Chinese Journal of Tissue Engineering Research ›› 2021, Vol. 25 ›› Issue (25): 4038-4044.doi: 10.12307/2021.017
Previous Articles Next Articles
Application status and prospect of induced pluripotent stem cell gene editing
Zhou Can, Yang Linan, Yang Kun, Liu Qi
- Department of Periodontology, Stomatological Hospital Affiliated to Zunyi Medical University, Zunyi 563003, Guizhou Province, China
-
Received:2020-09-12Revised:2020-09-15Accepted:2020-10-22Online:2021-09-08Published:2021-03-30 -
Contact:Liu Qi, MD, Professor, Department of Periodontology, Stomatological Hospital Affiliated to Zunyi Medical University, Zunyi 563003, Guizhou Province, China -
About author:Zhou Can, Master candidate, Department of Periodontology, Stomatological Hospital Affiliated to Zunyi Medical University, Zunyi 563003, Guizhou Province, China -
Supported by:the National Natural Science Foundation of China, No. 81860196 (to LQ); and National Natural Science Foundation of China, No. 81760199 (to YK)
CLC Number:
Cite this article
Zhou Can, Yang Linan, Yang Kun, Liu Qi. Application status and prospect of induced pluripotent stem cell gene editing[J]. Chinese Journal of Tissue Engineering Research, 2021, 25(25): 4038-4044.
share this article
Add to citation manager EndNote|Reference Manager|ProCite|BibTeX|RefWorks
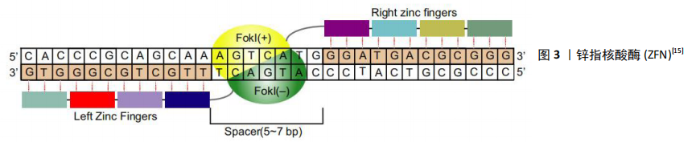
2.1 基因编辑工具 基因组编辑是一个利用自然细胞途径修复DNA断裂的过程,其中DNA断裂是基因组编辑的基础[13]。基因编辑依赖于各种核酸酶(也称“分子剪刀”)在基因组中特异性地引入位点,产生特异性DNA双链断裂(DSB)或单链DNA断裂(SSBs),随后诱导生物体通过非同源末端连接(NHEJ)或同源重组(HR)途径启动修复机制修复染色体特异性双链断裂,从而使基因组发生靶向配对安装,其范围可以从单个到数千个碱基对[14]。基因编辑可以针对性地编辑特定的基因组位点,用于纠正或功能上弥补遗传突变,敲除内源基因的表达或在精确的基因组位置插入转基因表达盒[15]。基于核酸酶的基因编辑系统,目前常用的基因工具包括聚簇的规则间隔的短回文重复序列(clustered regularly interspaced palindromic repeats/CRISPR-associated proteins,CRISPR),转录激活子样效应核酸酶(transcription activator-like nucleases,TALEN)和锌指核酸酶(zinc finger nucleases,ZFN)。随着基因编辑技术及基因编辑工具的发展,使各种改变宿主基因的操作变得容易[11]。 2.1.1 ZFN ZFN由特异性的锌指蛋白(zinc-finger protein,ZFP)结构域和非特异性的DNA裂解结构域(Ⅱ型限制性内切酶FokⅠ)组成,其中FokⅠ结构域发生二聚化是DNA裂解的前提条件[16]。ZFN被设计成一对能识别2个序列的单体位于目标位点侧,中间被5-7 bp间隔序列隔开,见图3[15],其中一个单体与正向链结合,另一个与反向链结合。单个锌指大约含有30个氨基酸,通常识别一个3 bp的DNA序列[17],每个锌指结构域通常以2个或多个锌指串联排列,结构域可以按比例结合更长的核苷酸序列,如3-6个锌指结构域用于生成单个ZFN亚基可与9-18 bp的DNA序列结合[18]。目前ZFN技术已经应用于多种生物的基因编辑[19]。CHANG等[20]利用ZFN技术在α-地中海贫血患者特异性的iPSCs插入野生型globin,从而获得包含正常globin的iPSCs以分化形成红细胞表达正常的β-球蛋白。 ZFN技术提供了快速对基因进行编辑的方法,但是其存在上下依赖效应[21],导致ZFN设计和筛选效率低、成本高,再加上其繁琐的程序限制了其成为实验室常规技术,从而使其在基因治疗领域中的应用受到限制。"
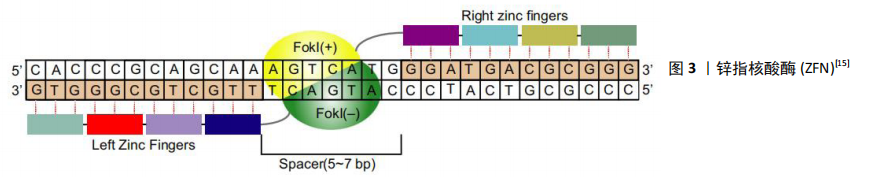
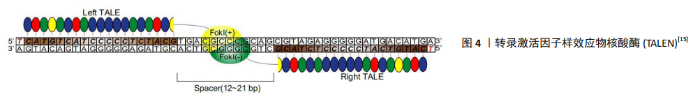
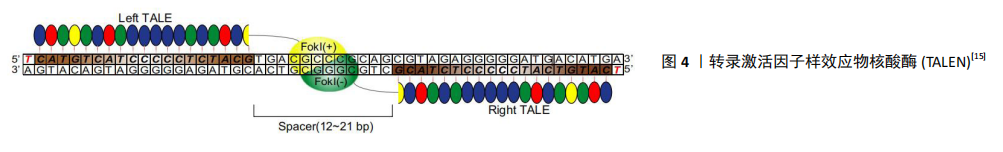
2.1.2 TALEN 与ZFN类似,TALEN由一个可设计的、特异性序列的DNA结合区域和一个来自FokⅠ的非特异性DNA裂解区域组成。不同的是,TALEN使用的DNA结构域来自植物病原性黄单胞菌属成员的DNA结合蛋白——转录激活因子样效应子(transcription activator-like effectors,TALEs)[22],从结构上讲可分为3个区域,分别为分泌信号的n端区域(NTD)、负责序列特异性DNA靶标识别的中心重复区(CRR)以及包含核定位信号(NLS)的CTD区域和转录激活区域(AD)。在TALEs结构中,每个重复单元由33-35个氨基酸组成,识别DNA中指定的一个目标碱基[23]。TALEs结构域不结合三联体核苷酸(遗传密码子),而是与特定的核苷酸序列结合,其与碱基对之间存在一对一的对应关系,见图4[15],具有识别单个特异性核苷酸而不受其周围其他结构域影响的能力[24]。 每个重复序列的特异性主要由位于12和13位的2个氨基酸赋予,称为重复可变二残基(repeat variable di-residue,RVDs)[25]。TALENs可以被设计成针对几乎任何给定的DNA序列,这是相对于其他类型核酸酶的一个关键优势。 2014年,ZHU等[26]通过TALENs插入标记基因,利用荧光蛋白 mCherry和不同神经转录因子LMX1a/FOXA2/OTX2的组合,发现可以优化多能干细胞向神经细胞分化所需要的转录因子。 TALENs较ZFNs更易于设计和建造,但序列长度通常为18-20 bp,更大的长度会限制TALEN的传递和表达,从而导致较低的特异性[24]。 "
