[1] ISOSAKA T, MATSUO T, YAMAGUCHI T, et al. Htr2a-Expressing Cells in the Central Amygdala Control the Hierarchy between Innate and Learned Fear. Cell. 2015;163(5):1153-1164.
[2] PICHAT J, IGLESIAS JE, YOUSRY T, et al. A Survey of Methods for 3D Histology Reconstruction. Med Image Anal. 2018;46:73-105.
[3] RAGAN T, KADIRI LR, VENKATARAJU KU,et al. Serial two-photon tomography for automated ex vivo mouse brain imaging. Nat Methods. 2012;9(3):255-258.
[4] DENK W, STRICKLER JH, WEBB WW.Two-photon laser scanning fluorescence microscopy. Science. 1990;248(4951):73-76.
[5] HILLMAN EMC, VOLETI V, LI W, et al. Light-Sheet Microscopy in Neuroscience. Annu Rev Neurosci. 2019;42:295-313.
[6] ZHU D, LARIN KV, LUO Q, et al. Recent progress in tissue optical clearing. Laser Photon Rev. 2013;7(5):732-757.
[7] WENNER M. The most transparent research. Nat Med. 2009;15(10): 1106-1109.
[8] SDOBNOV AY, DARVIN ME, GENINA EA, et al. Recent progress in tissue optical clearing for spectroscopic application. Spectrochim Acta A Mol Biomol Spectrosc. 2018;197(6):216-229.
[9] SUSAKI EA, UEDA HR. Whole-body and Whole-Organ Clearing and Imaging Techniques with Single-Cell Resolution: Toward Organism-Level Systems Biology in Mammals. Cell Chem Biol. 2016; 23(1):137-157.
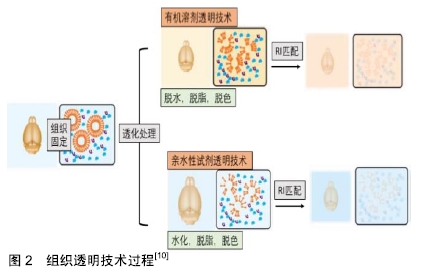
[10] TAINAKA K, KUNO A, KUBOTA SI, et al. Chemical Principles in Tissue Clearing and Staining Protocols for Whole-Body Cell Profiling. Annu Rev Cell Dev Biol. 2016;32(6):713-741.
[11] SPALTEHOLZ W. Uber das Durchsichtigmachen von menschlichen und tierischen Praparaten. (S Hierzel). 1914.
[12] GENINA EA, BASHKATOV AN, TUCHIN VV. Tissue optical immersion clearing. Expert Rev Med Devices. 2010;7(6):825-842.
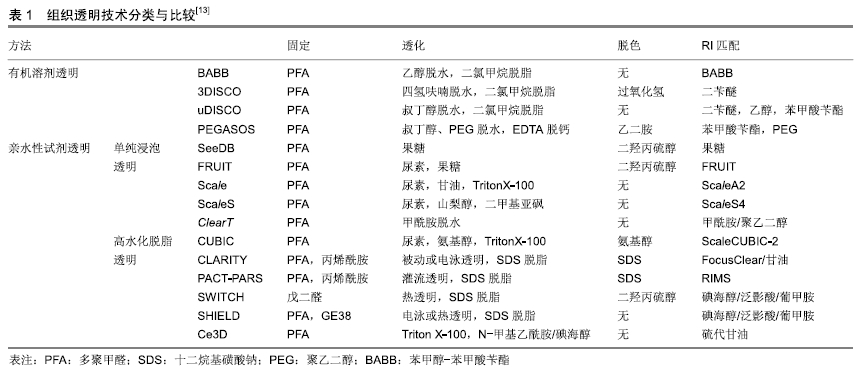
[13] RICHARDSON DS, LICHTMAN JW. SnapShot: Tissue Clearing. Cell. 2017;171(2):496-e1.
[14] DODT HU, LEISCHNER U, SCHIERLOH A, et al. Ultramicroscopy: three-dimensional visualization of neuronal networks in the whole mouse brain. Nat Methods. 2007;4(4):331-336.
