Chinese Journal of Tissue Engineering Research ›› 2026, Vol. 30 ›› Issue (24): 6390-6399.doi: 10.12307/2026.199
Previous Articles Next Articles
Keloid pathogenesis is correlated with fibroblast heterogeneity genes: single-cell transcriptomic analysis based on GEO database
Guo Tao1, Liu Yuxin2, Yan Meirong1, Wang Xiaoni1
- 1Department of Burn and Plastic Surgery, 2Department of Dermatology, Ningxia Medical University General Hospital, Yinchuan 750001, Ningxia Hui Autonomous Region, China
-
Received:2025-07-22Revised:2025-09-16Online:2026-08-28Published:2026-02-05 -
Contact:Wang Xiaoni, Associate chief physician, Department of Burn and Plastic Surgery, Ningxia Medical University General Hospital, Yinchuan 750001, Ningxia Hui Autonomous Region, China -
About author:Guo Tao, Attending physician, Department of Burn and Plastic Surgery, Ningxia Medical University General Hospital, Yinchuan 750001, Ningxia Hui Autonomous Region, China -
Supported by:Ningxia Natural Science Foundation, No. 2023AAC03609 (to YMR)
CLC Number:
Cite this article
Guo Tao, Liu Yuxin, Yan Meirong, Wang Xiaoni. Keloid pathogenesis is correlated with fibroblast heterogeneity genes: single-cell transcriptomic analysis based on GEO database[J]. Chinese Journal of Tissue Engineering Research, 2026, 30(24): 6390-6399.
share this article
Add to citation manager EndNote|Reference Manager|ProCite|BibTeX|RefWorks
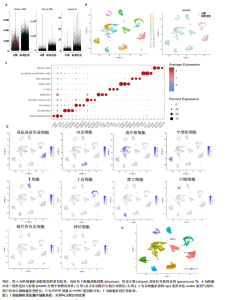
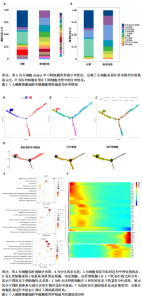
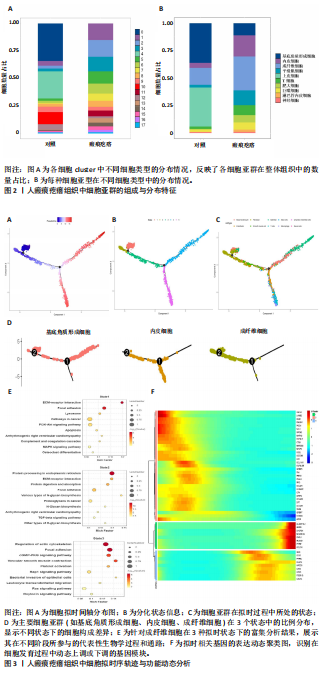
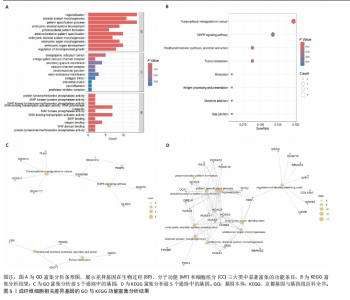
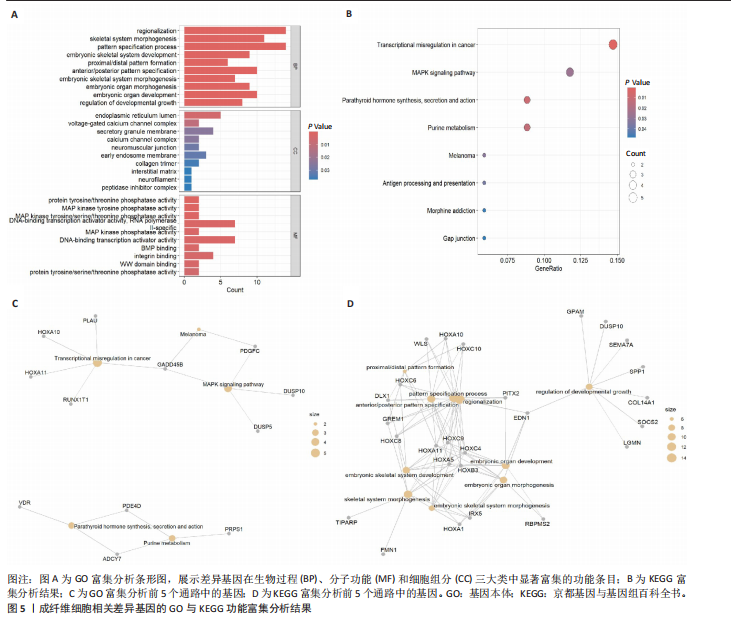
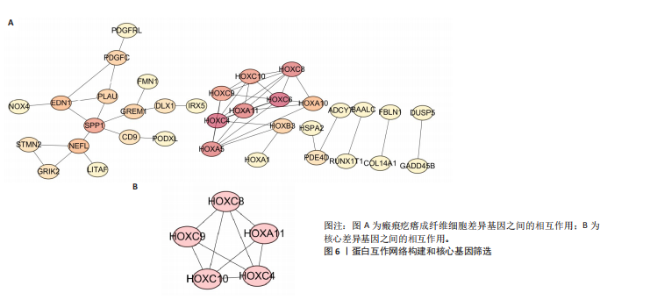
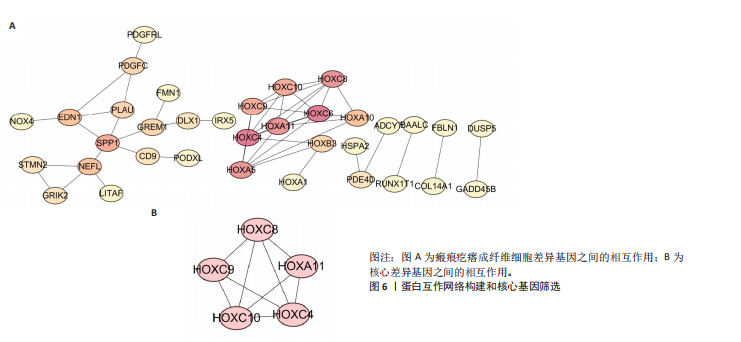
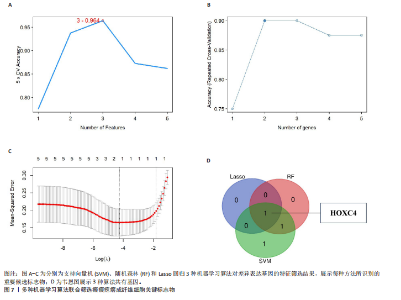
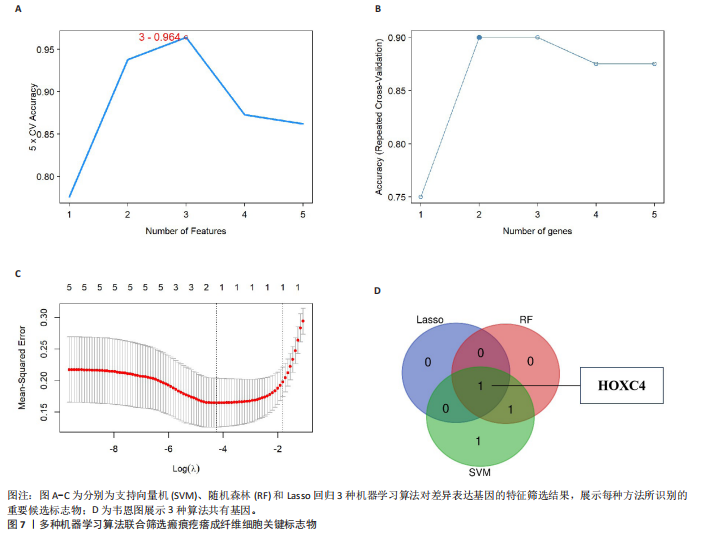
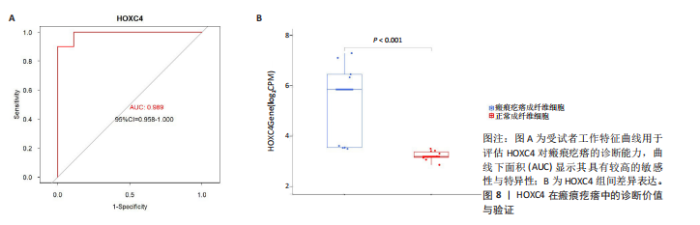
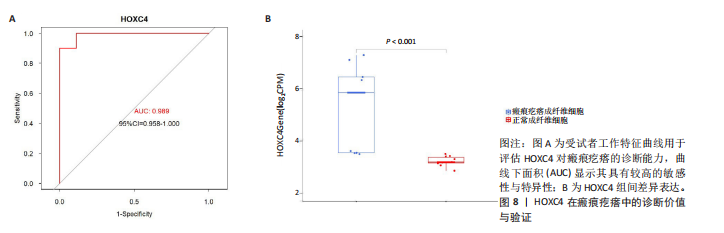
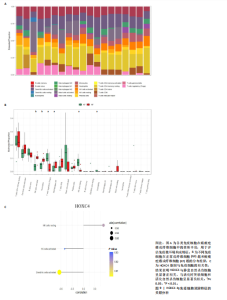
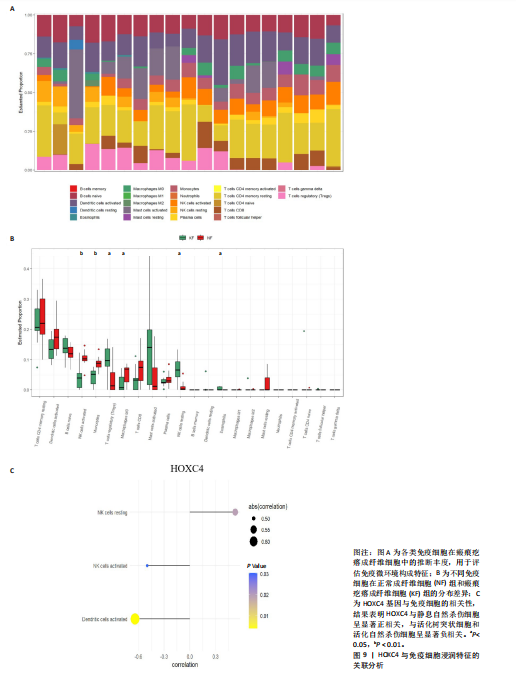