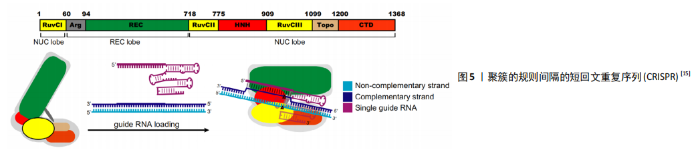
2.1.3 CRISPR 由于合成难度较大,ZFNs和TALENs的价格相对较贵。随着CRISPR-Cas9系统的发现和适应,基因组编辑变得更加容易。最早是通过细菌和古菌的免疫防御机制发现的CRISPR/Cas系统。细菌和古菌有一种被称为“有规律的集群间隔短回文重(CRISPR)/CRISPR-associated (Cas)”的自适应防御系统,该系统为RNA引导的DNA切割系统,以适应性形式存在,可抵抗细菌和古细菌中入侵的噬菌体或质粒[27]。 许多原核生物基因组都具有CRISPR,它们以CRISPR衍生的RNA形式来表达入侵病毒的短序列,称为crRNA。这些小RNA与蛋白质结合形成活性的核酸内切酶复合物,称为CrisprCas核酸内切酶复合物[28]。HECKL等[29]研究发现CrisprCas核酸内切酶复合物可产生定制的核酸酶以进行基因靶向,他通过将源自化脓链球菌(spCas9)的CRISPR/Cas9系统进行改造用于基因编辑,这是CRISPR系统第一次被改造用于基因编辑,涉及单个Crisper相关蛋白(Cas9)和2个RNA:靶向特异性RNA(target-specific CRISPR RNA,crRNA)和反式激活crRNA(Trans-activating crRNA,tracrRNA),这2个RNA结合成一个单嵌合引导RNA(single chimeric guide RNA,gRNA/sgRNA),有效地将Cas9活性引导至特定的靶标DNA,见图5[15]。crRNAs约由40个核苷酸序列组成,与tracr RNA结合激活可引导Cas9核酸酶将基因组中同源的双链DNA序列裂解。Cas9复合物的靶标识别要求存在一个特定序列基序,该基序应在gRNA靶向的20个核苷酸序列的下游,即原间隔子相邻基序(protospacer adjacent motif,PAM)。Crispr通过靶标特异性gRNAs实现了对序列的定制特异性,使其更高效、更容易获得目标基因。Crispr-Cas9复合物还可用于多种物种的基因编辑,包括哺乳动物和植物基因组[30-32]。MADSEN等[33]利用CRISPR/Cas9在人iPSCs中敲除了心肌细胞中的主要DNA甲基转移酶DNMT3A,从而比较基因敲除的心肌细胞和对照的人iPSCs来源的心肌细胞,发现人心肌细胞DNMT3A基因敲除对心肌细胞形态和功能的影响,进而表明DNA甲基化在心肌细胞的正常稳态和心脏应激过程中的重要作用,为心脏治疗发现了一个新的靶点。 CRISPR/Cas系统构建比ZFN和TALEN系统构建简便,普通实验室也可自行构建,显著提高了基因操作的简便性,并且CRISPR/Cas系统相对其他2种价格低,所以该系统为现在最常用的基因编辑工具。 2.2 应用 随着人类多能干细胞基因组编辑方法的发展,人类疾病遗传基础的研究也得到了迅速发展。体细胞的重新编程和基因组编辑代表了近年来能够彻底改变生物学和医学研究的2种技术,从而增强了干细胞在转化医学中的作用。在设计了正确的sgRNA之后,CRISPR/Cas9可通过基因敲除或敲入,基因干扰或激活以及其他与染色体相关的应用来靶向多种基因组序列,但保持剩余遗传背景不变[34]。iPSCs与CRISPR/Cas9的这些特点,使其既可用于干细胞的基础生物学研究,也可用于疾病建模和药物筛选的医学研究。 2.2.1 构建疾病模型 疾病模型是研究疾病发生、发展及药物作用机制的必要工具,iPSCs的基因编辑为其提供了更多的方法和选择。早期通过提取患者体内的多能干细胞进行研究,并通过病毒转基因表达和RNA干扰用于疾病特定基因的功能研究,但由于多能干细胞的伦理问题和个体差异性,以及病毒载体等插入位点突变导致转基因表达异常以及基因特异性较差时不能完全切除,现正在逐渐淡出疾病机制的研究[35-36]。 随着iPSCs的发现和基因编辑的发展,这些模型在变异特异性和细胞特异性的研究中已经取得了进展。 2011年,SEBASTTANO等[37]通过使用ZFN对来自2例镰状细胞性贫血患者的3个独立iPSCs系的β-球蛋白突变基因E6V进行了简单、高效和稳定的校正,结果表明,重编程的iPSCs和ZFN诱导的靶向基因可以联合使用来创建校正的患者源性iPSCs。随后,ZFN也广泛用于生成神经退行性疾病的iPSCs,从而构建模型[38]。TALEN也可用于产生多种疾病的iPSCs模型,包括杜氏肌营养不良症[39]、Niemann-Pick C型(NPC)病[40]、Lesch-Nyhan综合征[41]、X连锁严重联合免疫缺陷(SCID-X1)及各种血液病[42-44]。PARK等[45]通过提取具有反转基因型的血友病患者iPSCs,同时使用CRISPR-Cas9核酸酶将这些倒置染色体片段恢复后进行全基因组测序,分离出没有发生脱靶突变的频率高达6.7%的经反向校正的iPSCs,与其他血友病小鼠模型相比,校正后的iPSCs分化出来的内皮细胞成功表达了F8基因,同时在功能上恢复了Ⅷ因子缺乏的临床症状,从而为患者来源iPSCs进行染色体重排的功能校正提供了原理证明,并提出了潜在的治疗方法。 一般来说,利用合成核酸酶在iPSCs中建立同基因疾病模型有2种方法,即将疾病突变基因引入健康对照的iPSCs或纠正患者iPSCs中与疾病相关的基因突变。目前,利用iPSCs 建立的模型涉及了神经[46]、血液[47]、心血管[48]、胰腺以及肝脏等相关疾病[49-50],大多数为传统的2D模型。近年来,具有三维结构的体外模型更受各界学者青睐,因为其更接近生理状态,更适合于疾病病理研究。通过利用复杂器官芽的动态构图和构建3D结构及其相应的网络已成为现实,这种方法已经用于脑部异常和精神疾病的疾病建模中[51-52]。与外胚层组织结构的自组织类似,3D干细胞培养中的内胚层组织形成已被开发并应用于胃肠疾病建模[53]。但基于iPSCs技术的疾病模型有一个缺点,需要研究者区分致病的原因究竟是个别患者iPSCs突变还是受疾病本身背景的影响[54]。基因编辑工具可对关键致病位点进行基因编辑, 获得同基因背景对照, 从而排除研究过程中不同来源细胞系以及不同克隆间基因组或者表观多样性带来的研究干扰。Wang等[55]使用CRISPR/Cas9系统将致病基因SOD1和FUS基因校正后的患者特异iPSCs作为同基因背景对照, 来研究肌萎缩侧索硬化症。 2.2.2 药物开发 近年来,研究者们常基于iPSCs构建的疾病模型进行药物筛选试验从而开发对疾病有效的药物。