Chinese Journal of Tissue Engineering Research ›› 2020, Vol. 24 ›› Issue (20): 3269-3274.doi: 10.3969/j.issn.2095-4344.2532
Previous Articles Next Articles
Autophagy and epigenetic modification in inflammatory bowel disease
Guo Yajing1, Huang Yan2, Shi Yin2
- 1Yueyang Clinical Medical School, Shanghai University of Traditional Chinese Medicine, Shanghai 201203, China; 2Shanghai Institute of Acupuncture-Moxibustion and Meridian, Shanghai 200030, China
-
Received:2019-07-08Revised:2019-07-09Accepted:2019-09-02Online:2020-07-18Published:2020-04-14 -
Contact:Shi Yin, MD, Chief physician, Shanghai Institute of Acupuncture-Moxibustion and Meridian, Shanghai 200030, China -
About author:Guo Yajing, MD candidate, Yueyang Clinical Medical School, Shanghai University of Traditional Chinese Medicine, Shanghai 201203, China -
Supported by:the National Natural Science Foundation of China, No. 81674069; National Basic Research Program of China (973 Program), No. 2015CB554500
CLC Number:
Cite this article
Guo Yajing, Huang Yan, Shi Yin. Autophagy and epigenetic modification in inflammatory bowel disease[J]. Chinese Journal of Tissue Engineering Research, 2020, 24(20): 3269-3274.
share this article
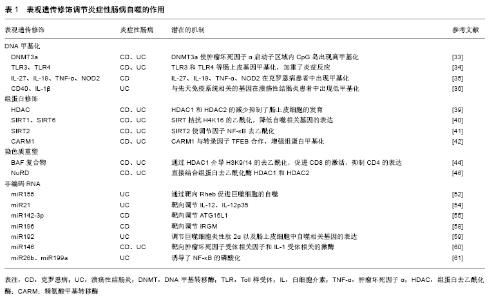
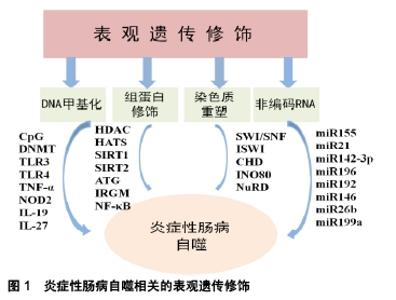
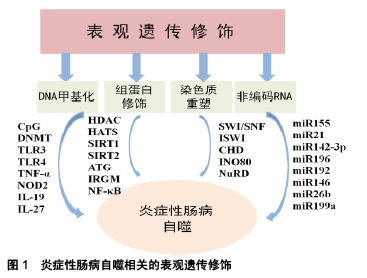
2.1 自噬与炎症性肠病 2.1.1 自噬与炎症性肠病肠道免疫 自噬是源自希腊语的术语,意思为“自食”,是泛素-蛋白酶体系统控制细胞内蛋白质降解的过程[8]。自噬是由于内在和外界环境压力而诱发的,包括营养和能量的条件限制、内质网应激、活性氧、激素失衡以及接触微生物等[9]。自噬的功能之一是控制先天免疫反应,对肠道免疫起到了重要调节作用,病原体激活炎症反应后导致免疫细胞的募集以激活适应性免疫应答[10]。巨噬细胞在吞噬微生物后激活烟酰胺腺嘌呤二核苷酸磷酸氧化酶(NADPH),产生活性氧、诱导型一氧化氮合成酶、一氧化氮,介导微生物吞噬体后的灭活[11],巨噬细胞自噬与肠道免疫应答相关,调节紊乱将会导致炎症性肠病的发生[12]。细胞因子如白细胞介素也参与了自噬,来自白细胞介素1家族(尤其是白细胞介素1β)在机体感染时,会产生先天性和适应性的免疫应答[13]。自噬通过影响白细胞介素1的分泌调节先天性淋巴细胞,CD4+T细胞是构成肠道淋巴细胞的最大群体,是肠道中宿主保护性和耐受性反应的中枢介质,CD4+T细胞的异常会造成慢性肠道炎症,导致炎症性肠病[14]。 自噬维持了肠上皮细胞和共生微生物群之间的平衡状态,对肠上皮细胞(包括Paneth细胞和杯状细胞)的特异性调控至关重要。Paneth细胞是肠道的先天免疫细胞,位于小肠的隐窝中,其顶部的颗粒含有抗微生物肽(溶菌酶、α-防御素和磷脂酶A2)、蛋白质、细胞因子、支架分子和蛋白酶等[15]。自噬基因缺陷小鼠Paneth细胞出现了异常颗粒,其可激活过氧化物酶受体信号的传导以及瘦素、脂联素的产生,进而造成肠黏膜的损伤[16]。Paneth细胞还分泌许多常见的免疫细胞因子,如干扰素β、白细胞介素1β、白细胞介素17和肿瘤坏死因子α,介导了肠道炎症反应[17]。在胃肠道中,杯状细胞存在于小肠和大肠中,杯状细胞可分泌黏蛋白,当黏蛋白分泌功能障碍或黏液层缺陷时,大量细菌到达上皮细胞并引发宿主过激的免疫反应,诱导炎症性肠病的发生[18]。 2.1.2 自噬相关基因与炎症性肠病 自噬相关基因(ATG)是细胞先天免疫的关键参与者,它们通过抗微生物选择性自噬(病原体消除)以及调控细胞溶质遍在样蛋白LC3的相关吞噬作用,引导LC3到内溶酶体区脂化,LC3在自噬激活后与自噬体膜上的磷脂酰乙醇胺结 合[19]。与ATG16样蛋白(ATG16L1)相关的炎症性肠病发生可归因于ATG16L1 T300A的变异,ATG16L1 T300A丧失了单核苷酸多态性,导致胱天蛋白酶介导的ATG16L1蛋白裂解增加,从而使自噬功能减退[20]。