Chinese Journal of Tissue Engineering Research ›› 2026, Vol. 30 ›› Issue (24): 6382-6389.doi: 10.12307/2026.190
Previous Articles Next Articles
Multi-omics approach unveils novel therapeutic targets for osteoporosis: integrated analysis of Asian and European gene-tissue expression consortium data
Chen Yongxi
- The First Affiliated Hospital of Guangxi University of Chinese Medicine, Nanning 530003, Guangxi Zhuang Autonomous Region, China
-
Received:2025-05-15Revised:2025-08-28Online:2026-08-28Published:2026-02-05 -
Contact:Chen Yongxi, the First Affiliated Hospital of Guangxi University of Chinese Medicine, Nanning 530003, Guangxi Zhuang Autonomous Region, Guangxi Province, China -
About author:Chen Yongxi, Master’s supervisor, Associate chief physician, the First Affiliated Hospital of Guangxi University of Chinese Medicine, Nanning 530003, Guangxi Zhuang Autonomous Region, Guangxi Province, China -
Supported by:Guangxi Project for the Development and Promotion of Appropriate Technologies in Traditional Chinese Medicine, No. GZSY-23-28 (to CYX); University-Level Scientific Research Project of Guangxi University of Chinese Medicine, No. 2022MS043 (to CYX)
CLC Number:
Cite this article
Chen Yongxi. Multi-omics approach unveils novel therapeutic targets for osteoporosis: integrated analysis of Asian and European gene-tissue expression consortium data[J]. Chinese Journal of Tissue Engineering Research, 2026, 30(24): 6382-6389.
share this article
Add to citation manager EndNote|Reference Manager|ProCite|BibTeX|RefWorks
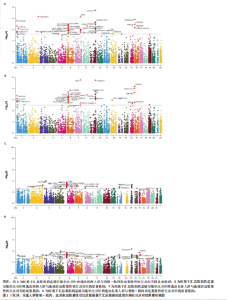
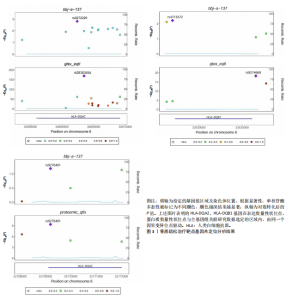
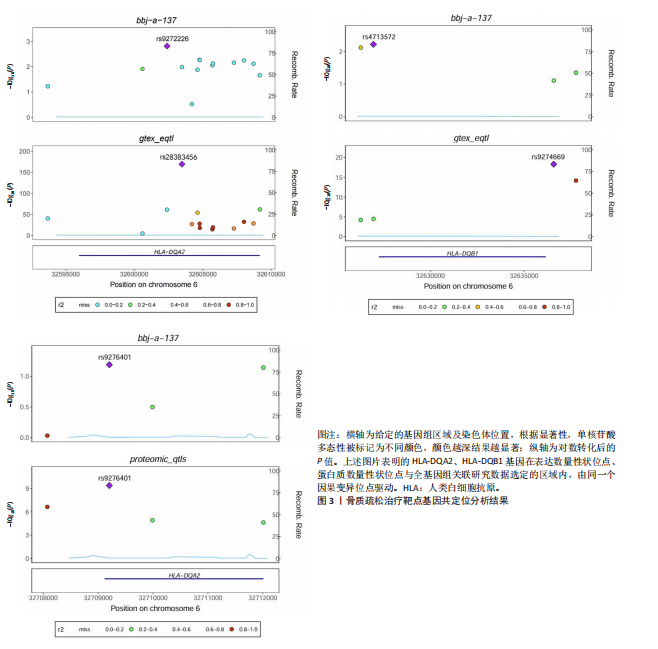
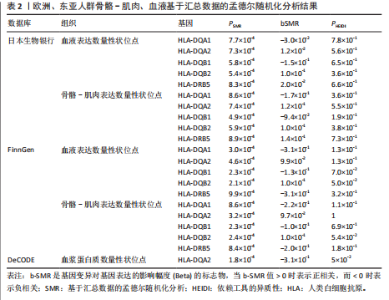
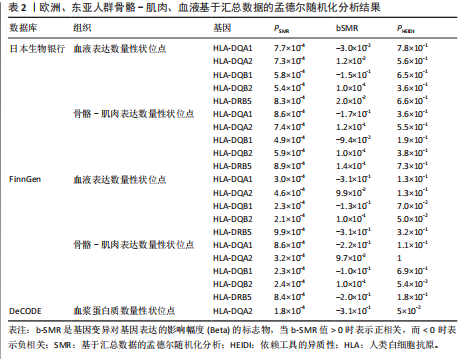
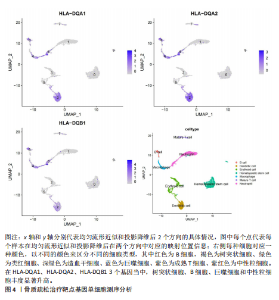
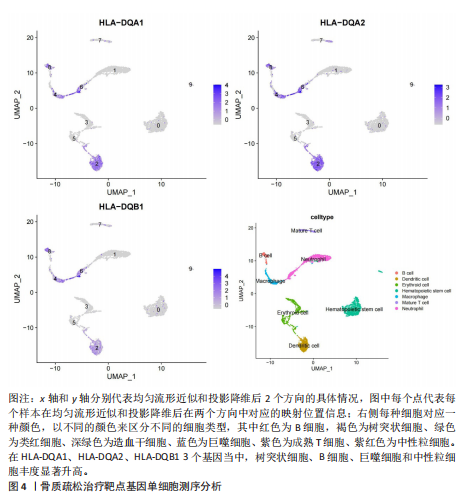
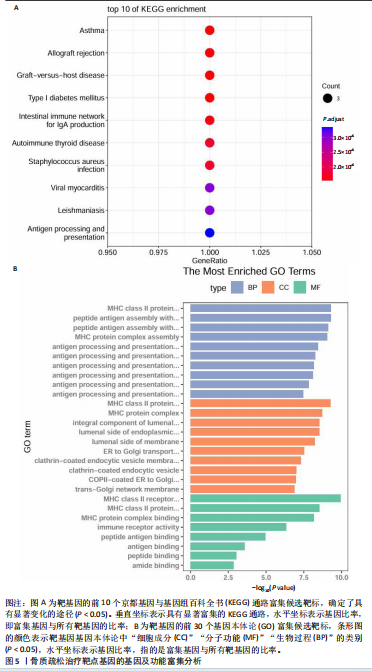
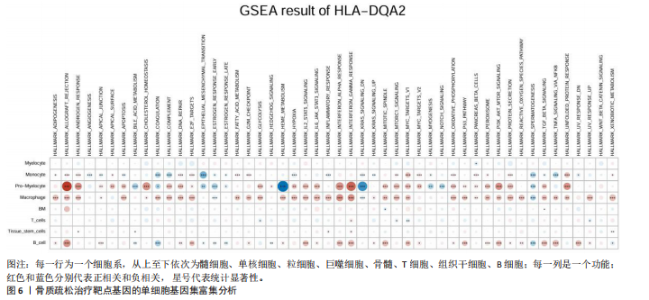
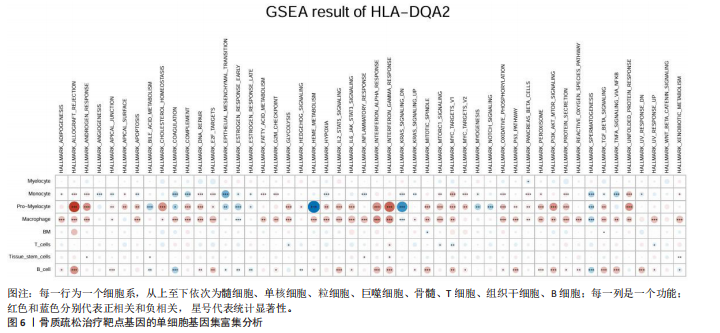
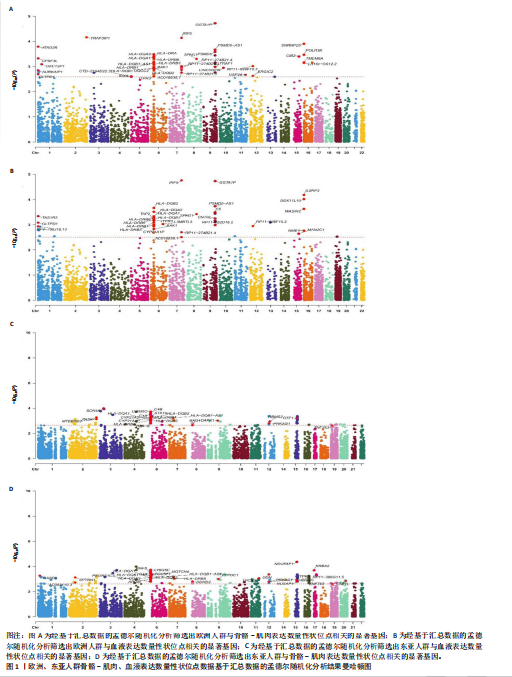
