Chinese Journal of Tissue Engineering Research ›› 2021, Vol. 25 ›› Issue (26): 4236-4242.doi: 10.12307/2021.127
Previous Articles Next Articles
N6-methyladenosine RNA methylation is involved in orthopedic related diseases
Chen Weijian1, Zhang Gangyu2, Lin Tianye2, Liang Du1, Wang Haibin3
- 1Guangzhou Bone Setting Hospital affiliated to Guangzhou University of Chinese Medicine, Guangzhou 510405, Guangdong Province, China; 2The First Clinical Medical College of Guangzhou University of Chinese Medicine, Guangzhou 510405, Guangdong Province, China; 3Department of Orthopedics, The First Affiliated Hospital of Guangzhou University of Chinese Medicine, Guangzhou 510405, Guangdong Province, China
-
Received:2020-08-07Revised:2020-08-11Accepted:2020-09-04Online:2021-09-18Published:2021-05-13 -
Contact:Wang Haibin, Chief physician, Doctoral supervisor, Department of Orthopedics, The First Affiliated Hospital of Guangzhou University of Chinese Medicine, Guangzhou 510405, Guangdong Province, China E-mail:whbtdzyyx@163.com -
About author:Chen Weijian, Master candidate, Guangzhou Bone Setting Hospital affiliated to Guangzhou University of Chinese Medicine, Guangzhou 510405, Guangdong Province, China -
Supported by:the National Natural Science Foundation of China in 2017 (General Program), No. 81774339 (to WHB); the National Natural Science Foundation of China in 2020 (General Program), No. 82074462 (to WHB)
CLC Number:
Cite this article
Chen Weijian, Zhang Gangyu, Lin Tianye, Liang Du, Wang Haibin. N6-methyladenosine RNA methylation is involved in orthopedic related diseases[J]. Chinese Journal of Tissue Engineering Research, 2021, 25(26): 4236-4242.
share this article
Add to citation manager EndNote|Reference Manager|ProCite|BibTeX|RefWorks
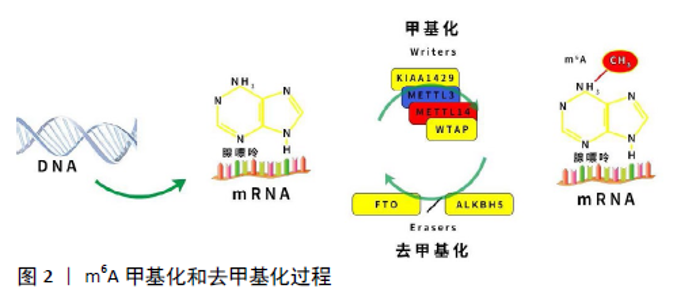