[15] ERTURK A, BECKER K, JAHRLING N,et al. Three-dimensional imaging of solvent-cleared organs using 3DISCO. Nat Protoc. 2012; 7(11):1983-1995.
[16] YANG L, FEUCHTINGER A, MOLLER W, et al. Three-Dimensional Quantitative Co-Mapping of Pulmonary Morphology and Nanoparticle Distribution with Cellular Resolution in Nondissected Murine Lungs. ACS Nano. 2019;13(2):1029-1041.
[17] BELLE M, GODEFROY D, COULY G, et al. Tridimensional Visualization and Analysis of Early Human Development. Cell. 2017; 169(1):161-73 e12.
[18] KUBOTA SI, TAKAHASHI K, NISHIDA J,et al. Whole-Body Profiling of Cancer Metastasis with Single-Cell Resolution. Cell Rep. 2017;20(1): 236-250.
[19] RENIER N, WU Z, SIMON DJ, et al. iDISCO: a simple, rapid method to immunolabel large tissue samples for volume imaging. Cell. 2014; 159(4):896-910.
[20] HAHN C, BECKER K, SAGHAFI S, et al. High resolution imaging of fluorescent whole mouse brains using stabilised organic media (sDISCO). J Biophotonics.2019;12(8):e201800368.
[21] PAN C, CAI R, QUACQUARELLI FP, et al. Shrinkage-mediated imaging of entire organs and organisms using uDISCO. Nat Methods. 2016; 13(10):859-867.
[22] LI Y, XU J, WAN P, et al. Optimization of GFP Fluorescence Preservation by a Modified uDISCO Clearing Protocol. Front Neuroanat.2018;12:67.
[23] CAI R, PAN C, GHASEMIGHARAGOZ A, et al. Panoptic imaging of transparent mice reveals whole-body neuronal projections and skull-meninges connections. Nat Neurosci. 2019;22(2):317-327.
[24] JING D, ZHANG S, LUO W, et al. Tissue clearing of both hard and soft tissue organs with the PEGASOS method. Cell Res. 2018;28(8): 803-818.
[25] HAMA H, KUROKAWA H, KAWANO H,et al.Scale: a chemical approach for fluorescence imaging and reconstruction of transparent mouse brain. Nat Neurosci. 2011;14(11):1481-1488.
[26] HAMA H, HIOKI H, NAMIKI K, et al. ScaleS: an optical clearing palette for biological imaging. Nat Neurosci. 2015;18(10):1518-1529.
[27] KE MT, FUJIMOTO S, IMAI T. SeeDB: a simple and morphology- preserving optical clearing agent for neuronal circuit reconstruction. Nat Neurosci. 2013;16(8):1154-1161.
[28] HOU B, ZHANG D, ZHAO S,et al. Scalable and DiI-compatible optical clearance of the mammalian brain. Front Neuroanat. 2015;9:19.
[29] CHUNG K, WALLACE J, KIM SY, et al. Structural and molecular interrogation of intact biological systems. Nature. 2013;497(7449): 332-337.
[30] SARITAS T, PUELLES VG, SU XT, et al. Optical Clearing in the Kidney Reveals Potassium-Mediated Tubule Remodeling.Cell Rep. 2018; 25(10):2668-75 e3.
[31] DU H, HOU P, WANG L, et al. Modified CLARITY Achieving Faster and Better Intact Mouse Brain Clearing and Immunostaining.Sci Rep. 2019; 9(1):10571.
[32] TOMER R, YE L, HSUEH B, et al. Advanced CLARITY for rapid and high-resolution imaging of intact tissues. Nat Protoc. 2014;9(7): 1682-1697.
[33] TREWEEK JB, CHAN KY, FLYTZANIS NC, et al. Whole-body tissue stabilization and selective extractions via tissue-hydrogel hybrids for high-resolution intact circuit mapping and phenotyping.Nat Protoc. 2015;10(11):1860-1896.
