Chinese Journal of Tissue Engineering Research ›› 2026, Vol. 30 ›› Issue (25): 6455-6462.doi: 10.12307/2026.475
Previous Articles Next Articles
Transcriptome sequencing analysis of tibial transverse transport in a rabbit model of diabetic foot ulcers
Li Haojie1, Xie Tongliang1, Li Rui1, Sun Zuyan1, Deng Jiang1, Xu Lin1, Huang Wenliang2
- 1The Third Affiliated Hospital of Zunyi Medical University, Zunyi 563000, Guizhou Province, China; 2Kweichow Moutai Hospital, Renhuai 564500, Guizhou Province, China
-
Received:2025-10-15Revised:2026-03-09Online:2026-09-08Published:2026-04-17 -
Contact:Huang Wenliang, PhD candidate, Master’s supervisor, Chief physician, Kweichow Moutai Hospital, Renhuai 564500, Guizhou Province, China -
About author:Li Haojie, MS candidate, The Third Affiliated Hospital of Zunyi Medical University, Zunyi 563000, Guizhou Province, China -
Supported by:Guizhou Provincial Science and Technology Program Project, No. ZK[2021]General 387 (to HWL); Guizhou Provincial Clinical Special Project, No. LC[2024]019 (to HWL); Kweichow Moutai Hospital Project, No. MTyk2025-37 (to HWL)
CLC Number:
Cite this article
Li Haojie, Xie Tongliang, Li Rui, Sun Zuyan, Deng Jiang, Xu Lin, Huang Wenliang. Transcriptome sequencing analysis of tibial transverse transport in a rabbit model of diabetic foot ulcers[J]. Chinese Journal of Tissue Engineering Research, 2026, 30(25): 6455-6462.
share this article
Add to citation manager EndNote|Reference Manager|ProCite|BibTeX|RefWorks
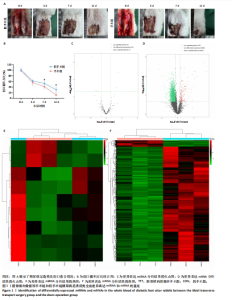
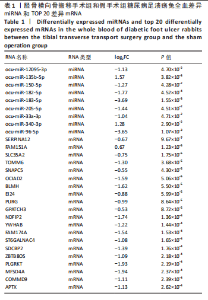
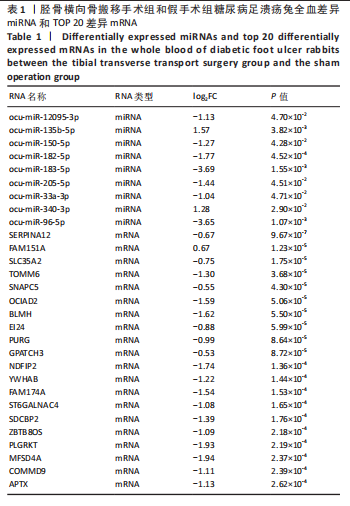
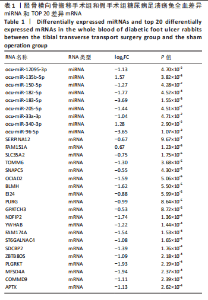
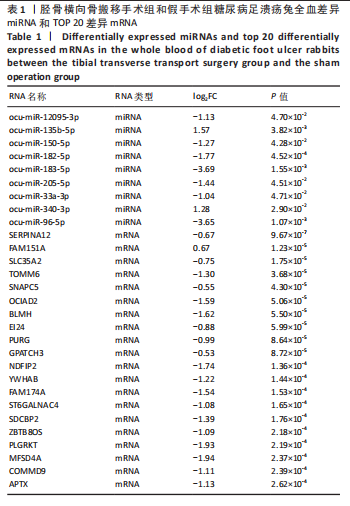
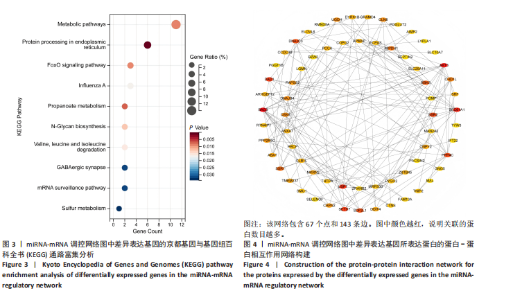
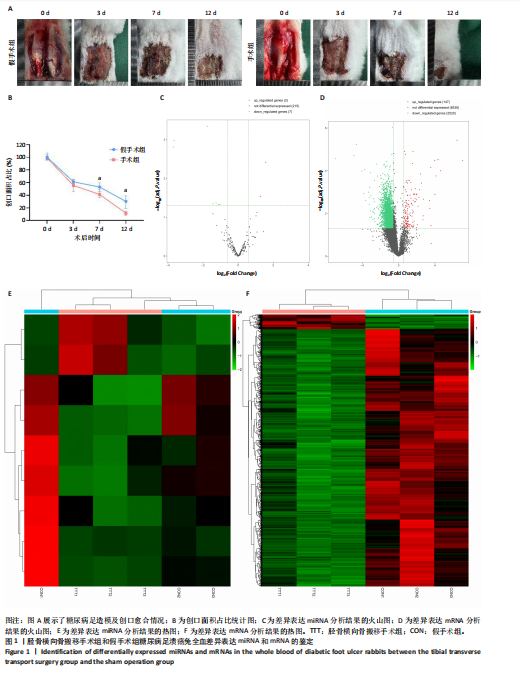
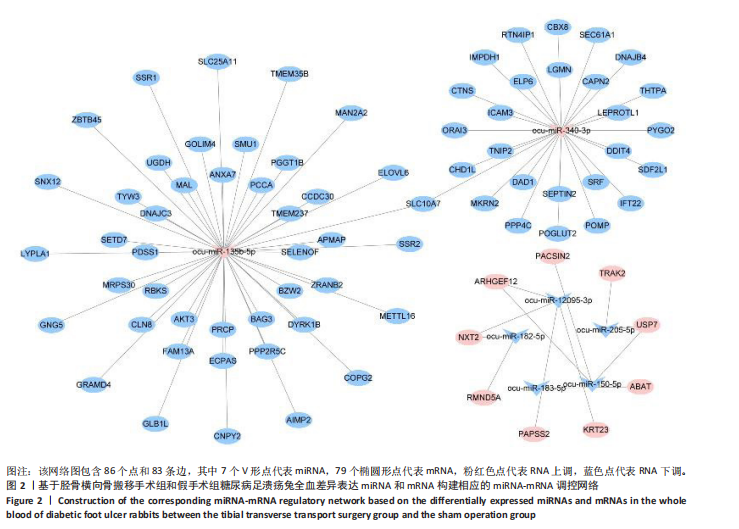
2.1 兔糖尿病足溃疡愈合情况、转录组测序以及全血差异miRNA和差异基因筛选 胫骨横向骨搬移术治疗显著促进了糖尿病足溃疡伤口的愈合,图1A,B。 采集手术组和假手术组兔外周血并进行全转录组测序。采用多层级质控体系确保数据可靠性。通过琼脂糖凝胶电泳和NanoDrop ND-1000分光光度计对总RNA样品进行完整性检测和纯度评估(A260/A280≈1.8-2.0,A260/A230 > 1.8)。文库构建阶段采用链特异性建库方法(KAPA Stranded RNA-Seq Kit),经Agilent 2100 Bioanalyzer验证文库片段分布(400-600 bp)并通过qPCR精确定量。测序数据质控显示所有样本Q30碱基比例均> 94%(Illumina NovaSeq 6000平台,PE150),比对率达62.93%-83.31%(Hisat2比对至OryCun2参考基因组)。 