研究发现巨噬细胞中ATG16L1条件性敲除后,肠道内的耶尔森氏菌、伤寒沙门氏菌的清除受损,增加了对共生微生物的免疫应答,产生了大量活性氧,导致肠道慢性炎症,同时树突状细胞中CD11+T细胞自噬的特异性遭到破坏,诱导了白细胞介素1β和肿瘤坏死因子α的产生,加剧了结肠炎[21];ATG16L1缺陷型骨髓细胞用脂多糖刺激后,可以观察到促炎细胞因子白细胞介素6、白细胞介素18的分泌增加[22];在ATG16L1缺陷型小鼠的Paneth细胞中出现了颗粒胞吐作用途径受损,增加了小鼠炎症递质的释放[23];有ATG16L1遗传缺陷的小鼠不能消除内质网应激诱导的反应,激活了核转录因子κB,导致自噬在肠黏膜屏障中的作用失调,引起炎症性肠病[24]。 IRGM也是一种关键的自噬相关因子,它可以通过吞噬凋亡细胞来预防炎症并保护细胞免受氧化应激,调节免疫应答[25]。IRGM不仅激发了基本的自噬信号的传导,而且还参与了核心自噬机制,通过与自噬基因NOD2结合,促进泛素化和自噬起始因子的组装[26];IRGM参与细胞内细菌的清除,例如结核分枝杆菌、单核细胞增生李斯特氏菌和鼠伤寒沙门氏菌,同时IRGM通过影响Toll样受体的信号通路来调节细胞因子干扰素γ水平,减少了由干扰素γ介导的CD4+T细胞的死亡,说明IRGM在清除细胞内病原体以及维持细胞稳态中有一定的作用[27];IRGM缺陷小鼠表现出Paneth细胞异常,抗微生物肽、α-防御素20和溶菌酶的mRNA水平降低,同时增加了结肠和回肠中炎症的易感性[28]。 2.2 表观遗传与炎症性肠病自噬 表观遗传是指DNA序列不发生变化,但基因表达却发生了可遗传的改变,主要包括DNA甲基化、组蛋白修饰、染色质重塑以及非编码RNA的调控,这种改变在细胞发育和增殖过程中能稳定的传递。 2.2.1 DNA甲基化与炎症性肠病自噬 DNA甲基化是DNA的天然修饰方式,在DNA起始、复制、转录、修复、失活等过程中对维持遗传信息的稳定性发挥重要作用。DNA甲基化是研究最广泛的表观遗传修饰,在该过程中甲基被共价添加到胞嘧啶-鸟嘌呤双核苷酸(cytosine phosphate guanin,CpG)的胞嘧啶中,当DNA链上两条胞嘧啶残基被甲基化时,发生完全甲基化,CpG双核苷酸通常在基因组中稀疏,但相对集中在称为“CpG岛”的特定区域,CpG岛含有大于50%CpG双核苷酸的200个碱基序列,在DNA甲基化中发挥重要作用[29]。基因组DNA甲基化由DNA甲基转移酶(DNA methyltransferase,DNMT)催化,根据结构和功能的差异将甲基转移酶归为3类,分别是DNMT1、DNMT3a和DNMT3b[30]。在细胞分裂期间,DNMT1优先与半甲基化的DNA作为甲基化底物,DNA链相对应的序列被甲基化,未甲基化的链恢复对称的DNA甲基化;DNMT3a是胚胎发育中的主要酶;DNMT3b能够甲基化着丝粒远端具有高密度的CpG位点[31]。 RPS6KA2是一种丝氨酸激酶家族中的核糖体激酶,调节多种细胞过程,包括细胞生长、运动、增殖和细胞周期。RPS6KA参与遗传转录的不同阶段,包括PI3K/Akt /mTOR信号通路的传导,参与了自噬过程,在克罗恩病中发挥一定的作用[32];DNMT1的过表达造成了巨噬细胞中CD68的表达增强,介导了促炎反应,DNMT3a在结合T细胞受体后通过白细胞介素4和干扰素γ启动子区域的甲基化影响T细胞的极化,同时DNMT3a使肿瘤坏死因子α启动子区域内的CpG岛出现高甲基化,在先天性和适应性免疫应答中发挥重要作用[33];参与先天免疫自噬的基因如Toll样受体3和Toll样受体4等肠上皮基因的甲基化,都加重了炎症反应[34];NIMMO等[35]在21例克罗恩病患者中发现了27 578个DNA甲基化的CpG位点,通过靶向基因甲基化和DNA甲基化微阵列技术,检测到了与克罗恩病相关的基因(包括白细胞介素27、白细胞介素19、肿瘤坏死因子α和NOD2)出现了DNA的甲基化修饰;TAMAN等[36]从溃疡性结肠炎患者的DNA甲基化基因中,发现肿瘤坏死因子受体超家族、CD40和白细胞介素1β也出现了低甲基化。以上研究结果提示DNA甲基化参与了炎症性肠病的自噬。 2.2.2 组蛋白修饰与炎症性肠病自噬 组蛋白是真核生物体细胞染色质中的碱性蛋白质,富含精氨酸和赖氨酸等碱性氨基酸,组蛋白八聚体来自于4种不同组蛋白中的2种形成(H2A,H2B,H3和H4),核小体是由组蛋白八聚体及缠绕其外周的DNA组成[37]。组蛋白在细胞中发挥多种功能,它们可以安全地储存DNA,可以通过形成合适的局部结构来促进转录,也可以通过紧密卷绕DNA来阻止转录。当DNA与常染色质或“开放”状态的组蛋白结合时,转录很容易发生;当组蛋白与DNA紧密结合时,即所谓的异染色质或“封闭”状态,DNA聚合酶和转录因子转录受到限制[38]。组蛋白修饰主要包括乙酰化、磷酸化和甲基化等不同修饰,对基因的转录起到不同的影响。 在发育早期,小鼠肠上皮细胞中组蛋白去乙酰化酶HDAC1和HDAC2的减少抑制了分泌细胞的发育,成人的肠上皮细胞也出现异常,组蛋白去乙酰化酶在自噬调节中发挥一定的作用[39]。组蛋白乙酰转移酶的作用是向H4K16添加乙酰基,促进染色质去浓缩,过表达的DNA调控组蛋白去乙酰化酶Sirtuin 1(SIRT1)来拮抗H4K16的乙酰化,从而降低自噬相关基因的表达,同时SIRT6也可通过相关基因(ATG12、ATG3和ATG7)来调节自噬,包括公认的炎症性肠病自噬相关基因IRGM[40];SIRT2能够使细胞因子产生的主要调节因子NF-κB去乙酰化,SIRT2敲除后小鼠巨噬细胞中NF-κB的p65亚基出现乙酰化,导致结肠炎的抗炎能力下降[41]。精氨酸甲基转移酶1 (coactivator associated arginine methyltransferase 1,CARM1)与转录因子TFEB合作,增强组蛋白甲基化,CARM1作为NF-κB信号传导中的特异性调节因子,诱导转录因子进入AMP-SKP-CARM1信号级联自噬相关反应,在先天性和适应性免疫应答中起着重要作用[42]。组蛋白修饰在炎症性肠病自噬中的作用,也是目前研究的热点。 2.2.3 染色质重塑与炎症性肠病自噬 染色质结构的改变与基因的转录、表达以及DNA复制、重组等基本生物学过程紧密相关,引起染色质变化的主要因素是ATP依赖的染色质重塑以及DNA甲基化、组蛋白修饰等。