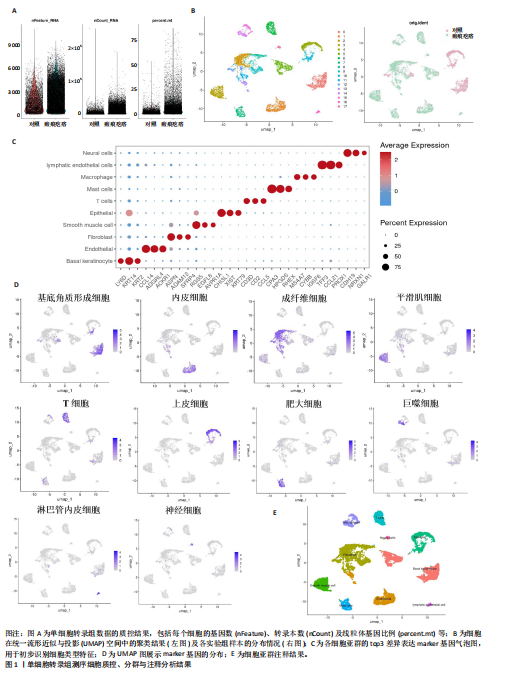
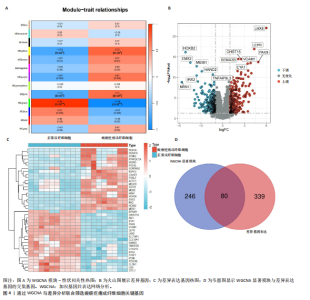
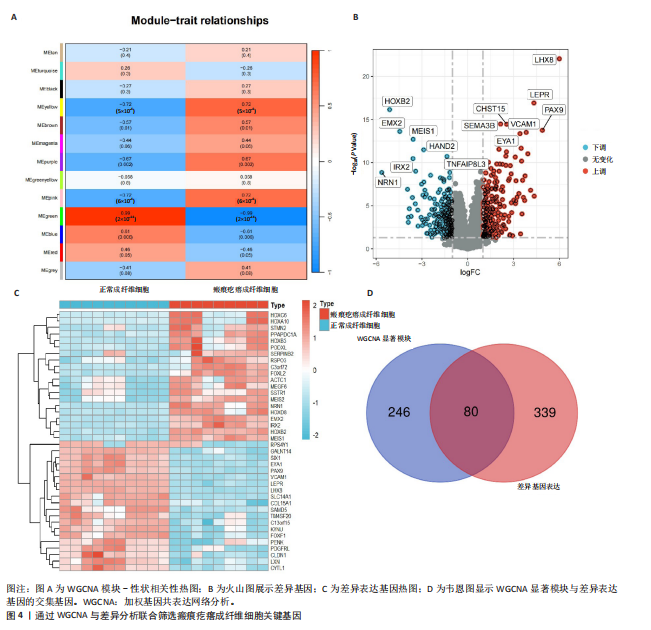
2.1 单细胞水平分析人瘢痕疙瘩的分子图谱 为了在单细胞分辨率下确定人瘢痕疙瘩组织的转录景观,对5个组织进行了单细胞转录组分析,经过严格的细胞过滤,收集了10 292个细胞(图1A),基于 Seurat 算法的无监督聚类,初步获得 18个细胞簇,使用UMAP算法展示了不同细胞类型在人瘢痕疙瘩及正常组织中的分布(图1B)。气泡图展示了每个细胞簇中前3个上调高表达基因(图1C)。通过相关marker基因成功注释出10个最终细胞簇,包括基底角质形成细胞(Basal keratinocyte,簇0,5)、内皮细胞(Endothelial,簇1,13)、成纤维细胞(Fibroblast,簇2,8,9,11,12)、平滑肌细胞(Smooth muscle cell,簇3)、T细胞(簇6)、上皮细胞(Epithelial,簇4,10)、肥大细胞(Mast cell,簇7)、巨噬细胞(Macrophage,14,15)、淋巴管内皮细胞(lymphatic endothelial cell,簇16)和神经细胞(Neural cells,簇17)(图1D,E)。 2.2 人瘢痕疙瘩细胞分布情况 为了进一步了解人瘢痕疙瘩组织中不同细胞的分布情况,分别展示了正常组织和人瘢痕疙瘩组织中各种细胞所占的比例。与对照组相比,内皮细胞、成纤维细胞、平滑肌细胞、T细胞、肥大细胞、巨噬细胞和淋巴管内皮细胞在人瘢痕疙瘩组织中的比例显著升高,差异有显著性意义(图2A,B)。尤其成纤维细胞在人瘢痕疙瘩中的数量为2 055,所占比例最大,在对照组中细胞数量为558,形成显著对比。 2.3 人瘢痕疙瘩亚群细胞拟时序分析 对所有亚群细胞进行拟时序分析,10种细胞主要分化为5种不同的状态(图3A-C)。其中基底角质形成细胞主要存在于状态4,位于分化终点;内皮细胞主要位于状态3和状态5,位于分化起点和分化过程中;成纤维细胞主要位于状态1、状态2和状态3,位于分化起点,具有较高的分化能力(图3D),状态1中的基因主要富集在细胞外基质受体相互作用、黏着、磷脂酰肌醇3激酶/蛋白激酶B信号通路等,提示成纤维细胞正处于早期激活阶段,通过基质感知和黏附信号启动迁移与激活行为;状态2中的基因主要富集在内质网中的蛋白质加工、细胞外基质受体相互作用、蛋白质消化和吸收等,推测该阶段对应成纤维细胞的高分泌活跃期,细胞正大量合成并分泌胶原与基质蛋白,是瘢痕形成的关键增生阶段;状态3中的基因主要富集在肌动蛋白细胞骨架的调节、环磷酸鸟苷-蛋白激酶 G信号通路和血管平滑肌收缩等,提示成纤维细胞已获得肌成纤维细胞特征,具备一定的收缩功能,可能对应瘢痕疙瘩发育中的成熟收缩期(图3E)。紧接着分析了在细胞分化过程中具有相似表达趋势的基因,其中FBLN2、LAMB2、ECM1等在分化起点显著高表达,ACADL、DHCR24和CCL7A1等在分化终点显著高表达,而S100A2、CXCL14、PTGDS等主要在决策分化过程中高度表达(图3F)。 2.4 加权基因共表达网络联合差异表达基因筛选瘢痕疙瘩成纤维细胞相关基因 下载转录组瘢痕疙瘩成纤维细胞数据集GSE145725,使用加权基因共表达网络鉴定出瘢痕疙瘩成纤维细胞相关的重要模块基因,最终得到13个模块,分别是“tan”“turquoise”“black”“yellow”“brown”“magenta”“purple”“greenyellow”“pink”“green”“blue”“red”和“grey”。其中黄色模块与瘢痕疙瘩成纤维细胞组(R=0.72,P=5×10-4)呈高度正相关;绿色模块与正常成纤维细胞组(R=0.99,P=2×10-14)呈高度正相关;因此黄色模块中的基因被认为与瘢痕疙瘩显著相关(图4A)。同时进行瘢痕疙瘩成纤维细胞相关差异基因筛选,一共鉴定出420个差异表达基因,其中上调基因212个,下调基因208个。火山图表明瘢痕疙瘩成纤维细胞组相对于正常成纤维细胞组的基因表达有显著差异(图4B),热图展示了差异最显著的上调基因和下调基因各20个(图4C),进一步对关键模块基因和差异表达基因取交集得到80个关键差异基因进行后续分析(图4D)。 