Tianzhu, Yu Menglei, Long Huibao, Wu Hao. Hypoxia preconditioning targets and downregulates miR-195 and promotes bone marrow mesenchymal stem cell survival and pro-angiogenic potential by activating MALAT1 [J]. Chinese Journal of Tissue Engineering Research, 2022, 26(7): 1005-1011. |
| [6] | Liang Xuezhen, Yang Xi, Li Jiacheng, Luo Di, Xu Bo, Li Gang. Bushen Huoxue capsule regulates osteogenic and adipogenic differentiation of rat bone marrow mesenchymal stem cells via Hedgehog signaling pathway [J]. Chinese Journal of Tissue Engineering Research, 2022, 26(7): 1020-1026. |
| [7] | Cui Xing, Sun Xiaoqi, Zheng Wei, Ma Dexin. Huangqin Decoction regulates autophagy to intervene with intestinal acute graft-versus-host disease in mice [J]. Chinese Journal of Tissue Engineering Research, 2022, 26(7): 1057-1062. |
| [8] | Fang Xiaolei, Leng Jun, Zhang Chen, Liu Huimin, Guo Wen. Systematic evaluation of different therapeutic effects of mesenchymal stem cell transplantation in the treatment of ischemic stroke [J]. Chinese Journal of Tissue Engineering Research, 2022, 26(7): 1085-1092. |
| [9] | Hu Weifan, Zheng Li, Li Dadi, Sun Yang, Zhao Fengchao. Overexpression of miR-25 downregulates titanium particle-induced osteoclast differentiation through the NFATc1 signaling pathway [J]. Chinese Journal of Tissue Engineering Research, 2022, 26(5): 682-687. |
| [10] | Yang Sidi, Wang Qian, Xu Nuo, Wang Ronghan, Jin Chuanqi, Lu Ying, Dong Ming. Biodentine enhances the proliferation and differentiation of osteoblasts through upregulating bone morphogenetic protein-2 [J]. Chinese Journal of Tissue Engineering Research, 2022, 26(4): 516-520. |
| [11] | Kang Kunlong, Wang Xintao. Research hotspot of biological scaffold materials promoting osteogenic differentiation of bone marrow mesenchymal stem cells [J]. Chinese Journal of Tissue Engineering Research, 2022, 26(4): 597-603. |
| [12] | Wei Zhoudan, Li Wenjin, Zhu Li, Wang Yu, Zhao Jiaoyang, Chen Yanan, Guo Dong, Hao Min. Platelet-rich fibrin as a material for alveolar ridge preservation significantly reduces the resorption of alveolar bone height and width after tooth extraction: a meta-analysis [J]. Chinese Journal of Tissue Engineering Research, 2022, 26(4): 643-648. |
| [13] | Feng Dongfei, He Hongxu, Xie Qi, Zhang Lili, Zhou Hui, Li Wei. Selection of key genes related to biological functions and regulation pathway in periodontal reconstruction [J]. Chinese Journal of Tissue Engineering Research, 2022, 26(2): 253-259. |
| [14] | Cao Wei, Mao Furong, Hu Xiaohua, Yang Xiaohong. N-6 methyladenosine RNA methylation regulates osteogenic and adipogenic differentiation of bone marrow mesenchymal stem cells [J]. Chinese Journal of Tissue Engineering Research, 2022, 26(2): 266-270. |
| [15] | Wu Saixuan, Zhang Mi, Dong Ming, Lu Ying, Niu Weidong. Regulatory role of Keap1/Nrf2/ARE signaling pathway in bone homeostasis [J]. Chinese Journal of Tissue Engineering Research, 2022, 26(2): 271-275. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||