LEE等[56] 第一次使用患者iPSCs构建了家族性自主神经功能障碍模型并对其进行了药物筛选,最终发现激动素植物激素可以减轻病理表型。随后,研究者们开始陆续对iPSCs进行了很多药物筛选试验,以测试不同化合物的效力和毒性。早期药物筛选均基于与疾病机制相关的靶标,成功率低,随着iPSCs的出现,表型筛选成为更多学者的选择[57]。患者来源iPSCs分化而来的细胞可能呈现分子和细胞表型,如果已知导致疾病表型的基因,则可以通过基因编辑方法来确定被选作药物筛选标准表型是否与疾病真正相关,并可以在模型和/或动物中进一步验证[58]。BURKHARDT等[59]通过使用散发性肌萎缩侧索硬化症患者的iPSCs进行疾病建模和药物筛选。散发性肌萎缩侧索硬化症患者最常见的病因是TDP-43(TAR DNA binding protein)的异常,研究者通过建立iPSCs模型在这些患者的运动神经元中发现TDP-43重新聚集,因此以TDP-43聚集作为药物筛选的标准,从1 757个活性化合物中筛选到了4类化合物,从而为散发性肌萎缩侧索硬化症的治疗带来了希望。 针对特定疾病的iPSCs的另一个应用是药物重新定位,即对已批准用于特定疾病的现有药物进行测试,以发现在其他疾病上的新应用。YAMASHITA等[60]研究发现成纤维细胞生长因子受体3(FGFR3)突变的软骨发育不全患者的iPSCs不能很好地分化成软骨组织,利用此模型从软骨缺陷表型中筛查软骨形成分化的iPSCs,确定了几种治疗心血管疾病的他汀类药物。研究发现,他汀类药物治疗可以挽救患者特异性的iPSCs模型和FGFR3骨骼发育不良小鼠模型,表明他汀类药物可以重新定位为软骨发育不良的候选药物。总体而言,基于iPSCs的药物筛选已被用于评估针对疾病的1 000多种化合物[61],除了测试药物或化合物的效力,仍然可以利用iPSCs来源模型测试不同遗传背景患者的不良药物反应,弥补使用传统动物模型进行药物安全性测定可能存在的缺陷。药物导致肝毒性损伤是临床药物开发中药物失败的常见原因[62]。研究表明,人类胚胎干细胞和iPSCs衍生肝细胞在肝中表达的细胞色素P450 3A4(CYP3A4)等功能分子可以吸收吲哚菁绿,并对已知的肝毒性药物产生反应,从而表明iPSCs衍生的肝细胞可以用于预测个体间肝脏药物代谢能力以及药物反应的差异[63]。 2.2.3 再生医学 再生医学于1992年被Kaiser[64]首次提出,是指通过一些手段再生或替代人类细胞、组织或器官,以恢复或建立正常的功能[65]。iPSCs用于再生医学主要是指重新编程获得的iPSCs通过定向分化的细胞注射,也称为细胞替代治疗。干细胞替代疗法为心脏再生治疗提供了巨大的潜力。然而,干细胞治疗的关键障碍之一是宿主环境中的缺血和炎症导致移植干细胞的显著损失。SIRISH等[66]用可溶性环氧化物水解酶抑制剂(SEHIs)抑制可溶性环氧化物水解酶(SEH)以减轻宿主心肌的炎症和纤维化,研究发现,CRISPR/Cas9介导的SEH基因沉默减少了血管紧张素Ⅱ攻击的hiPSC-CMS中caspase-3的裂解,提示SEH的敲除可以保护hiPSC-CMS免受凋亡的影响,为心脏干细胞治疗发现新的辅助手段。SNCA基因编码的α-突触核蛋白(α-syn)是一种小(14 kD)的突触前神经末梢蛋白,最初被确定为阿尔茨海默病淀粉样斑块中非淀粉样成分(NAC)的前体蛋白[67],其正常功能及其致病途径尚不清楚。CHEN等[68]利用常染色体显性突变的帕金森病患者的同源人类iPSCs来源的神经元、A53T和SNCA三倍体,以及通过基因组和表观基因组编辑的相应校正系,研究了SNCA在细胞核中的机制作用。研究结果表明SNCA突变通过捕获Ras相关核蛋白(RAN),阻止其转运维持正常核功能的关键核蛋白DNMT3A,以致于对细胞核产生不利影响。该结果指出α-突触核蛋白与RAN相互作用,从而侧向证明核质转运成分的缺陷可能是神经退行性疾病的一般病理机制驱动因素,为阿尔茨海默病的治疗提供新思路。β-球蛋白基因转移常被用作造血干细胞基因治疗的范例,但由于缺乏对经过基因校正的造血干细胞的选择、对治疗基因高水平表达的需求和细胞特异性转基因表达,研究者花费了大量时间对其进行研究[69]。随着基因编辑和iPSCs的开发,研究者将iPSCs与基因编辑结合进行血液疾病方面的研究。2007年,JACOB HANN等通过移植来自基因校正后iPSCs的造血祖细胞来校正SCD小鼠模型。自那以后,利用ZFNs、TALENs、CRISPR/ Cas靶向镰状细胞疾病患者iPSCs的点突变相关研究取得了进展。随后,研究者使用非整合载体从β地中海贫血患者获得了无整合iPSCs,并且通过基于TALEN的方法,ZFN介导的基因治疗或CRISPR/Cas9技术纠正了血红蛋白的突变[57]。iPSCs重新编程基于转录因子对体细胞的修饰,通过使用非整合策略或筛选安全整合位点,在限制与这一过程相关的肿瘤风险方面已经取得了巨大进展,但iPSCs的产生仍可能导致基因组不稳定。因此,这些细胞的突变状态和致瘤潜力需要在临床应用之前进行彻底测试[70]。 早在2014年,iPSCs的衍生品第一次进行临床试验[2],尽管iPSCs具有胚胎干细胞特性,但临床试验前仍需要解决这些问题。由于多能细胞在培养液中保存的时间较长,会积累核型异常和拷贝数变异,并失去杂合性[71]。因此,在临床使用之前,iPSCs衍生产品需要经过仔细的筛选,以确定是否存在潜在的风险基因改变,并经过严格的测试,以确保其纯度、质量和无菌性。"
| [1] TAKAHASHI K, YAMANAKA S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell. 2006;126(4): 663-676. [2] KIMBREL EA, LANZA R. Current status of pluripotent stem cells: moving the first therapies to the clinic. Nat Rev Drug Discov. 2015;14(10):681-692. [3] STRECKFUSS-BÖMEKE K, WOLF F, AZIZIAN A, et al. Comparative study of human-induced pluripotent stem cells derived from bone marrow cells, hair keratinocytes, and skin fibroblasts. Eur Heart J. 2013;34(33):2618-2629. [4] SI-TAYEB K, IDRISS S, CHAMPON B, et al. Urine-sample-derived human induced pluripotent stem cells as a model to study PCSK9-mediated autosomal dominant hypercholesterolemia. Dis Model Mech. 2016;9(1):81-90. [5] SPITALIERI P, TALARICO RV, BOTTA A, et al. Generation of Human Induced Pluripotent Stem Cells from Extraembryonic Tissues of Fetuses Affected by Monogenic Diseases. Cell Reprogram. 