2.5 关键差异基因的GO和KEGG富集分析 对黄色模块内的差异表达基因进行GO和KEGG富集分析,结果表明关键差异表达基因参与的主要生物学过程有区域化、骨骼系统形态发生、胚胎骨骼系统发育等;参与的主要细胞组分有内质网内腔、电压门控钙通道复合体、分泌颗粒膜和神经肌肉接头等;参与的分子功能主要有蛋白酪氨酸/苏氨酸磷酸酶活性、有丝分裂原活化蛋白激酶酪氨酸磷酸酶活性、DNA结合转录激活剂活性、骨形态发生蛋白结合等(图5A);参与的主要通路有癌症的转录失调、嘌呤代谢和丝裂原活化蛋白激酶信号通路等(图5B)。图5C,D分别展示了GO和KEGG富集分析前5个通路的基因,主要包括HOXA11,HOXA10,HOXC4,HOXC9等。 2.6 基因互作网络筛选核心基因 使用String数据库分析关键差异基因之间的相互作用,79个基因形成48条边,平均节点度为1.22,蛋白互作网络富集P值为1.11×10-16,不同的颜色表示不同的显著性,其中HOXC8,HOXC6,HOXA11,HOXC4和HOXA5显著性最高(图6A)。使用MCODE插件筛选网络中的核心基因,HOXC8,HOXC9,HOXA11,HOXC4和HOXC10处于核心位置(图6B),前后结果一致。 2.7 机器学习算法筛选瘢痕疙瘩相关标志物 支持向量机算法确定了3个瘢痕疙瘩相关标志物,即HOXC4,HOXA11和HOXC9(图7A)。随机森林算法确定了2个瘢痕疙瘩相关标志物,分别为HOXC4和HOXA11(图7B)。Lasso算法确定了1个瘢痕疙瘩相关标志物,为HOXC4 (图7C)。3种算法取交集最终得到HOXC4作为瘢痕疙瘩相关生物标志物(图7D)。 2.8 瘢痕疙瘩相关标志物HOXC4的诊断性能和鉴定 使用受试者工作特征曲线评估HOXC4在瘢痕疙瘩中的诊断价值,结果表明HOXC4(曲线下面积为0.989,95%CI为0.958-1.000)表现出令人满意的诊断性能(图8A)。同时验证了瘢痕疙瘩成纤维细胞和正常成纤维细胞中HOXC4 mRNA表达水平,结果表明与正常成纤维细胞组相比,HOXC4在瘢痕疙瘩成纤维细胞中高表达,差异有显著性意义(图8B)。 2.9 HOXC4与免疫细胞的相关性分析 为了解瘢痕疙瘩免疫浸润方面的特征,分析了22种免疫细胞的分布情况,在瘢痕数据集中,出现最多的免疫细胞分别为静息记忆CD4+T细胞、"
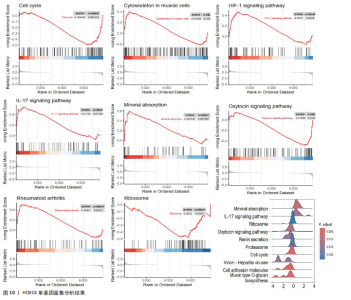
活化树突状细胞、幼稚B细胞和活化肥大细胞等(图9A)。与正常成纤维细胞组相比,瘢痕疙瘩成纤维细胞组调节性T细胞(Tregs)和静息自然杀伤细胞高表达,差异有显著性意义;而活化自然杀伤细胞、单核细胞、M0巨噬细胞和静息肥大细胞在瘢痕疙瘩成纤维细胞组低表达,差异有显著性意义(图9B)。进一步分析了关键基因HOXC4与免疫细胞的相关性,结果表明HOXC4与静息自然杀伤细胞呈显著正相关,与活化树突状细胞和活化自然杀伤细胞呈显著负相关(图9C)。 2.10 HOXC4单基因集富集分析 对瘢痕疙瘩的关键诊断基因HOXC4进行单基因集富集分析,结果表明富集的主要通路有细胞周期、肌肉细胞中的细胞骨架、缺氧诱导因子1信号通路、白细胞介素17信号通路、矿物质吸收、催产素信号通路、类风湿性关节炎和核糖体等,最后通过山脊图展示上调的前10个通路,见图10。"
| [1] HAWASH AA, INGRASCI G, NOURI K, et al. Pruritus in Keloid Scars: Mechanisms and Treatments. Acta Derm Venereol. 2021;101(10):adv00582. [2] KELLY AP. Keloids. Dermatol Clin. 1988;6(3): 413-424. [3] LEE SS, YOSIPOVITCH G, CHAN YH, et al. Pruritus, pain, and small nerve fiber function in keloids: a controlled study. J Am Acad Dermatol. 2004; 51(6):1002-1006. [4] DOHI T, PADMANABHAN J, AKAISHI S, et al. The Interplay of Mechanical Stress, Strain, and Stiffness at the Keloid Periphery Correlates with Increased Caveolin-1/ROCK Signaling and Scar Progression. Plast Reconstr Surg. 2019;144(1):58e-67e. [5] KASSI K, KOUAME K, KOUASSI A, et al. Quality of life in black African patients with keloid scars. Dermatol Reports. 