根据测序结果,以adj.P < 0.05和|logFC|≥0.5为筛选阈值,共鉴定出9个显著差异表达的miRNA和2 667个显著差异表达的mRNA,其中7个miRNA(ocu-miR-12095-3p,ocu-miR-150-5p,ocu-miR-182-5p,ocu-miR-183-5p,ocu-miR-205-5p,ocu-miR-33a-3p,ocu-miR-96-5p)和2 520个mRNA(GLIPR2,CLTA等)在手术组显著下调;2个miRNA(ocu-miR-135b-5p,ocu-miR-340-3p)和147个mRNA(SLC43A2,TRAK2等)在手术组显著上调。图1C,D分别展示了差异miRNA和差异mRNA分析结果的火山图。图1E,F分别展示了差异miRNA和差异mRNA分析结果的热图。表1展示了差异miRNA和TOP 20差异mRNA的详细信息。 2.2 miRNA-mRNA调控网络的构建 使用TargetScan和miRNADA软件预测miRNA的靶基因。将差异分析结果数据和预测的靶基因结果整合并使用Cytoscape软件(V3.10.0)构建miRNA-mRNA调控网络(图2)。该网络图包含86个点和83条边,其中7个V形点代表miRNA,79个椭圆形点代表mRNA,粉红色点代表RNA上调,蓝色点代表RNA下调。 根据miRNA-mRNA调控网络,SL10A7 (miR-135b-5p,miR-340-3p)、ARHGEF12(miR-12095-3p,miR-150-5p)、NXT2(miR-12095-3p,miR-182-5p)、USP7 (miR-205-5p,miR-150-5p)同时被2个miRNA调控。miR-12095-3p(ARHGEF12,NXT2,PAPSS2)和miR-150-5p(ARHGEF12,PACSIN2,USP7)同时调控3个靶基因。miR-182-5p (NXT2,RMND5A)和miR-205-5p(TRAK2,USP7)同时调控2个靶基因。 2.3 靶基因富集分析 利用KOBAS在线工具对miRNA-mRNA调控网络图中的79个靶基因进行京都基因与基因组百科全书富集分析,结果显示这些基因主要参与了代谢通路、内质网中的蛋白质加工、FoxO信号通路、丙酸代谢、N-聚糖生物合成等生物"
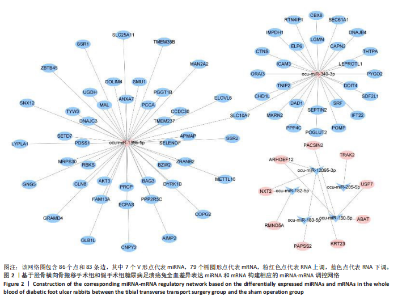
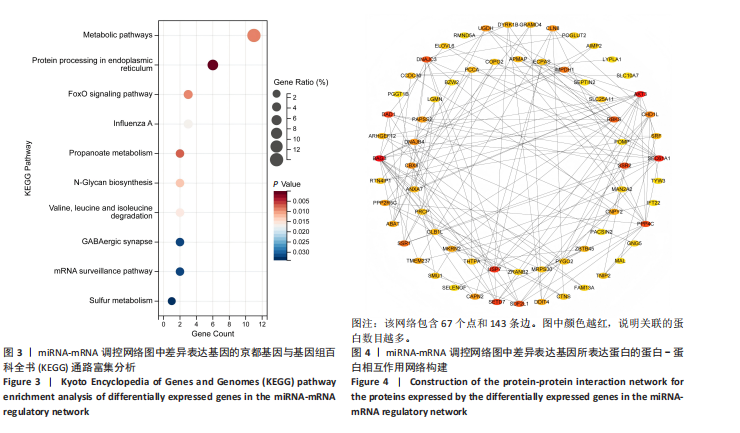
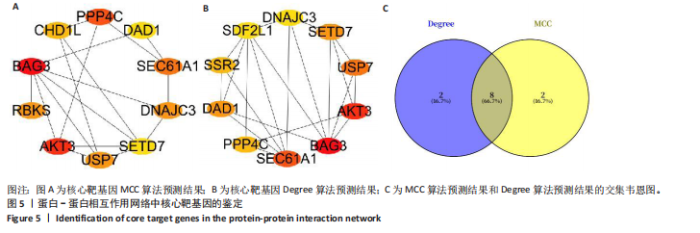
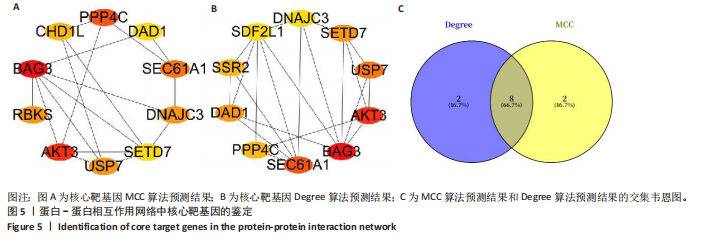
途径或信号通路(图3)。目前已有研究报道了p-FOXO在愈合组织中高表达,提示富集分析结果的准确性[20]。 2.4 蛋白-蛋白相互作用分析 利用String在线工具对miRNA-mRNA调控网络图中的79个靶基因进行蛋白-蛋白相互作用分析。使用Cytoscape软件进行可视化分析,得到了一个包含67个点和143条边的蛋白-蛋白相互作用调控网络图(图4)。图中颜色越红,说明关联的蛋白数目越多。 2.5 核心靶基因筛选 使用Cytohubba插件中的MCC和Degree算法鉴定网络图中的核心基因(图5A,B),最终BAG3、AKT3、PPP4C、SEC61A1、DNAJC3、USP7、DAD1、SETD7同时被两种算法鉴定为核心靶基因(图5C)。"
| [1] WANG X, YUAN CX, XU B, et al. Diabetic foot ulcers: Classification, risk factors and management. World J Diabetes. 2022;13(12):1049-1065. [2] 简喜超,邵靖杰,唐诗涵,等.外泌体促进糖尿病创面愈合:研究热点及演化趋势的可视化分析[J].中国组织工程研究,2026,30(19):5072-5081. [3] EDMONDS M, MANU C, VAS P. The current burden of diabetic foot disease. J Clin Orthop Trauma. 2021;17:88-93. [4] BURGESS JL, WYANT WA, ABDO ABUJAMRA B, et al. Diabetic Wound-Healing Science. Medicina (Kaunas). 2021;57(10):1072. [5] SHANG J, CHENG L, OU Q. Tibial cortex transverse transport surgery to treat diabetic foot ulcerations: what mechanism is involved in accelerated wound healing? Int J Surg. 2025;111(2):2323-2324. [6] ILIZAROV GA. The tension-stress effect on the genesis and growth of tissues. Part I. The influence of stability of fixation and soft-tissue preservation. Clin Orthop Relat Res. 1989;(238):249-281. [7] LIU Z, XU C, YU YK, et al. Twenty Years Development of Tibial Cortex Transverse Transport Surgery in PR China. Orthop Surg. 2022;14(6):1034-1048. [8] 李超,李娟,王栋,等.