2015;17(4):275-287. [6] SPITALIERI P, TALARICO VR, MURDOCCA M, et al. Human induced pluripotent stem cells for monogenic disease modelling and therapy. World J Stem Cells. 2016;8(4):118-135. [7] JODAT YA, KANG MG, KIAEE K, et al. Human-Derived Organ-on-a-Chip for Personalized Drug Development. Curr Pharm Des. 2018;24(45):5471-5486. [8] EGLEN RM, REISINE T. Human iPS Cell-Derived Patient Tissues and 3D Cell Culture Part 1: Target Identification and Lead Optimization. SLAS Technol. 2019;24(1):3-17. [9] YANG PC. Induced Pluripotent Stem Cell (iPSC)-Derived Exosomes for Precision Medicine in Heart Failure. Circ Res. 2018;122(5):661-663. [10] DOUDNA JA, CHARPENTIER E. Genome editing. The new frontier of genome engineering with CRISPR-Cas9. Science. 2014;346(6213):1258096. [11] GUPTA SK, SHUKLA P. Gene editing for cell engineering: trends and applications. Crit Rev Biotechnol. 2017;37(5):672-684. [12] RAN FA, HSU PD, WRIGHT J, et al. Genome engineering using the CRISPR-Cas9 system. Nat Protoc. 2013;8(11):2281-2308. [13] DELEIDI M, YU C. Genome editing in pluripotent stem cells: research and therapeutic applications. Biochem Biophys Res Commun. 2016;473(3): 665-674. [14] JINEK M, CHYLINSKI K, FONFARA I, et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science. 2012; 337(6096):816-821. [15] HODGES CA, CONLON RA. Delivering on the promise of gene editing for cystic fibrosis. Genes Dis. 2018;6(2):97-108. [16] BITINAITE J, WAH DA, AGGARWAL AK, et al. FokI dimerization is required for DNA cleavage. Proc Natl Acad Sci U S A. 1998;95(18):10570-10575. [17] WOLFE SA, NEKLUDOVA L, PABO CO. DNA recognition by Cys2His2 zinc finger proteins. Annu Rev Biophys Biomol Struct. 2000;29:183-212. [18] LEE J, CHUNG JH, KIM HM, et al. Designed nucleases for targeted genome editing. Plant Biotechnol J. 2016;14(2):448-462. [19] CUI X, JI D, FISHER DA, et al. Targeted integration in rat and mouse embryos with zinc-finger nucleases. Nat Biotechnol. 2011;29(1):64-67. [20] CHANG CJ, BOUHASSIRA EE. Zinc-finger nuclease-mediated correction of α-thalassemia in iPS cells. Blood. 2012;120(19):3906-3914. [21] SANDER JD, DAHLBORG EJ, GOODWIN MJ, et al. Selection-free zinc-finger-nuclease engineering by context-dependent assembly (CoDA). Nat Methods. 2011;8(1):67-69. [22] BOGDANOVE AJ, SCHORNACK S, LAHAYE T. TAL effectors: finding plant genes for disease and defense. Curr Opin Plant Biol. 2010;13(4):394-401. [23] MILLER JC, TAN S, QIAO G, et al. A TALE nuclease architecture for efficient genome editing. Nat Biotechnol. 2011;29(2):143-148. [24] GUILINGER JP, PATTANAYAK V, REYON D, et al. Broad specificity profiling of TALENs results in engineered nucleases with improved DNA-cleavage specificity. Nat Methods. 