2020;12(2):8312. [6] FERNANDES MG, DA SILVA LP, CERQUEIRA MT, et al. Mechanomodulatory biomaterials prospects in scar prevention and treatment. Acta Biomater. 2022;150:22-33. [7] WEI J, WANG Z, ZHONG C, et al. LncRNA MIR503HG promotes hypertrophic scar progression via miR-143-3p-mediated Smad3 expression. Wound Repair Regen. 2021;29(5): 792-800. [8] BIAZUS SOARES G, MAHMOUD O, YOSIPOVITCH G. Pruritus in keloid scars: mechanisms and treatments. Ital J Dermatol Venerol. 2023;158(5):401-407. [9] MUSKAT A, KOST Y, BALAZIC E, et al. Laser-Assisted Drug Delivery in the Treatment of Scars, Rhytids, and Melasma: A Comprehensive Review of the Literature. Aesthet Surg J. 2023;43(3):NP181-NP198. [10] BAGABIR R, BYERS RJ, CHAUDHRY IH, et al. Site-specific immunophenotyping of keloid disease demonstrates immune upregulation and the presence of lymphoid aggregates. Br J Dermatol. 2012;167(5):1053-1066. [11] SHARMA JR, LEBEKO M, KIDZERU EB, et al. In Vitro and Ex Vivo Models for Functional Testing of Therapeutic Anti-scarring Drug Targets in Keloids. Adv Wound Care (New Rochelle). 2019; 8(12):655-670. [12] 李政宇.基于综合生物信息学筛选瘢痕疙瘩关键生物标志物及SDC4泛癌分析[D].太原:太原理工大学,2023. [13] LV W, REN Y, HOU K, et al. Epigenetic modification mechanisms involved in keloid: current status and prospect. Clin Epigenetics. 2020;12(1):183. [14] XIE R, LI C, ZHAO T, et al. Integration of Flow Cytometry and Single-Cell RNA Sequencing Analysis to Explore the Fibroblast Subpopulations in Keloid that Correlate with Recurrence. Adv Wound Care (New Rochelle). 2025. doi: 10.1089/wound.2024.0262. [15] ZHONG C, SHI K, LI P, et al. Single-cell sequencing analysis and bulk-seq identify IGFBP6 and TNFAIP6 as novel differential diagnosis markers for postburn pathological scarring. Burns. 