2.1 RNA m6A甲基化修饰概述 RNA表观遗传修饰是在RNA水平调节基因表达的重要途径,动态可逆的表观遗传修饰可为疾病的早期干预提供科学依据。根据研究发现,RNA甲基化修饰占RNA修饰的60%以上,是RNA修饰最丰富的形式。常见的RNA甲基化修饰包括m6A、N1-甲基腺嘌呤、5-甲基胞嘧啶和假尿嘧啶核苷等[6]。m6A是真核生物mRNA最丰富的甲基化修饰,参与RNA代谢的调控,包括RNA的可变剪接、转运、翻译和降解等,从而调控各种细胞生理病理进程,如细胞增殖、分化、凋亡、侵袭以及DNA损伤修复等[7]。 早在数十年前,研究者们发现m6A修饰在多种真核生物中存在[1],其中在细菌、酵母、果蝇和哺乳动物等真核生物RNA中发现了m6A甲基化修饰水平,而且早期研究发现,m6A修饰占哺乳动物RNA腺苷酸的0.1%-0.4%和核糖核苷酸总甲基化的50%[8],揭示了m6A修饰在转录水平的普遍性。另外,早期鉴定的mRNA m6A修饰位点附近的序列具有高度保守性,主要出现在RRACH(R代表嘌呤A、G;A代表甲基化位置,H代表A、C、U)的腺嘌呤序列上[9]。尽管m6A早已在各种真核生物中被广泛发现,但由于m6A修饰并没有影响腺嘌呤和胸腺嘧啶的结合,无法运用基于标准的配对杂交或者基于测序的方法对m6A进行大规模检测,导致对于m6A甲基化修饰的研究陷入了停滞[10]。 直至2012年,DAN和KATE等两组研究团队开发出MeRIP-seq技术(亦称为m6A-seq技术)[11],通过利用高度特异性的m6A抗体免疫沉淀甲基化的mRNA,结合高通量测序以确定甲基化的转录本,用于大规模检测m6A修饰,同时揭示了m6A常发生在编码区和3’非翻译区中,特别是在终止密码子附近富集。随着RNA修饰检测技术的飞速发展,m6A的结构、功能和作用机制已被深入研究。当前研究发现,m6A修饰广泛存在于哺乳动物多个组织中,其中以肝脏、肾脏和大脑中m6A修饰水平最高[11],并且在骨组织、关节软骨、骨肿瘤组织以及骨髓间充质干细胞中均有不同水平的m6A修饰。另外还发现m6A参与神经系统发育和干细胞分化等多种生理过程,包括促进神经干细胞/祖细胞的自我更新和神经元生成、促进神经干细胞增殖和延长放射状胶质细胞的细胞周期等[12],并且还参与调控造血干细胞的定向分化[13],影响人体多种肿瘤增殖、侵袭和转移,同时还参与非编码RNA的相互调控[13]。另外,在骨组织细胞的研究中,发现m6A修饰参与调控骨髓间充质干细胞的成脂、成骨分化,并且有研究发现在骨质疏松症患者中骨组织总体m6A水平下降,其主要的m6A甲基化转移酶METTL3表达降低,揭示了m6A修饰相关的酶和调控蛋白参与了调节疾病的进程。 2.2 m6A甲基化修饰相关蛋白 目前,涉及m6A甲基化修饰的相关蛋白有3类:①甲基转移酶复合体(Writers),负责催化m6A甲基化;②负责催化m6A去甲基化的蛋白(Erasers);③识别m6A甲基化的结合蛋白(Readers)[15]。简而言之,“Writers”催化m6A修饰启动,促进RNA特定位点的甲基化,并通过不同的“Readers”来识别m6A修饰的RNA而产生不同的功能,包括RNA加工、核输出、翻译、衰变等。最后,依靠“Erasers”对m6A修饰进行去甲基化,从而使m6A甲基化过程变得动态、可逆,调节各种基因的表达,见图2。"
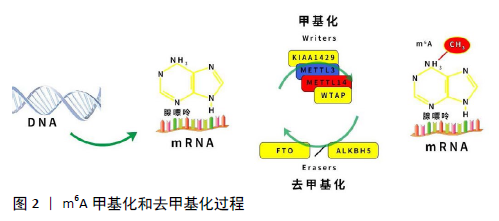

2.2.1 m6A甲基转移酶——Writers Writers是指将m6A甲基化“录入”特定的RNA位点中,从而实现 RNA甲基化修饰。m6A甲基化修饰主要由甲基转移酶复合体介导,该复合体主要包括:METTL3、METTL14、WTAP、KIAA29、RBM15/RBM15B、METTL16和ZC3H3蛋白[16]。METTL3是催化m6A甲基化的核心,并与METTL14形成异二聚体[17],促进m6A的沉积;METTL16参与催化m6A修饰;WTAP募集METTL3/METTL14异二聚体来提高m6A修饰水平[18]。除此之外,其他Writers主要通过胞间信号的传递,促进m6A复合体的功能,募集m6A复合体到达特定的RNA位点[19]。 2.2.2 m6A去甲基化酶——Erasers Erasers可“消除”RNA甲基化修饰信号,实现RNA去甲基化修饰。Erasers主要包括肥胖相关蛋白(obesity-associated protein, FTO)和ALKBH5,属于ALKB双加氧酶家族蛋白,并依赖于Fe2+和α-酮戊二酸来执行去甲基化功能[20]。FTO主要负责氧化去除细胞核和胞质中m6A甲基化,而ALKBH5仅负责氧化去除细胞核中甲基化的m6A[19]。 2.2.3 m6A结合蛋白——Readers Readers负责“识别”RNA甲基化修饰的信息,通过直接或间接结合RNA来参与下游RNA加工、降解等过程[21]。