2.1 表达数量性状位点分析结果 去除掉重复的基因后,从FinnGen数据库筛选出了34个与骨质疏松症具有显著因果关系的基因(PFDR < 0.05,PHEIDI > 0.05);从日本生物银行数据库筛选出了30个与骨质疏松症具有显著因果关系的基因(PFDR < 0.05,PHEIDI > 0.05)。进行了全面的文献检索,并将尚未报道过与骨质疏松症相关的基因以及在2个数据库分析结果中共有的基因作为显著结果,最后筛选出的5个基因分别为人类白细胞抗原(human leukocyte antigen,HLA)等位基因HLA-DQA1、HLA-DQA2、HLA-DRB5、HLA-DQB1、HLA-DQB2。详见图1,2及表2。 2.2 蛋白质数量性状位点验证 为了进一步确定筛选出的基因作为骨质疏松症治疗靶点的可能性,使用血浆蛋白质数量性状位点数据进行验证。在初步分析中,仅有基因HLA-DQA2在蛋白质水平上得到了验证,详见表2。 2.3 共定位分析结果 筛选出的5个基因中,HLA-DRB5、HLA-DQA1和HLA-DQB2无完整的摘要水平数据可用,因此无法进行共定位分析。其余2个基因HLA-DQA2(PPH4=0.98)和HLA-DQB1(PPH4 =0.98)在基因水平上具有共定位证据,表明上述基因与骨质疏松症风险之间存在共同因果变异;其中HLA-DQA2在蛋白质水平上同样具有共定位证据的支持(COLOC PPH4=0.80),共定位结果的区域关联详见图3。 2.4 骨质疏松症中细胞类型鉴定 骨质疏松症单细胞数据集分为7个细胞群,分别为B细胞、树突状细胞、类红细胞、造血干细胞、巨噬细胞、成熟T细胞、中性粒细胞;与其他细胞类型相比,树突状细胞、B细胞、巨噬细胞和中性粒细胞在骨质疏松免疫微环境中的丰度显著更高,详见图4。 2.5 基因富集分析及功能富集分析 对从基于汇总数据的孟德尔随机化分析中鉴定的基因进行功能富集分析;根据归一化富集评分,显示前10个KEGG通路和前30个基因本体论条目,详见图5。 2.6 单细胞基因集富集分析 结果显示,基因HLA-DQA2在T细胞及干扰素γ途径上活化,在Kras基因、亚铁血红素途径上抑制,详见图6。"
| [1] TROJNIAK J, SENDERA A, BANAŚ-ZĄBCZYK A, et al. The MicroRNAs in the Pathophysiology of Osteoporosis. Int J Mol Sci. 2024;25(11):6240. [2] ANAM AK, INSOGNA K. Update on Osteoporosis Screening and Management. Med Clin North Am. 2021;105:1117-1134. [3] MORRIS JA, KEMP JP, YOULTEN SE, et al. An atlas of genetic influences on osteoporosis in humans and mice. Nat Genet. 2019;51:258-266. [4] ESTRADA K, STYRKARSDOTTIR U, EVANGELOU E, et al. Genome-wide meta-analysis identifies 56 bone mineral density loci and reveals 14 loci associated with risk of fracture. Nat Genet. 2012;44:491-501. [5] KEMP JP, MORRIS JA, MEDINA-GOMEZ C, et al. Identification of 153 new loci associated with heel bone mineral density and functional involvement of GPC6 in osteoporosis. Nat Genet. 2017;49:1468-1475. [6] PIVIDORI M, RAJAGOPAL PS, BARBEIRA A, et al. PhenomeXcan: mapping the genome to the phenome through the transcriptome. Sci Adv. 2020;6:eaba2083. [7] GU XJ, SU WM, DOU M, et al. Expanding causal genes for Parkinson’s disease via multi-omics analysis. NPJ Parkinsons Dis. 2023;9(1):146. [8] AL-BARGHOUTHI BM, ROSENOW WT, DU KP, et al. Transcriptome-wide association study and eQTL colocalization identify potentially causal genes responsible for human bone mineral density GWAS associations. Elife. 2022;11:e77285. [9] SHIN HD, PARK BL, SHIN HJ, et al. Association of KCNQ1 polymorphisms with the gestational diabetes mellitus in Korean women. J Clin Endocrinol Metab. 