2024; 50(9):107255. [16] SZKLARCZYK D, KIRSCH R, KOUTROULI M, et al. The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 2023;51(D1):D638-D646. [17] WEI W, LI Y, HUANG T. Using Machine Learning Methods to Study Colorectal Cancer Tumor Micro-Environment and Its Biomarkers. Int J Mol Sci. 2023;24(13):11133. [18] UDDIN S, KHAN A, HOSSAIN ME, et al. Comparing different supervised machine learning algorithms for disease prediction. BMC Med Inform Decis Mak. 2019;19(1):281. [19] MOTAMEDI F, PÉREZ-SÁNCHEZ H, MEHRIDEHNAVI A, et al. Accelerating Big Data Analysis through LASSO-Random Forest Algorithm in QSAR Studies. Bioinformatics. 2022;38(2):469-475. [20] BIJL JJ, RIEGER E, VAN OOSTVEEN JW, et al. HOXC4, HOXC5, and HOXC6 expression in primary cutaneous lymphoid lesions. High expression of HOXC5 in anaplastic large-cell lymphomas. Am J Pathol. 1997;151(4):1067-1074. [21] WU S, CHEN H, ZUO L, et al. Suppression of long noncoding RNA MALAT1 inhibits the development of uveal melanoma via microRNA-608-mediated inhibition of HOXC4. Am J Physiol Cell Physiol. 2020;318(5):C903-C912. [22] ZHANG H, HAN B, TIAN S, et al. HOXC4 promotes proliferation of pancreatic cancer cells by increasing LDHA-mediated glycolysis. Aging (Albany NY). 2024;16(13):11103-11116. [23] HAHN JM, MCFARLAND KL, COMBS KA, et al. Analysis of HOX gene expression and the effects of HOXA9 overexpression in fibroblasts derived from keloid lesions and normal skin. Wound Repair Regen. 2021;29(5):777-791. [24] HAHN JM, GLASER K, MCFARLAND KL, et al. Keloid-derived keratinocytes exhibit an abnormal gene expression profile consistent with a distinct causal role in keloid pathology. Wound Repair Regen. 2013;21(4):530-544. [25] ZHANG N, GUO F, SONG Y. HOXC8/TGF-β1 positive feedback loop promotes liver fibrosis and hepatic stellate cell activation via activating Smad2/Smad3 signaling. Biochem Biophys Res Commun. 