胫骨横向骨搬移重建糖尿病足微循环的研究进展[J].中华骨与关节外科杂志,2025,18(4):362-367. [9] 冼呈,赵劲民,苏伟,等.外固定架骨搬移系统修复糖尿病足:功能与影像学评价[J].中国组织工程研究,2015,19(46):7539-7544. [10] HU XX, XIU ZZ, LI GC, et al. Effectiveness of transverse tibial bone transport in treatment of diabetic foot ulcer: A systematic review and meta-analysis. Front Endocrinol (Lausanne). 2023;13:1095361. [11] 柏小金,徐林,刘福英,等.基于HIF-1α/VEGF信号通路探讨Ilizarov技术治疗糖尿病足溃疡的机制[J].中国老年学杂志,2023,43(21):5300-5303. [12] HO PTB, CLARK IM, LE LTT. MicroRNA-Based Diagnosis and Therapy. Int J Mol Sci. 2022;23(13):7167. [13] SHANG R, LEE S, SENAVIRATHNE G, et al. microRNAs in action: biogenesis, function and regulation. Nat Rev Genet. 2023;24(12):816-833. [14] ZHAN C, LIU K, ZHANG Y, et al. Myocardial infarction unveiled: Key miRNA players screened by a novel lncRNA-miRNA-mRNA network model. Comput Biol Med. 2023;160:106987. [15] 陈霞,商宇伟,王林晓,等.维生素D缺乏模型小鼠子宫转录组的测序分析[J].中国组织工程研究,2022,26(26):4101-4106. [16] 涂明洋,李小磊,杨坚.自体PRP对糖尿病兔横向骨搬移术后局部血管再生及血管生长相关因子影响的实验研究[J].农垦医学,2024,46(6): 492-497. [17] YANG Y, LI Y, PAN Q, et al. Tibial cortex transverse transport accelerates wound healing via enhanced angiogenesis and immunomodulation. Bone Joint Res. 2022;11(4):189-199. [18] CHEN Y, CHEN L, LUN ATL, et al. edgeR v4: powerful differential analysis of sequencing data with expanded functionality and improved support for small counts and larger datasets. Nucleic Acids Res. 2025;53(2):gkaf018. [19] BU D, LUO H, HUO P, et al. KOBAS-i: intelligent prioritization and exploratory visualization of biological functions for gene enrichment analysis. Nucleic Acids Res. 2021;49(W1):W317-W325. [20] GARCÍA CORONADO PL, FRANCO MOLINA MA, ZÁRATE TRIVIÑO DG, et al. Exosomes isolated from IMMUNEPOTENT CRP, a hemoderivative, to accelerate diabetic wound healing. Front Bioeng Biotechnol. 2024; 12:1356028. [21] 田林, 时孝晴,段正兰,等.胫骨横向骨搬移术治疗糖尿病足疗效及安全性的Meta分析[J].中国组织工程研究,2021,25(20):3275-3280. [22] LIU N, DAI S, LIU D, et al. Application of Transverse Tibial Bone Transfer in Diabetic Foot Patients: A Comprehensive Evaluation of Postoperative Prognostic Factors and Risk Factors. Orthop Surg. 2025;17(9):2708-2716. [23] XU D, BAI C, HU R, et al. Exploring the Changes in IL-6 and Related Cytokines in Angiogenesis after Tibial Transverse Transplantation in Diabetic Foot Ulcers. Orthop Surg. 2024;16(9):2181-2190. [24] XIE