2014;11(4):429-435. [25] READ AC, RINALDI FC, HUTIN M, et al. Suppression of Xo1-Mediated Disease Resistance in Rice by a Truncated, Non-DNA-Binding TAL Effector of Xanthomonas oryzae. Front Plant Sci. 2016;7:1516. [26] ZHU F, GAMBOA M, FARRUGGIO AP, et al. DICE, an efficient system for iterative genomic editing in human pluripotent stem cells. Nucleic Acids Res. 2014;42(5):e34. [27] MAKAROVA KS, WOLF YI, IRANZO J, et al. Evolutionary classification of CRISPR-Cas systems: a burst of class 2 and derived variants. Nat Rev Microbiol. 2020;18(2):67-83. [28] BARRANGOU R, FREMAUX C, DEVEAU H, et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science. 2007;315(5819):1709-1712. [29] HECKL D, KOWALCZYK MS, YUDOVICH D, et al. Generation of mouse models of myeloid malignancy with combinatorial genetic lesions using CRISPR-Cas9 genome editing. Nat Biotechnol. 2014;32(9):941-946. [30] HISANO Y, SAKUMA T, NAKADE S, et al. Precise in-frame integration of exogenous DNA mediated by CRISPR/Cas9 system in zebrafish. Sci Rep. 2015;5:8841. [31] KONDO S, UEDA R. Highly improved gene targeting by germline-specific Cas9 expression in Drosophila. Genetics. 2013;195(3):715-721. [32] BELHAJ K, CHAPARRO-GARCIA A, KAMOUN S, et al. Plant genome editing made easy: targeted mutagenesis in model and crop plants using the CRISPR/Cas system. Plant Methods. 2013;9(1):39. [33] MADSEN A, HÖPPNER G, KRAUSE J, et al. An Important Role for DNMT3A-Mediated DNA Methylation in Cardiomyocyte Metabolism and Contractility. Circulation. 2020;142(16):1562-1578. [34] LYU C, SHEN J, WANG R, et al. Targeted genome engineering in human induced pluripotent stem cells from patients with hemophilia B using the CRISPR-Cas9 system. Stem Cell Res Ther. 2018;9(1):92. [35] ZHANG HX, ZHANG Y, YIN H. Genome Editing with mRNA Encoding ZFN, TALEN, and Cas9. Mol Ther. 2019;27(4):735-746. [36] JACKSON AL, LINSLEY PS. Recognizing and avoiding siRNA off-target effects for target identification and therapeutic application. Nat Rev Drug Discov. 2010;9(1):57-67. [37] SEBASTIANO V, MAEDER ML, ANGSTMAN JF, et al. In situ genetic correction of the sickle cell anemia mutation in human induced pluripotent stem cells using engineered zinc finger nucleases. Stem Cells. 2011;29(11):1717-1726. [38] FONG H, WANG C, KNOFERLE J, et al. Genetic correction of tauopathy phenotypes in neurons derived from human induced pluripotent stem cells. Stem Cell Reports. 2013;1(3):226-234. [39] LI HL, FUJIMOTO N, SASAKAWA N, et al. Precise correction of the dystrophin gene in duchenne muscular dystrophy patient induced pluripotent stem cells by TALEN and CRISPR-Cas9. Stem Cell Reports. 2015;4(1):143-154. [40] MAETZEL D, SARKAR S, WANG H, et al. Genetic and chemical correction of cholesterol accumulation and impaired autophagy in hepatic and neural cells derived from Niemann-Pick Type C patient-specific iPS cells. Stem Cell Reports. 2014;2(6):866-880. [41] FRANK S, SKRYABIN BV, GREBER B. A modified TALEN-based system for robust generation of knock-out human pluripotent stem cell lines and disease models. BMC Genomics. 