直接结合是指直接结合到RNA的m6A位点,如具有RNA结合结构域的YTH 结构域家族(YTHDF1、YTHDF2、YTHDF3、YTHDC1和YTHDC2),可直接结合细胞核和胞质中甲基化修饰的靶基因而发挥作用[22]。此外,间接RNA结合依赖于m6A甲基化诱导的RNA解螺旋,从而暴露隐藏的蛋白质结合位点并促进蛋白质与RNA结合[23]。 2.3 m6A在骨质疏松症中的研究进展 骨质疏松症的发病与骨代谢平衡的失调有关,骨吸收增强、骨形成不足、骨稳态发生改变均会引起骨质疏松[24]。骨髓间充质干细胞是成骨细胞和脂肪细胞的共同祖细胞,其成骨和成脂分化的平衡受多种因素严格调控,以维持骨代谢的平衡[25]。其中老年人和骨质疏松症患者的特点是骨量减少和脂肪组织过度积累。目前已有研究发现,在衰老或其他病理刺激(如激素失调)下,骨髓间充质干细胞向脂肪细胞的分化导致骨量和脂肪之间的失衡,导致骨质流失[26]。骨髓间充质干细胞的分化方向以及自我更新受细胞外基质和细胞内信号因子的影响,如转录因子过氧化物酶体增殖物激活受体γ2和RUNX2、氧化应激、机械应力及表观遗传修饰等[27-28],其中已有研究证明,表观遗传修饰中的DNA甲基化和翻译后组蛋白修饰参与了成骨相关基因的表达[29-30]。这些表观遗传程序对于骨质疏松症中的骨代谢紊乱以及骨重建等病理过程是必不可少的。近年来有大量研究表明,m6A修饰是骨质疏松症的潜在发病机制,m6A可通过其酶系统调节相关转录因子的表达,影响骨髓间充质干细胞、成骨细胞、脂肪细胞和破骨细胞的分化、增殖和凋亡[31-34]。 2.3.1 m6A修饰调节成骨细胞分化 TIAN等[31]首次研究发现,METTL3在诱导成骨分化的骨髓间充质干细胞中高度表达,而沉默METTL3后明显抑制了骨髓间充质干细胞的成骨分化潜能,且成骨分化标志物RUNX2、碱性磷酸酶、骨钙素以及Osterix表达均降低。此外,研究人员还发现,沉默METTL3后明显降低了骨髓间充质干细胞中血管内皮生长因子及其骨形成相关剪接变异体血管内皮生长因子A-164、血管内皮生长因子A-188的表达,已有研究证明血管内皮生长因子A-164在改善间充质干细胞增殖和软骨分化中起调节作用,而血管内皮生长因子A-188在促进间充质干细胞的成骨分化中发挥重要的作用[35]。另外,沉默METTL3基因后明显降低了骨髓间充质干细胞的Akt磷酸化水平,而PI3K-AKT信号传导在骨组织代谢和骨髓间充质干细胞的成骨分化中也起着至关重要的调节作用[36-38]。总而言之,METTL3通过调控血管内皮生长因子相关剪切变异体以及通过PI3K/AKT信号通路促进了骨髓间充质干细胞的成骨分化。 同样,WU等[39]通过CRISPR/Cas9技术敲除小鼠的METTL3基因,发现敲除小鼠的骨量减少,骨髓脂肪组织显著增加。此外,通过分离敲除小鼠的骨髓间充质干细胞进行体外成骨成脂潜能培养,发现成骨标志物RUNX2、Sp7、碱性磷酸酶和骨钙素的表达下调,而成脂因子过氧化物酶体增殖物激活受体γ、CEBPα、Adipoq、Plin1和CD36的表达增加,表明敲除METTL3基因后,小鼠的成骨分化潜能降低,而成脂分化增强。另外,为了保证实验的准确性,研究人员在卵巢切除小鼠产生的骨质疏松模型中过表达METTL3,发现其骨小梁密度和骨量减少量较对照组少,且发现过表达METTL3的小鼠成骨细胞数量增多;最后,通过MeRIP-Seq鉴定,METTL3/m6A通过甲状旁腺激素/甲状旁腺激素受体1信号轴影响骨髓间充质干细胞的成骨与成脂分化。 YAN等[32]在骨质疏松症患者的骨组织以及骨质疏松小鼠模型中除了发现m6A甲基化修饰水平外,还检测到METTL3和METTL14 mRNA的表达,METTL3/m6A甲基化可通过调控RUNX2的表达来诱导骨髓间充质干细胞的成骨分化。通过RNA点印迹和ELISA法检测,研究人员发现,骨质疏松症患者的骨组织以及骨质疏松小鼠中总体RNA的m6A、METTL3和METTL14 mRNA较对照组含量显著降低,而且其成骨细胞分化标志物骨钙素、骨形态发生蛋白2和碱性磷酸酶的表达水平低于对照组。此外,在转染METTL3过表达的骨髓间充质干细胞中发现总RNA的m6A水平以及成骨分化相关标记基因表达上升。随着研究的深入,研究人员还发现METTL3/m6A通过改变pre-miR-320的甲基化来降低miR-320的表达,进一步促进成骨转录因子RUNX2的表达从而促进成骨细胞的分化。