J, LIU X, WU B, et al. Bone transport induces the release of factors with multi-tissue regenerative potential for diabetic wound healing in rats and patients. Cell Rep Med. 2024;5(6):101588. [25] YU L, ZHANG D, YIN Y, et al. Tibial cortex transverse transport surgery improves wound healing in patients with severe type 2 DFUs by activating a systemic immune response: a cross-sectional study. Int J Surg. 2025; 111(1):257-272. [26] QIN W, LIU K, SU H, et al. Tibial cortex transverse transport promotes ischemic diabetic foot ulcer healing via enhanced angiogenesis and inflammation modulation in a novel rat model. Eur J Med Res. 2024;29(1): 155. [27] FANG Y, SUN S, WU J, et al. Alterations in the Levels of Urinary Exosomal MicroRNA-183-5p and MicroRNA-125a-5p in Individuals with Type 2 Diabetes Mellitus. Biomedicines. 2024;12(11):2608. [28] XU Y, XU J, CHEN S, et al. Identifying potential pathogenesis and immune infiltration in diabetic foot ulcers using bioinformatics and in vitro analyses. BMC Med Genomics. 2023;16(1):313. [29] ZHU L, ZHONG Q, YANG T, et al. Improved therapeutic effects on diabetic foot by human mesenchymal stem cells expressing MALAT1 as a sponge for microRNA-205-5p. Aging (Albany NY). 2019;11(24):12236-12245. [30] YANG M, GU Y. LncRNA DLEU1 promotes angiogenesis in diabetic foot ulcer wound healing by regulating miR-96-5p. Ir J Med Sci. 2024;193(1):241-247. [31] JIANG H, QIN H, YANG Q, et al. Effective delivery of miR-150-5p with nucleus pulposus cell-specific nanoparticles attenuates intervertebral disc degeneration. J Nanobiotechnology. 2024;22(1):292. [32] SU X, LAI T, TAO Y, et al. miR-33a-3p regulates METTL3-mediated AREG stability and alters EMT to inhibit pancreatic cancer invasion and metastasis. Sci Rep. 2023;13(1):13587. [33] HAO J, CHEN Y, YU Y. Circular RNA circ_0008360 Inhibits the Proliferation, Migration, and Inflammation and Promotes Apoptosis of Fibroblast-Like Synoviocytes by Regulating miR-135b-5p/HDAC4 Axis in Rheumatoid Arthritis. Inflammation. 2022;45(1):196-211. [34] WU W, LI Q, ZHU Z, et al. HTR1D functions as a key target of HOXA10-AS/miR-340-3p axis to promote the malignant outcome of pancreatic cancer via PI3K-AKT signaling pathway. Int J Biol Sci. 2022;18(9):3777-3794. [35] FENG J, WANG Y, XIANG S, et al. Applying GC-MS based serum metabolomic profiling to characterize two traditional Chinese medicine subtypes of diabetic foot gangrene. Front Mol Biosci. 