2023;662:39-46. [26] XIAO J, LI Y, LIU Y, et al. The involvement of homeobox-C 4 in predicting prognosis and unraveling immune landscape across multiple cancers via integrated analysis. Front Genet. 2022;13:1021473. [27] KIRK T, AHMED A, ROGNONI E. Fibroblast Memory in Development, Homeostasis and Disease. Cells. 2021;10(11):2840. [28] JIAO H, FAN J, CAI J, et al. Analysis of Characteristics Similar to Autoimmune Disease in Keloid Patients. Aesthetic Plast Surg. 2015;39(5):818-825. [29] WU J, DEL DUCA E, ESPINO M, et al. RNA Sequencing Keloid Transcriptome Associates Keloids With Th2, Th1, Th17/Th22, and JAK3-Skewing. Front Immunol. 2020;11:597741. [30] LEE CC, TSAI CH, CHEN CH, et al. An updated review of the immunological mechanisms of keloid scars. Front Immunol. 2023;14:1117630. [31] MARTIN CW, MUIR IF. The role of lymphocytes in wound healing. Br J Plast Surg. 1990;43(6):655-662. [32] LI X, AN T, YANG Y, et al. TLR9 activation in large wound induces tissue repair and hair follicle regeneration via γδT cells. Cell Death Dis. 2024;15(8):598. [33] XIA Y, WANG Y, SHAN M, et al. Advances in the pathogenesis and clinical application prospects of tumor biomolecules in keloid. Burns Trauma. 2022;10:tkac025. [34] DENG CC, XU XY, ZHANG Y, et al. Single-cell RNA-seq reveals immune cell heterogeneity and increased Th17 cells in human fibrotic skin diseases. Front Immunol. 2025;15: 1522076. [35] ZHANG X, WU X, LI D. The Communication from Immune Cells to the Fibroblasts in Keloids: Implications for Immunotherapy. Int J Mol Sci. 2023;24(20):15475. [36] SHAN M, WANG Y. Viewing keloids within the immune microenvironment. Am J Transl Res. 2022;14(2):718-727. [37] LI Y, LI M, QU C, et al. The Polygenic Map of Keloid Fibroblasts Reveals Fibrosis-Associated Gene Alterations in Inflammation and Immune Responses. Front Immunol. 2022;12:810290. |