染色质重塑复合物包括SWI/SNF、ISWI、CHD、INO80四大家族。SWI/SNF复合物首先被发现,其可促进核小体驱逐,导致H2A/H2B组蛋白二聚体的去除和剩余八聚体的进一步丧失[43]。人类SWI/SNF复合物存在2种方式,分别为BAF和PBAF,BAF复合物影响各种免疫细胞类型的发育,同时在T细胞发育的各个阶段发挥重要作用,它们接收不同的重塑染色质信号并调节转录。BAF复合物可通过HDAC1介导H3K9/14的去乙酰化,BRG1促进CD8的激活,而BRG1和BAF57抑制CD4的表达[44],与BAF复合物类似,ISWI复合物含有2个直系同源的ATP酶亚基,分别是SNF2H和SNF2L。 CHD复合物也是染色质重塑复合物中的一个家族,其N末端拥有一对chromo结构域,可以通过结合DNA、RNA和甲基化组蛋白H3的方式与染色质相互作用[45]。CHD家族分为三类亚家族,CHD亚家族Ⅱ的成员(CHD3和CHD4)形成的蛋白复合物NuRD,可以直接结合组蛋白去乙酰化酶HDAC1和HDAC2[46],NuRD主要与转录抑制有关,NuRD介导的抑制包括组蛋白H3在K9/14/27处的去乙酰化以及在基因启动子区域H3K4me3的去甲基化[47]。ATP依赖的染色质重塑的第4个主要家族为INO80,参与DNA的损伤,通过催化染色质中H2AZ(H2A的变体)/H2B二聚体置换H2A/H2B来控制H2AZ的水平,INO80和去乙酰化H2AZ之间的相互作用对于维持基因组的完整性至关重要[48]。ATP依赖的重塑复合物可以定位核小体,压缩染色质,促进不同DNA片段之间的相互作用,若此过程出现异常可导致包括自噬基因在内的表达紊乱,进而导致炎症性肠病的发生[49-50]。组蛋白修饰参与了染色质重塑复合物调节,进一步影响了自噬相关因子的表达。 2.2.4 非编码RNA与炎症性肠病自噬 非编码RNA在基因调控中发挥不同的作用,目前研究较多的是微小RNA(miRNAs),miRNAs是在真核生物中发现的一类内源性的具有转录后调控功能的非编码RNA,初级miRNAs(pri-miRNAs)是在RNA聚合酶Ⅱ作用下转录产生的。miRNAs具有多种编码方式,pri-miRNAs产生的miRNAs以簇的形式编码;一些miRNAs在内含子中编码,或在宿主蛋白的外显子中编码,它们可以与宿主基因共调节转录[51]。目前已知miR155、miR21、miR142-3p、miR223、miR106b、miR196、miR192、miR146、miR26b和miR199a等miRNAs均参与自噬相关基因的调控,与自噬过程相关。 miR155是首批确定在免疫中发挥作用的miRNAs之一,多与自噬过程相关[52],miR155靶向信号传导抑制因子SOCS-1,是脂多糖诱导的巨噬细胞活化的负调节因子,通过树突状细胞进行抗原呈递来激活免疫信号通路[53]。miR21在效应T细胞中表达最高,其潜在的靶标是白细胞介素12、白细胞介素12p35,白细胞介素12是参与适应性免疫应答的巨噬细胞和树突状细胞释放的关键细胞因子,对自噬有一定调节作用[54]。miR142-3p参与各种失调的炎症反应,其结合位点接近ATG16L1的末端,在化学诱导的急性结肠炎中miR142-3p显著升高,可能是由于miRNA的降解机制调节了ATG16L1,诱导包括炎症性肠病在内的肠道炎症和自噬[55];miR223是一种新的重要的自噬调节因子,ATG16L1是miR223的一个靶点,miR223的过表达减弱了ATG16L1蛋白和mRNA的水平[56];miR106b也可调节多种自噬相关基因,包括ATG16L1的表达[57];经典的miR196靶向调节IRGM,其水平决定靶细胞自噬或细胞死亡,对细胞内病原体的防御具有一定作用[58];miR192是溃疡性结肠炎相关的miRNA,调节巨噬细胞炎性肽2α以及肠上皮细胞中自噬相关基因的表达[59];同时miR146靶向肿瘤坏死因子受体相关因子和白细胞介素1受体相关的激酶[60];miR26b和miR199a诱导了NF-κB的磷酸化,在炎症性肠病的自噬中发挥作用[61],见表1,图1。 "
| [1] ASSADSANGABI A, EVANS CA, CORFE BM, et al. Application of Proteomics to Inflammatory Bowel Disease Research: Current Status and Future Perspectives. Gastroenterol Res Pract. 2019;2019: 1426954. [2] MANNINEN P, KARVONEN AL, HUHTALA H, et al. Mortality in ulcerative colitis and Crohn's disease. A population-based study in Finland. J Crohns Colitis. 2012;6(5):524-528. [3] HANSEN R, THOMSON JM, EL-OMAR EM, et al. The role of infection in the aetiology of inflammatory bowel disease. J Gastroenterol. 2010; 45(3):266-276. [4] KABAT AM, POTT J, MALOY KJ. The Mucosal Immune System and Its Regulation by Autophagy. Front Immunol. 2016;7:240. [5] EBBO M, CRINIER A, VÉLY F, et al. Innate lymphoid cells: major players in inflammatory diseases. Nat Rev Immunol. 