2024;11:1384307. [36] DENG P, LIANG H, WANG S, et al. Combined metabolomics and network pharmacology to elucidate the mechanisms of Dracorhodin Perchlorate in treating diabetic foot ulcer rats. Front Pharmacol. 2022;13:1038656. [37] XIAO Y, DONG X, CHEN C, et al. An integrated method for IgG N-glycans enrichment and analysis: Understanding the role of IgG glycosylation in diabetic foot ulcer. J Chromatogr B Analyt Technol Biomed Life Sci. 2024; 1233:123983. [38] LYTRIVI M, SENÉE V, SALPEA P, et al. DNAJC3 deficiency induces β-cell mitochondrial apoptosis and causes syndromic young-onset diabetes. Eur J Endocrinol. 2021;184(3):455-468. [39] ZHU Y, ZONG M, HU L, et al. USP7 deficiency promotes diabetic wound healing by repressing GATA3-mediated pro-inflammatory macrophage polarization. Mol Cell Endocrinol. 2025;599:112489. [40] MOHAMMED SA, GORICA E, ALBIERO M, et al. Targeting SETD7 Rescues Diabetes-Induced Impairment of Angiogenic Response by Transcriptional Repression of Semaphorin-3G. Diabetes. 2025;74(6):969-982. [41] LIANG Z, ZHANG S, ZOU Z, et al. Functional characterization of BAG3 in orange-spotted grouper (Epinephelus coioides) during viral infection. Fish Shellfish Immunol. 2022;122:465-475. [42] MARKUS T, LEY D, HANSSON SR, et al. Neuroprotective dobutamine treatment upregulates superoxide dismutase 3, anti-oxidant and survival genes and attenuates genes mediating inflammation. BMC Neurosci. 2018;19(1):9. [43] HUANG Q, CHU X, YANG C, et al. Bio-engineered microRNA-7 effectively interferes with the Akt3/p53 axis to suppress human non-small cell lung cancer. Cancer Cell Int. 2025;25(1):250. [44] HOU Q, YUAN J, LI S, et al. Autophagic degradation of DHCR7 activates AKT3 and promotes sevoflurane-induced hippocampal neuroinflammation in neonatal mice. Free Radic Biol Med. 2024;222:304-316. [45] HERRERA JL, KOMATSU M. Akt3 activation by R-Ras in an endothelial cell enforces quiescence and barrier stability of neighboring endothelial cells via Jagged1. Cell Rep. 2024;43(3):113837. [46] RAJA R, WU C, BASSOY EY, et al. PP4 inhibition sensitizes ovarian cancer to NK cell-mediated cytotoxicity via STAT1 activation and inflammatory signaling. J Immunother Cancer. 