2010;95(1):445-449. [10] CAO M, ZHANG L, CHEN T, et al. Genetic susceptibility to gestational diabetes mellitus in a Chinese population. Front Endocrinol (Lausanne). 2020;11:247. [11] YANG Y, HU P, ZHANG Q, et al. Single-cell and genome-wide Mendelian randomization identifies causative genes for gout. Arthritis Res Ther. 2024; 26(1):114. [12] WU J, SUN X, WU C, et al. Single-cell transcriptome analysis reveals liver injury induced by glyphosate in mice. Cell Mol Biol Lett. 2023;28(1):11. [13] JACOBS BM, TAYLOR T, AWAD A, et al. Summary-data-based Mendelian randomization prioritizes potential druggable targets for multiple sclerosis. Brain Commun. 2020;2(2):fcaa119. [14] GTEX CONSORTIUM. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science. 2020;369(6509):1318-1330. [15] LYON MS, ANDREWS SJ, ELSWORTH B, et al. The variant call format provides efficient and robust storage of GWAS summary statistics. Genome Biol. 2021;22(1):32. [16] ISHIGAKI K, AKIYAMA M, KANAI M. Large-scale genome-wide association study in a Japanese population identifies novel susceptibility loci across different diseases. Nat Genet. 2020;52(7): 669-679. [17] FERKINGSTAD E, SULEM P, ATLASON BA, et al. Large-scale integration of the plasma proteome with genetics and disease. Nat Genet. 2021; 53(12):1712-1721. [18] SKRIVANKOVA VW, RICHMOND RC, WOOLF BAR, et al. Strengthening the reporting of observational studies in epidemiology using Mendelian randomization: the STROBE-MR statement. JAMA. 2021;326(16):1614-1621. [19] WU Y, ZHANG CY, WANG L, et al. Genetic Insights of Schizophrenia via Single Cell RNA-Sequencing Analyses. Schizophr Bull. 2023;49(4):914-922. [20] BAI Y, WANG J, FENG X, et al. Identification of drug targets for Sjögren’s syndrome: multi-omics Mendelian randomization and colocalization analyses. Front Immunol. 2024;15:1419363. [21] ZHU Z, ZHANG F, HU H, et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat Genet. 2016;48(5):481-487. [22] WU Y, ZENG J, ZHANG F, et al. Integrative analysis of omics summary data reveals putative mechanisms underlying complex traits. Nat Commun. 