2017;17(11): 665-678. [6] SHIN HR, KIM H, KIM KI, et al. Epigenetic and transcriptional regulation of autophagy. Autophagy. 2016;12(11):2248-2249. [7] HUANG H, FANG M, JOSTINS L, et al. Fine-mapping inflammatory bowel disease loci to single-variant resolution. Nature. 2017;547(7662): 173-178. [8] IIDA T, YOKOYAMA Y, WAGATSUMA K, et al. Impact of Autophagy of Innate Immune Cells on Inflammatory Bowel Disease. Cells. 2018;8(1): E7. [9] KROEMER G, MARIÑO G, LEVINE B. Autophagy and the integrated stress response. Mol Cell. 2010;40(2):280-293. [10] NETEA-MAIER RT, PLANTINGA TS, VAN DE VEERDONK FL, et al. Modulation of inflammation by autophagy: Consequences for human disease. Autophagy. 2016;12(2):245-260. [11] BRÜNE B, DEHNE N, GROSSMANN N, et al. Redox control of inflammation in macrophages. Antioxid Redox Signal. 2013;19(6):595-637. [12] WU Y, YAO J, XIE J, et al. The role of autophagy in colitis-associated colorectal cancer. Signal Transduct Target Ther. 2018;3:31. [13] PERAL DE CASTRO C, JONES SA, NÍ CHEALLAIGH C, et al. Autophagy regulates IL-23 secretion and innate T cell responses through effects on IL-1 secretion. J Immunol.2012;189(8): 4144-4153. [14] GOLDBERG R, PRESCOTT N, LORD GM, et al. The unusual suspects--innate lymphoid cells as novel therapeutic targets in IBD. Nat Rev Gastroenterol Hepatol. 2015;12(5):271-283. [15] CULLEN TW, SCHOFIELD WB, BARRY NA, et al. Gut microbiota. Antimicrobial peptide resistance mediates resilience of prominent gut commensals during inflammation. Science. 2015;347(6218):170-175. [16] CADWELL K, LIU JY, BROWN SL, et al. A key role for autophagy and the autophagy gene Atg16l1 in mouse and human intestinal Paneth cells. Nature. 2008;456(7219):259-263. [17] TAKAHASHI N, VANLAERE I, DE RYCKE R, et al. IL-17 produced by Paneth cells drives TNF-induced shock. J Exp Med. 2008;205(8): 1755-1761. [18] ITOH H, BECK PL, INOUE N, et al. A paradoxical reduction in susceptibility to colonic injury upon targeted transgenic ablation of goblet cells. J Clin Invest. 1999;104(11):1539-1547. [19] MIZUSHIMA N, NODA T, YOSHIMORI T, et al. A protein conjugation system essential for autophagy. Nature. 1998;395(6700):395-398. [20] VANDUSSEN KL, LIU TC, LI D, et al. Genetic variants synthesize to produce paneth cell phenotypes that define subtypes of Crohn's disease. Gastroenterology. 2014;146(1):200-209. [21] ZHANG H, ZHENG L, CHEN J, et al. The protection role of Atg16l1 in CD11c+dendritic cells in murine colitis. Immunobiology. 