| [1] | Liu Hongjie, Mu Qiuju, Shen Yuxue, Liang Fei, Zhu Lili. Metal organic framework/carboxymethyl chitosan-oxidized sodium alginate/platelet-rich plasma hydrogel promotes healing of diabetic infected wounds [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(8): 1929-1939. |
| [2] | Lai Jiaming, , Song Yuling, Chen Zixi, Wei Jinghuan, Cai Hao, , Li Guoquan, . Screening of diagnostic markers for endothelial cell Senescence in mice with radiation-induced heart disease and analysis of immune infiltration [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(6): 1450-1463. |
| [3] | Peng Tuanhui, Song Hongming, Yang Ling, Ding Xiaoge, Meng Pengjun. Effects of long-term endurance exercise on kl/FGF23 axis and calcium-phosphorus metabolism in naturally aging mice [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(5): 1089-1095. |
| [4] | Bao Zhuoma, Hou Ziming, Jiang Lu, Li Weiyi, Zhang Zongxing, Liu Daozhong, Yuan Lin. Effect and mechanism by which Pterocarya hupehensis skan total flavonoids regulates the proliferation, migration and apoptosis of fibroblast-like synoviocytes [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(4): 816-823. |
| [5] | Guo Jiachen, Gao Jun, Dai Wenhao, Liao Huayuan, Jiang You, Zhang Xi . Effect of compressive stress microenvironment on cytokines during fracture healing [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(4): 908-916. |
| [6] | Li Feihong, Wang Linrong, Cheng Leping. Role of fibrosis in tissue injury repair [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(22): 5814-5831. |
| [7] | Chen Ling, Mao Qiuhua, Xu Pu, Zhang Wenbo. Effect of water-soluble matrix of nano-pearl powder on proliferation, migration and apoptosis of mouse fibroblasts#br# [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(2): 338-344. |
| [8] | Shi Yuxin, Kaiwusail · Tursun, Liu Jia. Effects of basic fibroblast growth factor-loaded composite bioscaffold on angiogenesis of dental pulp stem cells [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(13): 3343-3349. |