2018;9(1):918. [23] HUANG X, SHEN R, ZHENG Z. Unraveling genetic threads: Identifying novel therapeutic targets for allergic rhinitis through Mendelian randomization. World Allergy Organ J. 2024; 17(7):100927. [24] WANG Z, LI X, YANG J, et al. Single-cell RNA sequencing deconvolutes the in vivo heterogeneity of human bone marrow-derived mesenchymal stem cells. Int J Biol Sci. 2021; 17(15):4192-4206. [25] CAO G, XUAN X, LI Y, et al. Single-cell RNA sequencing reveals the vascular smooth muscle cell phenotypic landscape in aortic aneurysm. Cell Commun Signal. 2023;21(1):113. [26] SRIVASTAVA RK, SAPRA L, MISHRA PK. Osteometabolism: Metabolic Alterations in Bone Pathologies. Cells. 2022;11(23):3943. [27] CHAPMAN NM, BOOTHBY MR, CHI H. Metabolic coordination of T cell quiescence and activation. Nat Rev Immunol. 2020;20(1):55-70. [28] SCHIRMER M, SMEEKENS SP, VLAMAKIS H, et al. Linking the Human Gut Microbiome to Inflammatory Cytokine Production Capacity. Cell. 2016;167(4):1125-1136.e8. [29] YASUDA H. Discovery of the RANKL/RANK/OPG system. J Bone Miner Metab. 2021;39(1):2-11. [30] SRIVASTAVA S, RASOOL M. Genetics, epigenetics and autoimmunity constitute a Bermuda triangle for the pathogenesis of rheumatoid arthritis. Life Sci. 2024;357: 123075. [31] 中山大学.中华民族人MHC(HLA)二类基因多态性及其与疾病相关性研究.2006. [32] 北京市结核病胸部肿瘤研究所. HLA-DR、DQ基因与Ⅰ型糖尿病(IDDM)易感性研究.2002. [33] ZHANG Z, CHEN H, WANG XP, et al. Association between major histocompatibility complex class II molecules and postmenopausal osteoporosis. Chin J Osteoporos. 2024;30(2):270-274. [34] LIU H, HE J, BAGHERI-YARMAND R, et al. Osteocyte CIITA aggravates osteolytic bone lesions in myeloma. Nat Commun. 2022;13(1):3684. [35] BENASCIUTTI E, MARIANI E, OLIVA L, et al. MHC class II transactivator is an in vivo regulator of osteoclast differentiation and bone homeostasis co-opted from adaptive immunity. J Bone Miner Res. 2014;29(2):290-303. [36] WANG X, ZHANG X, HAN Y, et al. Role of the major histocompatibility complex class II protein presentation pathway in bone immunity imbalance in postmenopausal osteoporosis. Front Endocrinol (Lausanne). 2022;13:876067. [37] SUN X, CHEN B, QI Y, et al. Multi-omics Mendelian randomization integrating GWAS, eQTL and pQTL data revealed GSTM4 as a potential drug target for migraine. J Headache Pain. 2024;25(1):117. [38] MA Y, ZHOU Y, JIANG D, et al. Integration of human organoids single-cell transcriptomic profiles and human genetics repurposes critical cell type-specific drug targets for severe COVID-19. Cell Prolif. 2024;57(3):e13558. |