2013;14:773. [42] MENON T, FIRTH AL, SCRIPTURE-ADAMS DD, et al. Lymphoid regeneration from gene-corrected SCID-X1 subject-derived iPSCs. Cell Stem Cell. 2015; 16(4):367-372. [43] SUN N, ZHAO H. Seamless correction of the sickle cell disease mutation of the HBB gene in human induced pluripotent stem cells using TALENs. Biotechnol Bioeng. 2014;111(5):1048-1053. [44] PARK CY, KIM J, KWEON J, et al. Targeted inversion and reversion of the blood coagulation factor 8 gene in human iPS cells using TALENs. Proc Natl Acad Sci U S A. 2014;111(25):9253-9258. [45] PARK CY, KIM DH, SON JS, et al. Functional Correction of Large Factor VIII Gene Chromosomal Inversions in Hemophilia A Patient-Derived iPSCs Using CRISPR-Cas9. Cell Stem Cell. 2015;17(2):213-220. [46] DEROSA BA, VAN BAAREN JM, DUBEY GK, et al. Derivation of autism spectrum disorder-specific induced pluripotent stem cells from peripheral blood mononuclear cells. Neurosci Lett. 2012;516(1):9-14. [47] MOTAZEDIAN A, BRUVERIS FF, KUMAR SV, et al. Multipotent RAG1+ progenitors emerge directly from haemogenic endothelium in human pluripotent stem cell-derived haematopoietic organoids. Nat Cell Biol. 2020;22(1):60-73. [48] ITZHAKI I, MAIZELS L, HUBER I, et al. Modelling the long QT syndrome with induced pluripotent stem cells. Nature. 2011;471(7337):225-229. [49] KIM MJ, LEE EY, YOU YH, et al. Generation of iPSC-derived insulin-producing cells from patients with type 1 and type 2 diabetes compared with healthy control. Stem Cell Res. 2020;48:101958. [50] RASHID ST, CORBINEAU S, HANNAN N, et al. Modeling inherited metabolic disorders of the liver using human induced pluripotent stem cells. J Clin Invest. 2010;120(9):3127-3136. [51] LANCASTER MA, RENNER M, MARTIN CA, et al. Cerebral organoids model human brain development and microcephaly. Nature. 2013;501(7467):373-379. [52] MARIANI J, COPPOLA G, ZHANG P, et al. FOXG1-Dependent Dysregulation of GABA/Glutamate Neuron Differentiation in Autism Spectrum Disorders. Cell. 2015;162(2):375-390. [53] DEDHIA PH, BERTAUX-SKEIRIK N, Zavros Y, et al. Organoid Models of Human Gastrointestinal Development and Disease. Gastroenterology. 2016;150(5): 1098-1112. [54] HOCKEMEYER D, JAENISCH R. Induced Pluripotent Stem Cells Meet Genome Editing. Cell Stem Cell. 2016;18(5):573-586. [55] WANG L, YI F, FU L, et al. CRISPR/Cas9-mediated targeted gene correction in amyotrophic lateral sclerosis patient iPSCs. Protein Cell. 2017;8(5):365-378. [56] LEE G, PAPAPETROU EP, KIM H, et al. Modelling pathogenesis and treatment of familial dysautonomia using patient-specific iPSCs. Nature. 2009;461 (7262):402-406. [57] VINCENT F, LORIA P, PREGEL M, et al. Developing predictive assays: the phenotypic screening “rule of 3”. Sci Transl Med. 2015;7(293):293ps15. [58] MATSA E, BURRIDGE PW, YU KH, et al. Transcriptome Profiling of Patient-Specific Human iPSC-Cardiomyocytes Predicts Individual Drug Safety and Efficacy Responses In Vitro. Cell Stem Cell. 2016;19(3):311-325. [59] BURKHARDT MF, MARTINEZ FJ, WRIGHT S, et al. A cellular model for sporadic ALS using patient-derived induced pluripotent stem cells. Mol Cell Neurosci. 