[34] DABROWSKA N, JOSHI S, WILLIAMSON J,et al. Parallel pathways of seizure generalization. Brain. 2019;142(8):2336-2351.
[35] GREENBAUM A, CHAN KY, DOBREVA T, et al. Bone CLARITY: Clearing, imaging, and computational analysis of osteoprogenitors within intact bone marrow. Sci Transl Med. 2017;9(387). pii: eaah6518.
[36] BERNAL L, CISNEROS E, GARCIA-MAGRO N,et al. Immunostaining in whole-mount lipid-cleared peripheral nerves and dorsal root ganglia after neuropathy in mice. Sci Rep. 2019;9(1):8374.
[37] MURRAY E, CHO JH, GOODWIN D, et al. Simple, Scalable Proteomic Imaging for High-Dimensional Profiling of Intact Systems. Cell. 2015; 163(6):1500-1514.
[38] PARK YG, SOHN CH, CHEN R,et al. Protection of tissue physicochemical properties using polyfunctional crosslinkers. Nat Biotechnol. 2018;168(6):129-136
[39] SUSAKI EA, TAINAKA K, PERRIN D, et al. Whole-brain imaging with single-cell resolution using chemical cocktails and computational analysis. Cell. 2014;157(3):726-739.
[40] HASEGAWA S, SUSAKI EA, TANAKA T, et al. Comprehensive three-dimensional analysis (CUBIC-kidney) visualizes abnormal renal sympathetic nerves after ischemia/reperfusion injury. Kidney Int. 2019; 96(1):129-138.
[41] TAINAKA K, MURAKAMI TC, SUSAKI EA, et al. Chemical Landscape for Tissue Clearing Based on Hydrophilic Reagents. Cell Rep. 2018;2 4(8):2196-210 e9.
[42] CHEN L, LI G, LI Y, et al. UbasM: An effective balanced optical clearing method for intact biomedical imaging. Sci Rep. 2017;7(1):12218.
[43] LAI HM, LIU AKL, NG HHM,et al. Next generation histology methods for three-dimensional imaging of fresh and archival human brain tissues. Nat Commun. 2018;9(1):1066.
[44] LI W, GERMAIN RN, GERNER MY. Multiplex, quantitative cellular analysis in large tissue volumes with clearing-enhanced 3D microscopy (Ce3D). Proc Natl Acad Sci U S A. 2017;114(35):E7321- E7330.
[45] LI W, GERMAIN RN, GERNER MY. High-dimensional cell-level analysis of tissues with Ce3D multiplex volume imaging. Nat Protoc. 2019;14(6):1708-1033.
[46] BOSSOLANI GDP, PINTELON I, DETREZ JD, et al. Comparative analysis reveals Ce3D as optimal clearing method for in toto imaging of the mouse intestine. Neurogastroenterol Motil. 2019;31(5):e13560.
[47] MERTZ J. Optical sectioning microscopy with planar or structured illumination. Nat Methods. 2011;8(10):811-819.
[48] LI X, YU B, SUN Q, et al. Generation of a whole-brain atlas for the cholinergic system and mesoscopic projectome analysis of basal forebrain cholinergic neurons. Proc Natl Acad Sci U S A. 2018;115(2): 415-420.
[49] COMBS CA, SMIRNOV A, CHESS D, et al. Optimizing multiphoton fluorescence microscopy light collection from living tissue by noncontact total emission detection (epiTED). J Microsc. 2011;241(2): 153-161.
[50] RICHARDSON DS, LICHTMAN JW. Clarifying Tissue Clearing. Cell. 2015;162(2):246-257.
[51] SHARPE J, AHLGREN U, PERRY P, et al. Optical projection tomography as a tool for 3D microscopy and gene expression studies. Science. 2002;296(5567):541-545.
[52] ANDERSON GA, WONG MD, YANG J, et al. 3D imaging, registration, and analysis of the early mouse embryonic vasculature. Dev Dyn. 2013;242(5):527-538.
[53] ZENG W, PIRZGALSKA RM, PEREIRA MM,et al. Sympathetic neuro-adipose connections mediate leptin-driven lipolysis. Cell. 2015; 163(1):84-94.