| [1] | Chen Huiting, Zeng Weiquan, Zhou Jianhong, Wang Jie, Zhuang Congying, Chen Peiyou, Liang Zeqian, Deng Weiming. Tail anchoring technique of vertebroplasty in treatment of osteoporotic vertebral compression fractures with intravertebral cleft: a finite element analysis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(9): 2145-2152. |
| [2] | Zeng Xuan, Weng Rui, Ye Shicheng, Tang Jiadong, Mo Ling, Li Wenchao. Two lumbar rotary manipulation techniques in treating lumbar disc herniation: a finite element analysis of biomechanical differences [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(9): 2153-2161. |
| [3] | Cheng Qisheng, Julaiti·Maitirouzi, Xiao Yang, Zhang Chenwei, Paerhati·Rexiti. Finite element analysis of novel variable-diameter screws in modified cortical bone trajectory of lumbar vertebrae [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(9): 2162-2171. |
| [4] | Liu Wenlong, Dong Lei, Xiao Zhengzheng, Nie Yu. Finite element analysis of tibial prosthesis loosening after fixed-bearing unicompartmental knee arthroplasty for osteoporosis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(9): 2191-2198. |
| [5] | Chen Long, Wang Xiaozhen, Xi Jintao, Lu Qilin. Biomechanical performance of short-segment screw fixation combined with expandable polyetheretherketone vertebral body replacement in osteoporotic vertebrae [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(9): 2226-2235. |
| [6] | Hu Xiongke, Liu Shaohua, Tan Qian, Liu Kun, Zhu Guanghui. Shikonin intervention with bone marrow mesenchymal stem cells improves microstructure of femur in aged mice [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(7): 1609-1615. |
| [7] | Wen Guangwei, Zhen Yinghao, Zheng Taikeng, Zhou Shuyi, Mo Guoye, Zhou Tengpeng, Li Haishan, Lai Yiyi. Effects and mechanisms of isoginkgetin on osteoclastogenesis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(6): 1348-1358. |