另外,他们还通过沉默METTL3来检测m6A水平是否影响骨髓间充质干细胞的成骨分化,结果表明转染si-METTL3的骨髓间充质干细胞中成骨相关基因RUNX2、骨钙素和碱性磷酸酶的表达显著降低,表明骨髓间充质干细胞中成骨分化能力减弱。 另外,还有研究发现,在骨质疏松症中,FTO是决定骨髓间充质干细胞分化方向的调节因子,通过影响骨髓中GDF11-C/EBP mRNA信号轴激活,促进Smad2/3磷酸化刺激破骨细胞生成和抑制成骨细胞分化而导致骨质疏松的发展[40-42]。 2.3.2 m6A修饰调节破骨细胞分化 LI等[33]研究发现,在破骨细胞系中的RAW264.7、骨髓单核巨噬细胞中均检测到m6A以及METTL3的表达,且在诱导破骨细胞分化后,m6A水平和METTL3表达均有上升。另外,研究人员用RANKL和巨噬细胞集落刺激因子处理si-METTL3的骨髓单核巨噬细胞,发现其分化的破骨细胞的骨吸收活性降低,破骨细胞分化相关基因c-Fos、Nfatc1和Dcstamp以及骨吸收相关基因Ctsk、Acp5的mRNA和蛋白表达水平均下降。研究人员还发现METTL3沉默的细胞中JNK、P38、ERK、IKKα/β、P65、IκBα和AKT的磷酸化水平均降低,这表明了METTL3可能通过激活MAPK、核因子κB和AKT信号通路来调节RANKL诱导的破骨细胞分化。 2.3.3 m6A修饰调节脂肪细胞分化 YAO等[34]研究发现,METTL3表达与猪骨髓间充质干细胞成脂分化关系呈负相关,METTL3以m6A/YTHDF2依赖的方式靶向调控JAK1/STAT5/C/EBPβ途径,从而抑制骨髓间充质干细胞的成脂分化。此外,沉默METTL3基因可通过m6A显著促进猪骨髓间充质干细胞的成脂过程和JAK1蛋白的表达。 SHEN等[43]研究发现,FTO/m6A通过影响过氧化物酶体增殖物激活受体γ的去甲基化从而激活GDF11-FTO-PPARγ信号轴,促进骨髓间充质干细胞向成脂分化,而抑制成骨分化,从而导致骨质疏松发展过程中的骨形成能力减弱,导致骨量和脂肪间的失衡。 2.4 m6A修饰在骨关节炎中的研究进展 骨关节炎是一种慢性进行性关节疾病,其发病率随着年龄的增长而增高,其特征是关节软骨的破损、软骨下硬化、骨赘形成[44]。进行性软骨退变是骨关节炎发病的重要病理改变,包括软骨细胞的凋亡和软骨细胞合成的细胞外基质成分的改变[45]。LIU等[46]研究发现,通过m6A甲基化定量检测及qRT-PCR检测,在用白细胞介素1β处理的鼠软骨细胞祖细胞系ATDC5细胞中METTL3和m6A的水平上调。此外,研究人员通过转染shMETTL3到ATDC5细胞中,显著降低白细胞介素1β诱导的软骨细胞凋亡,同时降低了炎症因子白细胞介素8、白细胞介素6、白细胞介素12和肿瘤坏死因子α的mRNA和蛋白表达水平,基质金属蛋白酶13和X型胶原的mRNA水平显著降低,而Ⅱ型胶原和聚集蛋白聚糖的mRNA水平显著升高,结果均表明METTL3的沉默抑制了白细胞介素1β诱导的炎症反应和细胞外基质合成。最后,为了确保实验结果的准确性,研究人员通过注射甲基化抑制剂环亮氨酸到白细胞介素1β处理的ATDC5细胞中,发现甲基化抑制剂环亮氨酸处理可降低骨关节炎软骨组织中白细胞介素8、白细胞介素6 mRNA的上调,且可提高X型胶原和Ⅱ型胶原的表达。综上所述,METTL3/m6A可通过介导炎症反应和细胞外基质降解促进骨关节炎进展。 同样,ZHANG等[47]研究发现,METTL3 mRNA和蛋白水平在成骨分化过程中表达增加而在炎症刺激后降低。另外,研究人员还发现,在转染METTL3沉默后的成骨细胞中,Smad1,5,9的磷酸化水平降低,Smad7和Smurf1的表达增加,表明了METTL3可能在炎症环境中通过Smad信号通路参与调节成骨分化。已有研究证明,成骨细胞通过激活MAPK和核因子κB信号通路来调节下游促炎细胞因子(如白细胞介素6、白细胞介素12和肿瘤坏死因子α)表达[48-49],通过Western blot检测IKKα/β、p65、IκBα、ERK、p38和JNK的磷酸化水平,发现在转染METTL3沉默的成骨细胞中p-ERK、p-p38、p-JNK和p-p65蛋白表达水平升高,揭示了沉默METTL3可激活MAPK信号通路,从而促进脂多糖处理的成骨细胞中促炎细胞因子的表达。综上,METTL3/m6A可作为抑制软骨细胞凋亡、促进细胞外基质合成、抑制促炎因子表达的潜在治疗靶点。 2.5 m6A修饰在其他骨病的研究进展 跌倒、交通事故和骨质疏松症等均会增加骨折的风险,尽管外固定和内固定在治疗骨折方面取得了良好的效果,但也难免存在一些失败的情况,另外骨折愈合的具体机制至今尚未研究清楚,先前已有大量研究证实了骨折愈合是由多种因素介导的复杂生物过程,其中包括了促进和抗骨折愈合的细胞因子、非编码RNA等[50-52]。