[54] BAN S, CHO NH, MIN E, et al. Label-free optical projection tomography for quantitative three-dimensional anatomy of mouse embryo. J Biophotonics. 2019;12(7):e201800481.
[55] KELLER PJ, AHRENS MB. Visualizing whole-brain activity and development at the single-cell level using light-sheet microscopy. Neuron. 2015;85(3):462-483.
[56] WU Y, WAWRZUSIN P, SENSENEY J, et al. Spatially isotropic four-dimensional imaging with dual-view plane illumination microscopy. Nat Biotechnol. 2013;31(11):1032-1038.
[57] LERNER TN, SHILYANSKY C, DAVIDSON TJ, et al. Intact-Brain Analyses Reveal Distinct Information Carried by SNc Dopamine Subcircuits. Cell. 2015;162(3):635-647.
[58] TAINAKA K, KUBOTA SI, SUYAMA TQ, et al. Whole-body imaging with single-cell resolution by tissue decolorization. Cell. 2014;159(4): 911-924.
[59] MANO T, ALBANESE A, DODT HU, et al. Whole-Brain Analysis of Cells and Circuits by Tissue Clearing and Light-Sheet Microscopy. J Neurosci. 2018;38(44):9330-9337.
[60] ERTURK A, MAUCH CP, HELLAL F, et al. Three-dimensional imaging of the unsectioned adult spinal cord to assess axon regeneration and glial responses after injury. Nat Med. 2011;18(1):166-171.
[61] MURAKAMI TC, MANO T, SAIKAWA S, et al. A three-dimensional single-cell-resolution whole-brain atlas using CUBIC-X expansion microscopy and tissue clearing. Nat Neurosci. 2018;21(4):625-637.
[62] LONG F. Building strong bones: molecular regulation of the osteoblast lineage. Nat Rev Mol Cell Biol. 2011;13(1):27-38.
[63] KLAG KA, HORTON WA. Advances in treatment of achondroplasia and osteoarthritis. Hum Mol Genet. 2016;25(R1):R2-8.
[64] BERKE IM, MIOLA JP, DAVID MA, et al. Seeing through Musculoskeletal Tissues: Improving In Situ Imaging of Bone and the Lacunar Canalicular System through Optical Clearing. PLoS One. 2016;11(3):e0150268.
[65] NEU CP, NOVAK T, GILLILAND KF,et al. Optical clearing in collagen- and proteoglycan-rich osteochondral tissues. Osteoarthritis Cartilage. 2015;23(3):405-413.
[66] ACAR M, KOCHERLAKOTA KS, MURPHY MM, et al. Deep imaging of bone marrow shows non-dividing stem cells are mainly perisinusoidal. Nature. 2015;526(7571):126-130.
[67] GREENBAUM A, CHAN KY, DOBREVA T, et al. Bone CLARITY: Clearing, imaging, and computational analysis of osteoprogenitors within intact bone marrow. Sci Transl Med. 2017;9(387):68-82.
[68] SAADATPOUR Z, REZAEI A, EBRAHIMNEJAD H, et al. Imaging techniques: new avenues in cancer gene and cell therapy. Cancer Gene Ther. 2017;24(1):1-5.
[69] KOLESOVÁ H, BARTOŠ M, HSIEH WC, et al. Novel approaches to study coronary vasculature development in mice. Developmental Dynamics. 2018;247(8):1018-1027.
[70] LEE E,CHOI J, JO Y,et al.ACT-PRESTO: Rapid and consistent tissue clearing and labeling method for 3-dimensional (3D) imaging. Sci Rep. 2016;6:18631.
[71] YANG B, TREWEEK JB, KULKARNI RP,et al.Single-cell phenotyping within transparent intact tissue through whole-body clearing. Cell. 2014;158(4):945-958.
[72] WANG K, MILKIE DE, SAXENA A, et al. Rapid adaptive optical recovery of optimal resolution over large volumes. Nat Methods. 2014;11(6):625-628.
|