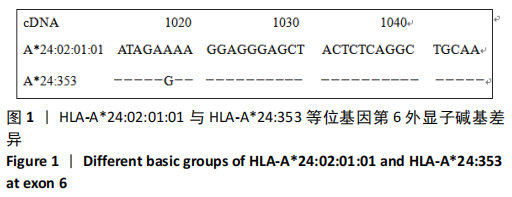
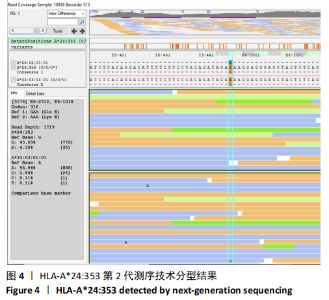
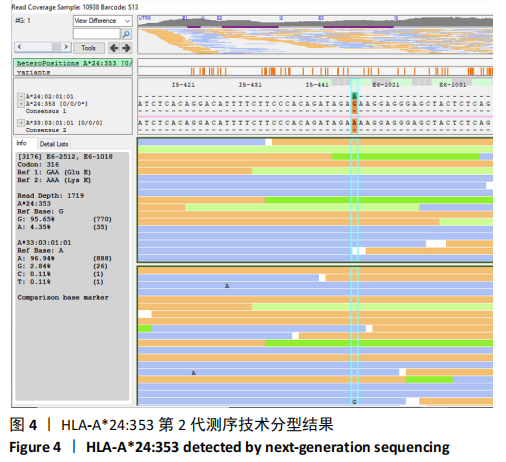
[1] GAO SQ, XU YP, HE LM, et al. The full length genomic sequence of a novel HLA-A*24 allele, HLA-A*24:353, identified in a patient with hepatitis B infection. HLA. 2017;89(5):304-305.
[2] MATTSSON J, RINGDÉN O, STORB R. Graft failure after allogeneic hematopoietic cell transplantation. Biol Blood Marrow Transplant. 2008;14(1 Suppl 1):165-170.
[3] RUBINSTEIN P, CARRIER C, SCARADAVOU A, et al. Outcomes among 562 recipients of placental-blood transplants from unrelated donors. N Engl J Med. 1998;339(22):1565-1577.
[4] HURLEY CK, BAXTER LOWE LA, et al. National Marrow Donor Program HLA-matching guidelines for unrelated marrow transplants. Biol Blood Marrow Transplant. 2003;9(10):610-615.
[5] MORISHIMA Y, SASAZUKI T, INOKO H, et al. The clinical significance of human leukocyte antigen (HLA) allele compatibility in patients receiving a marrow transplant from serologically HLA-A, HLA-B, and HLA-DR matched unrelated donors. Blood. 2002;99(11):4200-4206.
[6] APPS R, MENG Z, DEL PRETE GQ, et al. Relative expression levels of the HLA class-I proteins in normal and HIV-infected cells. J Immunol. 2015;194(8):3594-3600.
[7] FÜRST D, NEUCHEL C, TSAMADOU C, et al. HLA Matching in Unrelated Stem Cell Transplantation up to Date. Transfus Med Hemother. 2019;46(5):326-336.
[8] 刘晓华,迟晓云,焦淑贤.新等位基因HLA-A*24:233与HLA-A*26:89的分析确认[J].中国组织工程研究,2015,19(6):950-954.
[9] ULRICH S, POSCH U, HELMBERG W, et al. HLA-A*68:02:11, a new HLA-A*68 allele identified during family HLA typing. HLA. 2016;88(4):197-198.
[10] PARK Y, KIM H, IM J, et al. Identification of a novel allele, HLA-A*02:01:131, by full-length genomic sequencing. HLA. 2017;90(6):360-361.
[11] BALAS A, LÓPEZ-HERNÁNDEZ R, MORENO-HIDALGO MA, et al. Description of the new HLA-A*11:288 allele found in a healthy individual from the Canary Islands. HLA. 2018;92(4):239-240.
[12] KHOR SS, HITOMI Y, OMAE Y, et al. Detection of the novel HLA-A allele, HLA-A*02:01:01:58, in a Japanese individual. HLA. 2019;94(5):435-436.
[13] WANG F, WANG L, ZHANG W, et al. Identification of the novel HLA-A*02 allele, HLA-A*02:725. HLA. 2020;95(5):476-478.
[14] 鲍自谦,王大明,邓志辉,等.中国南方汉族人群中一个常见型新等位基因HLA-C*08:22的基因频率调查和分析[J].中国实验血液学杂志,2011,19(6): 950-954.
[15] ROBINSON J, BARKER DJ, GEORGIOU X, et al. IPD-IMGT/HLA Database. Nucleic Acids Res. 2020;48(D1):D948-D955.
[16] MACK SJ, CANO P, HOLLENBACH JA, et al. Common and well-documented HLA alleles: 2012 update to the CWD catalogue. Tissue Antigens. 2013;81(4):194-203.
[17] 安仕萍,闫莉娜.测序技术在人类白细胞抗原基因分型领域的应用进展[J].医学综述, 2007,13(15):1189-1191.
[18] HOSOMICHI K, SHIINA T, TAJIMA A, et al. The impact of next-generation sequencing technologies on HLA research. J Hum Genet. 2015;60(11):665-673.
[19] GOODWIN S, MCPHERSON JD, MCCOMBIE WR. Coming of age: ten years of next-generation sequencing technologies. Nat Rev Genet. 2016;17(6):333-351.
[20] 张弦,张艳玲,王建玲,等. HLA-A/B/C/DRB1/DQB1对非血缘造血干细胞移植影响的分析[J].中国实验血液学杂志,2011,19(1):143-148.
[21] GLUCKMAN E, FUENTE J, CAPPELLI B, et al. The role of HLA matching in unrelated donor hematopoietic stem cell transplantation for sickle cell disease in Europe. Bone Marrow Transplant. 2020. doi: 10.1038/s41409-020-0847-z. Online ahead of print.
[22] SABER W, OPIE S, RIZZO JD, et al. Outcomes after matched unrelated donor versus identical sibling hematopoietic cell transplantation in adults with acute myelogenous leukemia. Blood. 2012;119(17):3908-3916.
[23] PETERSDORF EW, MALKKI M, HOROWITZ MM, et al. Mapping MHC haplotype effects in unrelated donor hematopoietic cell transplantation. Blood. 2013; 121(10):1896-1905.
[24] ZHENG C, ZHU X, TANG B, et al. Comparative analysis of unrelated cord blood transplantation and HLA-matched sibling hematopoietic stem cell transplantation in children with high-risk or advanced acute leukemia. Ann Hematol. 2015;94(3): 473-480.
|