2017;222(7): 831-841. [22] WILDENBERG ME, VOS AC, WOLFKAMP SC, et al. Autophagy attenuates the adaptive immune response by destabilizing the immunologic synapse. Gastroenterology. 2012;142(7):1493-1503.e6. [23] CADWELL K, PATEL KK, KOMATSU M, et al. A common role for Atg16L1, Atg5 and Atg7 in small intestinal Paneth cells and Crohn disease. Autophagy. 2009;5(2):250-252. [24] ADOLPH TE, TOMCZAK MF, NIEDERREITER L, et al. Paneth cells as a site of origin for intestinal inflammation. Nature. 2013;503(7475): 272-276. [25] GRÉGOIRE IP, RABOURDIN-COMBE C, FAURE M. Autophagy and RNA virus interactomes reveal IRGM as a common target. Autophagy. 2012;8(7):1136-1137. [26] CHAUHAN S, MANDELL MA, DERETIC V. IRGM governs the core autophagy machinery to conduct antimicrobial defense. Mol Cell. 2015;58(3):507-521. [27] FENG CG, WEKSBERG DC, TAYLOR GA, et al. The p47 GTPase Lrg-47 (Irgm1) links host defense and hematopoietic stem cell proliferation. Cell Stem Cell. 2008;2(1):83-89. [28] LIU B, GULATI AS, CANTILLANA V, et al. Irgm1-deficient mice exhibit Paneth cell abnormalities and increased susceptibility to acute intestinal inflammation. Am J Physiol Gastrointest Liver Physiol. 2013;305(8):G573-584. [29] BEN-AVRAHAM D, MUZUMDAR RH, ATZMON G. Epigenetic genome-wide association methylation in aging and longevity. Epigenomics. 2012;4(5):503-509. [30] LOW D, MIZOGUCHI A, MIZOGUCHI E. DNA methylation in inflammatory bowel disease and beyond. World J Gastroenterol. 2013;19(32):5238-5249. [31] WIENHOLZ BL, KARETA MS, MOAREFI AH, et al. DNMT3L modulates significant and distinct flanking sequence preference for DNA methylation by DNMT3A and DNMT3B in vivo. PLoS Genet. 2010;6(9):e1001106. [32] PANCHOLI S, LYKKESFELDT AE, HILMI C, et al. ERBB2 influences the subcellular localization of the estrogen receptor in tamoxifen-resistant MCF-7 cells leading to the activation of AKT and RPS6KA2. Endocr Relat Cancer. 2008;15(4):985-1002. [33] JONES B, CHEN J. Inhibition of IFN-gamma transcription by site-specific methylation during T helper cell development. EMBO J. 2006;25(11):2443-2452. [34] KRAICZY J, NAYAK K, ROSS A, et al. Assessing DNA methylation in the developing human intestinal epithelium: potential link to inflammatory bowel disease. Mucosal Immunol. 2016;9(3):647-658. [35] NIMMO ER, PRENDERGAST JG, ALDHOUS MC, et al. Genome-wide methylation profiling in Crohn's disease identifies altered epigenetic regulation of key host defense mechanisms including the Th17 pathway. Inflamm Bowel Dis. 2012;18(5):889-899. [36] TAMAN H, FENTON CG, HENSEL IV, et al. Genome-wide DNA Methylation in Treatment-naïve Ulcerative Colitis. J Crohns Colitis. 