2022;10(12):e005026. [47] LIAO FH, SHUI JW, HSING EW, et al. Protein phosphatase 4 is an essential positive regulator for Treg development, function, and protective gut immunity. Cell Biosci. 2014;4:25. [48] CAO C, ZHOU S, HU J. Long noncoding RNA MAGI2-AS3/miR-218-5p/GDPD5/SEC61A1 axis drives cellular proliferation and migration and confers cisplatin resistance in nasopharyngeal carcinoma. Int Forum Allergy Rhinol. 2020;10(8):1012-1023. [49] FA X, SONG P, FU Y, et al. Long non-coding RNA VPS9D1-AS1 facilitates cell proliferation, migration and stemness in hepatocellular carcinoma. Cancer Cell Int. 2021;21(1):131. [50] MORI S, KIMURA R, MORIHARA H, et al. Suppression of Dad1 induces cardiomyocyte death by weakening cell adhesion. Am J Physiol Cell Physiol. 2025;328(1):C95-C106. |
| [1] | Wang Zhengye, Liu Wanlin, Zhao Zhenqun. Advance in the mechanisms underlying miRNAs in steroid-induced osteonecrosis of the femoral head [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(5): 1207-1214. |
| [2] | Luan Chuankai, Zhu Lei. Role and mechanism of exercise-regulated miRNAs in cardiac remodeling after acute myocardial infarction [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(34): 9024-9031. |
| [3] | Tian Yu, Guo Ying, Yiershatijiang Aniwaer, Aihematijiang Refuhaiti, Maierdana Maimaitireyimu. Molecular mechanisms by which Fusobacterium nucleatum regulates colonic polyp formation in mice [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(29): 7603-7611. |
| [4] | He Yanyan, Ge Jirong, Li Shengqiang, Chen Xuan, Huang Jingwen, Huang Xiaobin, Xue Lipeng. Transcriptomic analysis of expression and function of differential genes in traditional Chinese medicine syndromes of postmenopausal osteoporosis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(25): 6446-6454. |
| [5] | Wang Zhengye, Liu Wanlin, Zhao Zhenqun. Mechanisms of miRNAs involved in cartilage development: new strategies and targets [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(24): 6289-6296. |
| [6] | Wu Lingjie, Zheng Kaiyuan, Wang Guangrong, Yin Chong . Strategies for the application of miRNA-targeted therapy in the treatment of osteoporosis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(22): 5792-5803. |
| [7] | Zhong Zhuolan, Peng Zhina, Tian Xiaohong, Han Cuifei, Zhang Zihan, Chu Jiaqi. Acanthopanax exosome-like nanovesicles promote osteogenic differentiation of human bone marrow mesenchymal stem cells [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(19): 4825-4835. |