2013;56:355-364. [60] YAMASHITA A, MORIOKA M, KISHI H, et al. Statin treatment rescues FGFR3 skeletal dysplasia phenotypes. Nature. 2014;513(7519):507-511. [61] XU X, LEI Y, LUO J, et al. Prevention of β-amyloid induced toxicity in human iPS cell-derived neurons by inhibition of Cyclin-dependent kinases and associated cell cycle events. Stem Cell Res. 2013;10(2):213-227. [62] WILLIAMS DP. Modeling hepatic drug metabolism and toxicity: where are we heading? Future Med Chem. 2014;6(7):725-727. [63] TAKAYAMA K, INAMURA M, KAWABATA K, et al. Efficient generation of functional hepatocytes from human embryonic stem cells and induced pluripotent stem cells by HNF4alpha transduction. Mol Ther. 2012;20(1): 127-137. [64] KAISER LR. The future of multihospital systems. Top Health Care Financ. 1992;18(4):32-45. [65] MASON C, DUNNILL P. A brief definition of regenerative medicine. Regen Med. 2008;3(1):1-5. [66] SIRISH P, THAI PN, LEE JH, et al. Suppression of inflammation and fibrosis using soluble epoxide hydrolase inhibitors enhances cardiac stem cell-based therapy. Stem Cells Transl Med. 2020 Aug 13. doi: 10.1002/sctm.20-0143. [67] UÉDA K, FUKUSHIMA H, MASLIAH E, et al. Molecular cloning of cDNA encoding an unrecognized component of amyloid in Alzheimer disease. Proc Natl Acad Sci U S A. 1993;90(23):11282-11286. [68] CHEN V, MONCALVO M, TRINGALI D, et al. The mechanistic role of alpha-synuclein in the nucleus: impaired nuclear function caused by familial Parkinson’s disease SNCA mutations. Hum Mol Genet. 2020;29(18):3107-3121. [69] SII-FELICE K, NEGRE O, BRENDEL C, et al. Innovative Therapies for Hemoglobin Disorders. BioDrugs. 2020;34(5):625-647. [70] DOULATOV S, VO LT, CHOU SS, et al. Induction of multipotential hematopoietic progenitors from human pluripotent stem cells via respecification of lineage-restricted precursors. Cell Stem Cell. 2013;13(4): 459-470. [71] LUND RJ, NÄRVÄ E, LAHESMAA R. Genetic and epigenetic stability of human pluripotent stem cells. Nat Rev Genet. 2012;13(10):732-744. [72] SEAH YF, EL FARRAN CA, WARRIER T, et al. Induced Pluripotency and Gene Editing in Disease Modelling: Perspectives and Challenges. Int J Mol Sci. 2015;16(12):28614-28634. |
| [1] | Pu Rui, Chen Ziyang, Yuan Lingyan. Characteristics and effects of exosomes from different cell sources in cardioprotection [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(在线): 1-. |
| [2] | Lin Qingfan, Xie Yixin, Chen Wanqing, Ye Zhenzhong, Chen Youfang. Human placenta-derived mesenchymal stem cell conditioned medium can upregulate BeWo cell viability and zonula occludens expression under hypoxia [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(在线): 4970-4975. |
| [3] | Zhang Xiumei, Zhai Yunkai, Zhao Jie, Zhao Meng. Research hotspots of organoid models in recent 10 years: a search in domestic and foreign databases [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(8): 1249-1255. |