2018;12(11):1338-1347. [37] LUGER K, MÄDER AW, RICHMOND RK, et al. Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature. 1997;389(6648): 251-260. [38] RANSOM M, DENNEHEY BK, TYLER JK. Chaperoning histones during DNA replication and repair. Cell. 2010;140(2):183-195. [39] ROOSTAEE A, BENOIT YD, BOUDJADI S, et al. Epigenetics in Intestinal Epithelial Cell Renewal. J Cell Physiol. 2016;231(11):2361-2367. [40] GARCIA-PETERSON LM, NDIAYE MA, SINGH CK, et al. SIRT6 histone deacetylase functions as a potential oncogene in human melanoma. Genes Cancer. 2017;8(9-10):701-712. [41] LO SASSO G, MENZIES KJ, MOTTIS A, et al. SIRT2 deficiency modulates macrophage polarization and susceptibility to experimental colitis. PLoS One. 2014;9(7):e103573. [42] SHIN HJ, KIM H, OH S, et al. AMPK-SKP2-CARM1 signalling cascade in transcriptional regulation of autophagy. Nature. 2016;534(7608): 553-557. [43] DECHASSA ML, SABRI A, PONDUGULA S, et al. SWI/SNF has intrinsic nucleosome disassembly activity that is dependent on adjacent nucleosomes. Mol Cell. 2010;38(4):590-602. [44] HOTA SK, BRUNEAU BG. ATP-dependent chromatin remodeling during mammalian development. Development. 2016;143(16):2882-2897. [45] HALL JA, GEORGEL PT. CHD proteins: a diverse family with strong ties. Biochem Cell Biol. 2007;85(4):463-476. [46] LE GUEZENNEC X, VERMEULEN M, BRINKMAN AB, et al. MBD2/NuRD and MBD3/NuRD, two distinct complexes with different biochemical and functional properties. Mol Cell Biol. 2006;26(3): 843-851. [47] YAMADA T, YANG Y, HEMBERG M, et al. Promoter decommissioning by the NuRD chromatin remodeling complex triggers synaptic connectivity in the mammalian brain. Neuron. 2014;83(1):122-134. [48] PAPAMICHOS-CHRONAKIS M, WATANABE S, RANDO OJ, et al. Global regulation of H2A.Z localization by the INO80 chromatin- remodeling enzyme is essential for genome integrity. Cell. 2011; 144(2):200-213. [49] KALITSIS P, ZHANG T, MARSHALL KM, et al. Condensin, master organizer of the genome. Chromosome Res. 2017;25(1):61-76. [50] RANKIN S, DAWSON DS. Recent advances in cohesin biology. F1000Res. 2016;5. pii: F1000 Faculty Rev-1909. [51] WEBER MJ. New human and mouse microRNA genes found by homology search. FEBS J. 2005;272(1):59-73. [52] WANG J, YANG K, ZHOU L, et al. MicroRNA-155 promotes autophagy to eliminate intracellular mycobacteria by targeting Rheb. PLoS Pathog. 2013;9(10):e1003697. [53] EVEL-KABLER K, SONG XT, ALDRICH M, et al. SOCS1 restricts dendritic cells' ability to break self tolerance and induce antitumor immunity by regulating IL-12 production and signaling. J Clin Invest. 