| [8] | Li Tianbo, Yu Zeyang, Qin Xinyuan, Wang Jiangning, Gao Lei. Exosomes derived from human umbilical cord mesenchymal stem cells in treatment of diabetic foot ulcers [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(19): 4897-4901. |
| [9] | Liu Yingzhao, Ma Yanxia, Lin Yaofa, Zhang Guoqiao, Miao Weiliang, Jia Yanli, Chi Chenshen, Song Wangsheng, Li Di, Liu Chenglong, Zhang Haonan. miR-9 regulates the differentiation of neural stem cells in mouse cerebral cortex [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(19): 4942-4948. |
| [10] | Liu Yuxuan, Guan Dongsheng, Wang Jing, Ren Yihan. Exosomal miRNA as an early diagnostic biomarker and potential therapeutic target for cerebral small vessel disease [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(19): 5000-5006. |
| [11] | Wang Xinyue, Li Hongli, Guo Chunhui, Chen Jibing, Yu Hua. Changes in the expression of six microRNAs in ovarian tissue from animal models of premature ovarian failure and in peripheral blood of patients with premature ovarian failure [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(18): 4675-4684. |
| [12] | Li Wenhui, Fan Weijing, Liu Guobin. Impact of Zi-Zhu ointment on the miRNA expression profile in mouse models of diabetic ulcers: a high-throughput sequencing analysis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(17): 4337-4346. |
| [13] | Wumiti·Taxi, Wang Lining, Li Muzhe, Sun Jie, Chen Shuangliu, Zhu Yihua, Zhou Shijie, Ma Yong, Guo Yang. Wenshen Tongluo Zhitong decoction regulates the bone fat differentiation balance of bone marrow mesenchymal stem cells through exosomal miR-342-3p [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(13): 3258-3269. |
| [14] | Jiang Yidi, Zhao Jianwei, Zhou Jue, Lyu Jinpeng, Wang Datao, Li Xunsheng, Yue Zhigang, Cui Bo, Sun Hongmei. Regulation of antler stem cell exosomes miRNA-145 on inflammatory chondrocytes [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(13): 3288-3297. |
| [15] | Lu Biqiong, Wei Zhongjian. Skeletal muscle-derived exosome-mediated regulation of bone formation and role of exercise intervention [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(13): 3379-3391. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||