Chinese Journal of Tissue Engineering Research ›› 2013, Vol. 17 ›› Issue (7): 1251-1258.doi: 10.3969/j.issn.2095-4344.2013.07.020
Previous Articles Next Articles
Improving PCR efficiency in the promoter region of serum amyloid A-activating transcription factor
Cao Xi-mei1, Liang Jun-hong2, Zhang Chao2, Wan Dong-fang2, Meng Xiao-ping2, Guo Da-wei2
- 1 Department of Histology and Embryology, Shanxi Medical University, Taiyuan 030001, Shanxi Province, China
2 School of Forensic Medicine, Shanxi Medical University, Taiyuan 030001, Shanxi Province, China
-
Received:2012-07-24Revised:2012-08-27Online:2013-02-12Published:2013-02-12 -
Contact:Guo Da-wei, Doctor, Professor, Doctoral supervisor, School of Forensic Medicine, Shanxi Medical University, Taiyuan 030001, Shanxi Province, China guo8dawei@yahoo.com.cn -
About author:Cao Xi-mei★, Master, Lecturer, Department of Histology and Embryology, Shanxi Medical University, Taiyuan 030001, Shanxi Province, China caoximei@163.com
CLC Number:
Cite this article
Cao Xi-mei, Liang Jun-hong, Zhang Chao, Wan Dong-fang, Meng Xiao-ping, Guo Da-wei. Improving PCR efficiency in the promoter region of serum amyloid A-activating transcription factor[J]. Chinese Journal of Tissue Engineering Research, 2013, 17(7): 1251-1258.
share this article
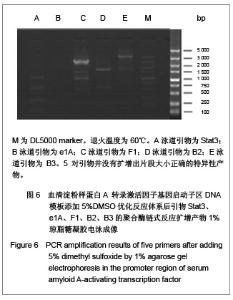
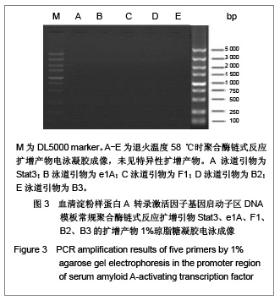
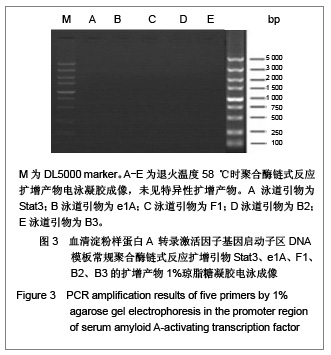
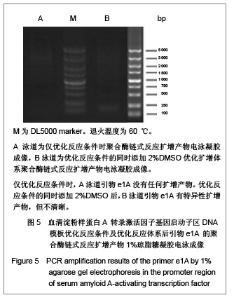
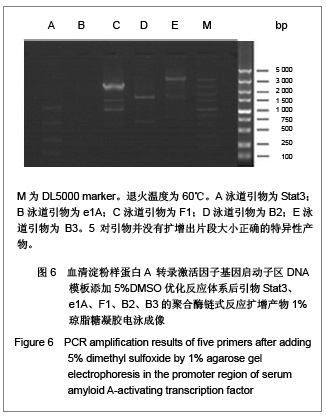
优化反应条件后,实验中低于60 ℃的退火温度几乎没有扩增产物,将退火温度升至高于60 ℃时伴随扩增产物出现非特异性扩增,实验证实退火温度为60 ℃时扩增最理想。优化反应条件后,退火温度为60 ℃时的扩增产物电泳后凝胶成像可见图4,5。引物Stat3没有扩增产物,引物F1、B2均可见特异性扩增产物,引物B3扩增产物非常弱,引物e1A没有任何扩增产物。 2.4 THP-1细胞基因组DNA优化反应条件及优化反应体系后的聚合酶链式反应扩增 优化反应条件的同时,在聚合酶链式反应扩增体系中加入3种不同浓度的增强剂DMSO以优化反应体系,结果表明反应体系中添加5%或10%DMSO易生成非特异性扩增产物,添加2%DMSO能起到优化扩增效率的作用。添加5%DMSO的扩增产物电泳后凝胶成像见图6。"
| [1] Ray BK, Murphy R, Ray P, et al. SAF-2, a splice variant of SAF-1,acts as a negative regulation of transcription. J Biol Chem. 2002;277(48):46822-46830.[2] Ray A, Dhar S, Shakya A, et al. SAF-3, a novel splice variant of the SAF-1/MAZ/Pur-1 family, is expressed during inflammation. FEBS J. 2009;276(15):4276-4286. [3] Ray A, Shakya A, Kumar D, et al. Inflammation-responsive transcription factor SAF-1 activity is linked to the development of amyloid A amyloidosis. J Immunol. 2006; 177(4):2601-2609.[4] Song J, Ugai H, Nakata-Tsutsui H, et al. Transcriptional regulation by zinc-finger proteins Sp1 and MAZ involves interactions with the same cis-elements. Int J Mol Med. 2003;11(5):547-553.[5] Auwerx J. The human leukemia cell line, THP-1: a multifacetted model for the study of monocyte-macrophage differentiation. Experientia. 1991;47(1):22-31. [6] Daigneault M, Preston JA, Marriott HM, et al. The identification of markers of macrophage differentiation in PMA-stimulated THP-1 cells and monocyte-derived macrophages. PLoS One. 2010;5(1): e8668. [7] Qin Z. The use of THP-1 cells as a model for mimicking the function and regulation of monocytes and macrophages in the vasculature. Atherosclerosis. 2012;221(1):2-11. [8] Zhou L, Shen LH, Hu LH, et al. Retinoid X receptor agonists inhibit phorbol-12-myristate-13-acetate (PMA)-induced differentiation of monocytic THP-1 cells into macrophages. Mol Cell Biochem. 2010;335(1-2):283-289. [9] Saiki RK, Gelfand DH, Stoffel S, et al. Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science. 1988;239(4839):487-491. [10] Kramer MF, Coen DM. Enzymatic amplification of DNA by PCR: standard procedures and optimization. Curr Protoc Cytom. 2006; Appendix 3:Appendix 3K.[11] Whitney SE, Sudhir A, Nelson RM, et al. Principles of rapid polymerase chain reactions: mathematical modeling and experimental verification. Comput Biol Chem. 2004;28(3): 195-209. [12] Vosberg HP. The polymerase chain reaction: an improved method for the analysis of nucleic acids. Hum Genet. 