在这些调控因素中,生物因素对骨折愈合影响较为显著,已有研究表明通过调节或补充细胞外基质蛋白、改变局部免疫微环境,靶向输送生长因子及成骨诱导因子等能够有效治疗骨延迟愈合或骨不连等骨折并发症[50-53]。MI 等[54]研究发现,骨折后的前7 d内METTL3、WTAP和KIAA1429的水平显著降低,研究人员还将质粒METTL3局部注入小鼠股骨骨折部位,发现成骨相关基因骨形态发生蛋白2、RUNX2在质粒METTL3组中的表达明显降低,同时在细胞水平验证中得到了相似的结果,表明METTL3在骨折愈合过程中起负调节作用。最后,研究人员还证明了METTL3通过依赖于m6A的pri-miRNA加工方式靶向调控成骨细胞相关的miR-7212-5p来抑制骨折愈合过程中的成骨能力。 骨肉瘤是儿童和青壮年最常见的原发性恶性肿瘤,其特点是恶性程度高,并能迅速损害周围组织进行转移,早期骨肉瘤细胞转移是导致骨肉瘤患者预后不良的主要因素之一。虽然目前骨肉瘤的治疗已有一定发展,包括手术切除联合全身化疗或放疗,但是骨肉瘤患者的生存率依然很低。此外,接受手术治疗的骨肉瘤患者中约有80%会复发,严重影响了患者的预后[55]。因此,有必要进一步研究骨肉瘤的分子致癌机制,寻找骨肉瘤的新治疗靶点。目前已有大量研究发现m6A在各器官肿瘤中的分子靶点和调控作用,其中包括通过m6A依赖性机制增加RNA衰变、稳定性或增强翻译等,影响多器官肿瘤增殖、侵袭和转移,如FTO在急性髓细胞白血病和脑肿瘤中显示抗肿瘤作用[56];ALKBH5在乳腺癌中显示促进肿瘤的作用[57];METTL3在急性髓细胞白血病、肝癌和肺癌的发生发展中起促进作用;而METTL14在促进和抑制肿瘤发生等方面具有双重作用等[58]。然而m6A在骨肉瘤中的作用及其表观遗传修饰机制的相关研究较少。MIAO等[59]研究发现,在人类骨肉瘤组织和细胞系中,总m6A水平和METTL3的表达水平均升高,在HOS和SAOS-2细胞中沉默METTL3抑制了骨肉瘤细胞的增殖、迁移和侵袭能力。研究人员还发现METTL3沉默降低了m6A水平和淋巴增强因子1的RNA表达,进而抑制了Wnt/β-catenin信号通路的活性,以上结果均表明了METTL3通过调节淋巴增强因子1的m6A水平从而激活Wnt/β-catenin信号来促进骨肉瘤细胞的进展。同样,ZHOU等[60]研究发现,METTL3定位于骨肉瘤细胞的胞浆和胞核内,在SAOS-2和MG63细胞中沉默METTL3可显著抑制m6A水平,抑制细胞增殖、分化、侵袭和迁移,促进细胞凋亡。此外,研究人员还发现METTL3基因敲除后抑制了ATAD2的表达,而ATAD2基因沉默后抑制了SAOS-2和MG63细胞的增殖和侵袭,而其过表达则显著增强了细胞的增殖和侵袭,以上均表明了METTL3通过调节ATAD2 m6A甲基化来促进骨肉瘤的生长和转移,同时LI等[61]研究发现,FTO和METTL14的低表达以及METTL3,HNRNPA2B1和YTHDF3的高表达与骨肉瘤的不良预后有关,并通过生物信息学分析发现,m6A调控蛋白可能通过体液免疫应答和细胞周期途径参与骨肉瘤进程,提示m6A甲基化修饰可能是骨肉瘤潜在的治疗靶点,为靶向治疗人骨肉瘤提供了新的思路和突破口。"

| [1] DUBIN DT, TAYLOR RH. The methylation state of poly A-containing messenger RNA from cultured hamster cells. Nucleic Acids Res. 1975; 2(10):1653-1668. [2] 张翔, 杜娟, 陈雅慧, 等. mRNA m6A甲基化修饰异常与疾病的研究进展[J]. 生命的化学,2019,39(2):255-261. [3] DESROSIERS R, FRIDERICI K, ROTTMAN F. Identification of methylated nucleosides in messenger RNA from Novikoff hepatoma cells. Proc Natl Acad Sci U S A. 1974;71(10):3971-3975. [4] HOROWITZ S, HOROWITZ A, NILSEN TW, et al. Mapping of N6-methyladenosine residues in bovine prolactin mRNA. Proc Natl Acad Sci U S A. 1984;81(18):5667-5671. [5] KRUG RM, MORGAN MA, SHATKIN AJ. Influenza viral mRNA contains internal N6-methyladenosine and 5’-terminal 7-methylguanosine in cap structures. J Virol. 