2006;116(1):90-100. [54] WU H, NEILSON JR, KUMAR P, et al. miRNA profiling of naïve, effector and memory CD8 T cells. PLoS One. 2007;2(10):e1020. [55] ZHAI Z, WU F, DONG F, et al. Human autophagy gene ATG16L1 is post-transcriptionally regulated by MIR142-3p. Autophagy. 2014;10(3): 468-479. [56] LI Y, ZHOU D, REN Y, et al. Mir223 restrains autophagy and promotes CNS inflammation by targeting ATG16L1. Autophagy. 2019;15(3): 478-492. [57] HUANG Y, CHUANG AY, RATOVITSKI EA. Phospho-ΔNp63α/miR- 885-3p axis in tumor cell life and cell death upon cisplatin exposure. Cell Cycle. 2011;10(22):3938-3947. [58] LU C, CHEN J, XU HG, et al. MIR106B and MIR93 prevent removal of bacteria from epithelial cells by disrupting ATG16L1-mediated autophagy. Gastroenterology. 2014;146(1):188-199. [59] WU F, GUO NJ, TIAN H, et al. Peripheral blood microRNAs distinguish active ulcerative colitis and Crohn's disease. Inflamm Bowel Dis. 2011; 17(1):241-250. [60] TAGANOV KD, BOLDIN MP, CHANG KJ, et al. NF-kappaB-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc Natl Acad Sci U S A. 2006; 103(33):12481-12486. [61] POLYTARCHOU C, HOMMES DW, PALUMBO T, et al. MicroRNA214 Is Associated With Progression of Ulcerative Colitis, and Inhibition Reduces Development of Colitis and Colitis-Associated Cancer in Mice. Gastroenterology. 2015;149(4):981-992. |
| [1] | Duan Liyun, Cao Xiaocang. Human placenta mesenchymal stem cells-derived extracellular vesicles regulate collagen deposition in intestinal mucosa of mice with colitis [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(7): 1026-1031. |
| [2] | Ma Zetao, Zeng Hui, Wang Deli, Weng Jian, Feng Song. MicroRNA-138-5p regulates chondrocyte proliferation and autophagy [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(5): 674-678. |
| [3] | Xie Yang, Zhang Shujiang, Liu Menglan, Luo Ying, Yang Yang, Li Zuoxiao. Mechanism by which rapamycin protects spinal cord neurons in experimental autoimmune encephalomyelitis mice [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(5): 695-700. |
| [4] | Liu Bo, Chen Xianghe, Yang Kang, Sun Changliang, Yu Huilin, Lu Pengcheng. Epigenetic reprogramming and exercise regulation of bone metabolism disorders [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(20): 3210-3218. |
| [5] | Tong Jie, Liao Ying, Chen Zhengyu, Sun Guanghua . Osteoarthritic chondrocyte autophagy and regulation of mitogen-activated protein kinase signaling pathway [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(20): 3246-3251. |