1989;83(1):1-15.[13] Chang PL, Hsieh WS, Chiang CL, et al. Identification of individual DNA molecule of Mycobacterium tuberculosis by nested PCR-RFLP and capillary electrophoresis. Talanta. 2008;77(1):182-188. [14] Housley DJ, Zalewski ZA, Beckett SE, et al. Design factors that influence PCR amplification success of cross-species primers among 1147 mammalian primer pairs. BMC Genomics. 2006;7:253.[15] Harris S, Jones DB. Optimisation of the polymerase chain reaction. Br J Biomed Sci. 1997;54(3):166-173.[16] Orpana AK, Ho TH, Stenman J. Multiple heat pulses during PCR extension enabling amplification of GC-rich sequences and reducing amplification bias. Anal Chem. 2012;84(4): 2081-2087.[17] Chouljenko V, Jayachandra S, Rybachuk G, et al. Efficient long-PCR site-specific mutagenesis of a high GC template. Biotechniques. 1996;21(3):472-474,476-478,480.[18] Rychlik W, Spencer WJ, Rhoads RE. Optimization of the annealing temperature for DNA amplification in vitro. Nucleic Acids Res. 1990;18(21):6409-6412. [19] Wu DY, Ugozzoli L, Pal BK, et al. The effect of temperature and oligonucleotide primer length on the specificity and efficiency of amplification by the polymerase chain reaction. DNA Cell Biol. 1991;10(3):233-238. [20] Shaffer AL, Wojnar W, Nelson W. Amplification, detection, and automated sequencing of gibbon interleukin-2 mRNA by Thermus aquaticus DNA polymerase reverse transcription and polymerase chain reaction. Anal Biochem. 1990;190(2): 292-296.[21] Eggerding FA. A one-step coupled amplification and oligonucleotide ligation procedure for multiplex genetic typing. PCR Methods Appl. 1995;4(6):337-345.[22] Jensen MA, Fukushima M, Davis RW. DMSO and betaine greatly improve amplification of GC-rich constructs in de novo synthesis. PLoS One. 2010;5(6):e11024. [23] Hubé F, Reverdiau P, Iochmann S, et al. Improved PCR method for amplification of GC-rich DNA sequences. Mol Biotechnol. 2005;31(1):81-84. [24] Musso M, Bocciardi R, Parodi S, et al. Betaine, dimethyl sulfoxide, and 7-deaza-dGTP, a powerful mixture for amplification of GC-rich DNA sequences. J Mol Diagn. 2006;8(5):544-550.[25] Pratyush DD, Tiwari S, Kumar A, et al. A new approach to touch down method using betaine as co-solvent for increased specificity and intensity of GC rich gene amplification. Gene. 2012;497(2):269-272. [26] Bachmann HS, Siffert W, Frey Ulrich. Successful amplification of extremely GC-rich promoter regions using a novel ‘slowdown PCR’ technique. Pharmacogenetics. 2003;13(12): 759-766.[27] Frey UH, Bachmann HS, Peters J, et al. PCR-amplification of GC-rich regions: 'slowdown PCR'. Nat Protoc. 2008;3(8):1312-1317.[28] Hung T, Mak K, Fong K. A specificity enhancer for polymerase chain reaction. Nucleic Acids Res. 1990;18(16):4953.[29] Kovárová M, Dráber P. New specificity and yield enhancer of polymerase chain reactions. Nucleic Acids Res. 2000;28(13): E70.[30] Markoulatos P, Siafakas N, Moncany M. Multiplex polymerase chain reaction: a practical approach. J Clin Lab Anal. 2002;16(1): 47-51. |
| [1] | Liu Bo, Chen Xianghe, Yang Kang, Yu Huilin, Lu Pengcheng. Mechanism of DNA methylation in exercise intervention for osteoporosis [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(5): 791-797. |
| [2] | Liu Bo, Chen Xianghe, Yang Kang, Sun Changliang, Yu Huilin, Lu Pengcheng. Epigenetic reprogramming and exercise regulation of bone metabolism disorders [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(20): 3210-3218. |
| [3] |
Zhang Cong, Zhao Yan, Du Xiaoyu, Du Xinrui, Pang Tingjuan, Fu Yining, Zhang Hao, Zhang Buzhou, Li Xiaohe, Wang Lidong.