| [4] | Wang Xianyao, Guan Yalin, Liu Zhongshan. Strategies for improving the therapeutic efficacy of mesenchymal stem cells in the treatment of nonhealing wounds [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(7): 1081-1087. |
| [5] | Wan Ran, Shi Xu, Liu Jingsong, Wang Yansong. Research progress in the treatment of spinal cord injury with mesenchymal stem cell secretome [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(7): 1088-1095. |
| [6] | Liao Chengcheng, An Jiaxing, Tan Zhangxue, Wang Qian, Liu Jianguo. Therapeutic target and application prospects of oral squamous cell carcinoma stem cells [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(7): 1096-1103. |
| [7] | Xie Wenjia, Xia Tianjiao, Zhou Qingyun, Liu Yujia, Gu Xiaoping. Role of microglia-mediated neuronal injury in neurodegenerative diseases [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(7): 1109-1115. |
| [8] | Li Shanshan, Guo Xiaoxiao, You Ran, Yang Xiufen, Zhao Lu, Chen Xi, Wang Yanling. Photoreceptor cell replacement therapy for retinal degeneration diseases [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(7): 1116-1121. |
| [9] | Jiao Hui, Zhang Yining, Song Yuqing, Lin Yu, Wang Xiuli. Advances in research and application of breast cancer organoids [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(7): 1122-1128. |
| [10] | Wang Shiqi, Zhang Jinsheng. Effects of Chinese medicine on proliferation, differentiation and aging of bone marrow mesenchymal stem cells regulating ischemia-hypoxia microenvironment [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(7): 1129-1134. |
| [11] | Zeng Yanhua, Hao Yanlei. In vitro culture and purification of Schwann cells: a systematic review [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(7): 1135-1141. |
| [12] | Kong Desheng, He Jingjing, Feng Baofeng, Guo Ruiyun, Asiamah Ernest Amponsah, Lü Fei, Zhang Shuhan, Zhang Xiaolin, Ma Jun, Cui Huixian. Efficacy of mesenchymal stem cells in the spinal cord injury of large animal models: a meta-analysis [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(7): 1142-1148. |
| [13] | Hou Jingying, Yu Menglei, Guo Tianzhu, Long Huibao, Wu Hao. Hypoxia preconditioning promotes bone marrow mesenchymal stem cells survival and vascularization through the activation of HIF-1α/MALAT1/VEGFA pathway [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(7): 985-990. |
| [14] | Shi Yangyang, Qin Yingfei, Wu Fuling, He Xiao, Zhang Xuejing. Pretreatment of placental mesenchymal stem cells to prevent bronchiolitis in mice [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(7): 991-995. |
| [15] | Liang Xueqi, Guo Lijiao, Chen Hejie, Wu Jie, Sun Yaqi, Xing Zhikun, Zou Hailiang, Chen Xueling, Wu Xiangwei. Alveolar echinococcosis protoscolices inhibits the differentiation of bone marrow mesenchymal stem cells into fibroblasts [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(7): 996-1001. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||