| [6] | Chen Xinling, Wang Shenglan. Cell autophagy, pathway, regulation and its multiple correlations with pulmonary hypertension [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(2): 311-316. |
| [7] | Jiang Tao, Ling Cuimin, Chen Qingzhen, Yang Bingxuan, Lin Yanping, Shao Min. Icariin prevents osteoporosis by activating autophagy and promoting osteoblast differentiation [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(17): 2643-2649. |
| [8] | Lai Shuaiwei, Zhang Shasha, Liu Xiaoyun, Haniya Mazhar, Amber Naz, He Lin, Zhu Hongxin. Ultraviolet resistance-associated gene (Uvrag) deficiency promotes cellular senescence in the heart [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(14): 2241-2246. |
| [9] | Zhang Xujian, Zhao Zhenqun, Liu Wanlin. The role of endoplasmic reticulum stress in the pathogenesis of steroid-induced avascular necrosis of the femoral head [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(11): 1759-1765. |
| [10] | Liu Xin , Du Bin, Gao Lili, Sun Guangquan, Wang Xu, Yuan Peng, Lin Xuanye. Effect of Yougui Decoction on autophagy and fate of bone marrow mesenchymal stem cells in rats with glucocorticoid-associated femoral head necrosis [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(1): 20-25. |
| [11] | Xian Guoyan, Chen Weishen, Zhang Ziji, Ye Yongyu, Pan Baiqi, Zheng Linli, Sheng Puyi. Construction of a stable Atg5 gene knockdown cell line in RAW 264.7 cells by lentivirus infection [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(1): 84-89. |
| [12] | Zhu Ning, Yang Xinming, Ruan Jianwei. Triptolide improves spinal cord injury recovery via upregulation of autophagy and inhibition of apoptosis in Thy-yfp transgenic mice [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(在线): 1-. |
| [13] | Xu Guofeng, Li Xuebin, Tang Yifan, Zhao Yin, Zhou Shengyuan, Chen Xiongsheng, Jia Lianshun. The role of autophagy in ossification of the human ligamentum flavum [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(8): 1174-1181. |
| [14] | Han Yanfu, Tao Ran, Sun Tianjun. Autophagy has an early protective effect against fibroblast injury induced by advanced glycation end products [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(35): 5619-5624. |
| [15] | Zhu Ning, Yang Xinming, Ruan Jianwei. Triptolide improves spinal cord injury recovery via upregulation of autophagy and inhibition of apoptosis in Thy-YFP transgenic mice [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(35): 5650-5655. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||