Biomechanical analysis of the lumbar spine and pelvis in adolescent
idiopathic scoliosis with lumbar major curve |
| [4] | Xu Guofeng, Li Xuebin, Tang Yifan, Zhao Yin, Zhou Shengyuan, Chen Xiongsheng, Jia Lianshun. The role of autophagy in ossification of the human ligamentum flavum [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(8): 1174-1181. |
| [5] | He Yujie, Wang Haiyan, Li Zhijun, Li Xiaohe, Cai Yongqiang, Dai Lina, Xu Yangyang, Wang Yidan, Xu Xuebin. Digital measurements of the anatomical parameters of pedicle-rib unit screw fixation in thoracic vertebrae of preschoolers [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(6): 869-876. |
| [6] | Sun Jian, Fang Chao, Gao Fei, Wei Laifu, Qian Jun. Clinical efficacy and complications of short versus long segments of internal fixation for the treatment of degenerative scoliosis: a meta-analysis [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(3): 438-445. |
| [7] | Xu Yangyang, Zhang Kai, Li Zhijun, Zhang Yunfeng, Su Baoke, Wang Xing, Wang Lidong, Wang Yidan, He Yujie, Li Kun, Wang Haiyan, Li Xiaohe. Morphological analysis of optimal selection of lumbar pedicle screws in adolescents aged 12-15 years [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(21): 3321-3328. |
| [8] | Guo Yajing, Huang Yan, Shi Yin. Autophagy and epigenetic modification in inflammatory bowel disease [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(20): 3269-3274. |
| [9] | Qin Haikuo, Luo Shixing. Correlation of cortical bone thickness and X-ray gray value in different planes of proximal femur with brittle fracture of female hip [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(18): 2867-2872. |
| [10] | Huang Tianji, Yang Shengdong, Lin Hao, Zhang Chunyang, Deng Zhongqi, Zhong Weiyang, Luo Xiaoji. Mapping knowledge domains of bibliometrics regarding percutaneous vertebroplasty and percutaneous kyphoplasty based on VOSviewer [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(15): 2410-2417. |
| [11] | Qu Renfei, Mai Yuying, Chen Xiaowei, Hu Huanying, Chen Guozhi, Li Dongdong, Liao Hongbing. Relationship between lactic acid concentration and osteoclast differentiation of mouse monocytes [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(14): 2177-2183. |
| [12] | Song Xudong, He Yunwu, Li Yonglin, Chen Jing, Hu Junlan. Ultrasound-guided paravertebral nerve block for zoster-associated pain: a Meta-analysis [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(11): 1797-1804. |
| [13] | Zhong Yanping, Zou Hongyan, Quan Zhanrou, Deng Zhihui, Hong Wenxu. Analysis of full-length sequence and 18 point mutations of HLA-B in a leukemia patient and her family [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(1): 77-82. |
| [14] | Liu Yapu, Lin Junyu, Yang Zhou, Wu Xiuhua, Wu Xiaoliang, Zhu Qing’an. Three-dimensional visualization and quantitative analysis of microvessels in rat cervical spinal cord using barium sulfate perfusion combined with micro-CT scanning [J]. Chinese Journal of Tissue Engineering Research, 2019, 23(36): 5836-5840. |
| [15] | Gao Yangyang, Che Xianda, Han Pengfei, Liang Bin, Li Pengcui. Clinical efficacy of unicompartmental knee arthroplasty between robotic-assisted and conventional manual methods: a meta-analysis [J]. Chinese Journal of Tissue Engineering Research, 2019, 23(36): 5889-5895. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||