| [9] | Wu Xianyuan, Zhang Nini, Huang Guilin. Gene transfection technology and tissue fibrosis repair [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(13): 3424-3434. |
| [10] | Wu Zhijing, Li Jiali, Zhang Jiaxin, Wang Tangrong, Zheng Yuzhou, Sun Zixuan. Alpha-ketoglutarate engineered small extracellular vesicles delay skin aging [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(1): 120-129. |
| [11] | Han Haihui, Ran Lei, Meng Xiaohui, Xin Pengfei, Xiang Zheng, Bian Yanqin, Shi Qi, Xiao Lianbo. Targeting fibroblast growth factor receptor 1 signaling to improve bone destruction in rheumatoid arthritis [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(9): 1905-1912. |
| [12] | Li Jun, Gong Jingjing, Sun Guobin, Guo Rui, Ding Yang, Qiang Lijuan, Zhang Xiaoli, Fang Zhanhai . miR-27a-3p promotes the proliferation of human hypertrophic scar fibroblasts by regulating mitogen-activated protein kinase signaling pathway [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(8): 1609-1617. |
| [13] | Chen Yuning, Jiang Ying, Liao Xiangyu, Chen Qiongjun, Xiong Liang, Liu Yue, Liu Tong. Buqi Huoxue Compounds intervene with the expression of related factors and autophagy related proteins in a rat model of cerebral ischemia/reperfusion [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(6): 1152-1158. |
| [14] | Zhang Debao, Wang Peng, Li Kun, Zhang Shaojie, Li Zhijun, Li Shuwen, Wu Yimin. Epidural fibrous scar formation in rabbits following autologous ligamentum flavum intervention [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(6): 1168-1175. |
| [15] | Han Haihui, Meng Xiaohu, Xu Bo, Ran Le, Shi Qi, Xiao Lianbo. Effect of fibroblast growth factor receptor 1 inhibitor on bone destruction in rats with collagen-induced arthritis [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(5): 968-977. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||