| [8] | Wu Zhilin, , He Qin, Wang Pingxi, Shi Xian, Yuan Song, Zhang Jun, Wang Hao . DYRK2: a novel therapeutic target for rheumatoid arthritis combined with osteoporosis based on East Asian and European populations [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(6): 1569-1579. |
| [9] | Zhang Haiwen, Zhang Xian, Xu Taichuan, Li Chao. Bibliometric and visual analysis of the research status and trends of senescence in osteoporosis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(6): 1580-1591. |
| [10] | Huang Jie, Zeng Hao, Wang Wenchi, Lyu Zhucheng, Cui Wei. Visualization analysis of literature on the effect of lipid metabolism on osteoporosis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(6): 1558-1568. |
| [11] | Yang Zhijie, Zhao Rui, Yang Haolin, Li Xiaoyun, Li Yangbo, Huang Jiachun, Lin Yanping, Wan Lei, HuangHongxing. Postmenopausal osteoporosis: predictive values of muscle mass, grip strength, and appendicular skeletal muscle index [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(5): 1073-1080. |
| [12] | Zhou Jian, Zhang Tao, Zhou Weili, Zhao Xingcheng, Wang Jun, Shen Jie, Qian Li, Lu Ming. Effects of resistance training on quadriceps mass and knee joint function in patients with osteoporosis and sarcopenia [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(5): 1081-1088. |
| [13] | Cao Wenqi, Feng Xiuzhi, Zhao Yi, Wang Zhimin, Chen Yiran, Yang Xiao, Ren Yanling. Effect of macrophage polarization on osteogenesis-angiogenesis coupling in type 2 diabetic osteoporosis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(4): 917-925. |
| [14] | Zeng Hao, Sun Pengcheng, Chai Yuan, Huang Yourong, Zhang Chi, Zhang Xiaoyun. Association between thyroid function and osteoporosis: genome-wide data analysis of European populations [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(4): 1019-1027. |
| [15] | Wang Meng, Lu Tan, Li Minjie, Liu Zhicheng, Guo Xiaoyong. Finite element analysis of stress distribution of anchors at different implantation depths under different bone density conditions in rotator cuff tears [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(3): 561-569. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||