1976;20(1):45-53. [6] TRAUBE FR, CARELL T. The chemistries and consequences of DNA and RNA methylation and demethylation. RNA Biol. 2017;14(9):1099-1107. [7] HUANG H, WENG H, CHEN J. The Biogenesis and Precise Control of RNA m(6)A Methylation. Trends Genet. 2020;36(1):44-52. [8] WEI CM, GERSHOWITZ A, MOSS B. Methylated nucleotides block 5’ terminus of HeLa cell messenger RNA. Cell. 1975;4(4):379-386. [9] DOMINISSINI D, MOSHITCH-MOSHKOVITZ S, SCHWARTZ S, et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature. 2012;485(7397):201-206. [10] CAO G, LI H B, YIN Z, et al. Recent advances in dynamic m6A RNA modification. Open Biol. 2016;6(4):160003. [11] MEYER KD, SALETORE Y, ZUMBO P, et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3’ UTRs and near stop codons. Cell. 2012;149(7):1635-1646. [12] LI M, ZHAO X, WANG W, et al. Ythdf2-mediated m(6)A mRNA clearance modulates neural development in mice. Genome Biol. 2018;19(1):69. [13] ZHANG C, CHEN Y, SUN B, et al. m(6)A modulates haematopoietic stem and progenitor cell specification. Nature. 2017;549(7671):273-276. [14] WANG X, LU Z, GOMEZ A, et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature. 2014;505(7481): 117-120. [15] ALDERMAN MR, XIAO AZ. N(6)-Methyladenine in eukaryotes. Cell Mol Life Sci. 2019;76(15):2957-2966. [16] LENCE T, PAOLANTONI C, WORPENBERG L, et al. Mechanistic insights into m(6)A RNA enzymes. Biochim Biophys Acta Gene Regul Mech. 2019;1862(3):222-229. [17] LIU J, YUE Y, HAN D, et al. A METTL3-METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation. Nat Chem Biol. 2014;10(2):93-95. [18] PING XL, SUN BF, WANG L, et al. Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Res. 2014;24(2):177-189. [19] LIAO S, SUN H, XU C. YTH Domain: A Family of N(6)-methyladenosine (m(6)A) Readers. Genomics Proteomics Bioinformatics. 2018;16(2): 99-107. [20] ZHAO W, QI X, LIU L, et al. Epigenetic Regulation of m(6)A Modifications in Human Cancer. Mol Ther Nucleic Acids. 2020;19:405-412. [21] YUE Y, LIU J, HE C. RNA N6-methyladenosine methylation in post- transcriptional gene expression regulation. Genes Dev. 2015;29(13): 1343-1355. [22] LIU ZX, LI LM, SUN HL, et al. Link Between m6A Modification and Cancers. Front Bioeng Biotechnol. 2018;6:89. [23] LIU N, DAI Q, ZHENG G, et al. N(6)-methyladenosine-dependent RNA structural switches regulate RNA-protein interactions. Nature. 2015;518(7540):560-564. [24] ANDERSEN TL, SONDERGAARD TE, SKORZYNSKA KE, et al. A physical mechanism for coupling bone resorption and formation in adult human bone. Am J Pathol. 2009;174(1):239-247. [25] KIM M, KIM C, CHOI YS, et al. Age-related alterations in mesenchymal stem cells related to shift in differentiation from osteogenic to adipogenic potential: implication to age-associated bone diseases and defects. Mech Ageing Dev. 2012;133(5):215-225. [26] CAMPI G, CRISTOFARO F, PANI G, et al. Heterogeneous and self-organizing mineralization of bone matrix promoted by hydroxyapatite nanoparticles. Nanoscale. 2017;9(44):17274-17283. [27] BATEMAN ME, STRONG AL, MCLACHLAN JA, et al. The Effects of Endocrine Disruptors on Adipogenesis and Osteogenesis in Mesenchymal Stem Cells: A Review. Front Endocrinol (Lausanne). 2016;7:171. [28] NAKASHIMA K, DE CROMBRUGGHE B. Transcriptional mechanisms in osteoblast differentiation and bone formation. Trends Genet. 2003; 19(8):458-466. [29] DELGADO-CALLE J, SAÑUDO C, SÁNCHEZ-VERDE L, et al. Epigenetic regulation of alkaline phosphatase in human cells of the osteoblastic lineage. Bone. 2011;49(4):830-838. [30] YE L, FAN Z, YU B, et al. Histone Demethylases KDM4B and KDM6B Promote Osteogenic Differentiation of Human MSCs. Cell Stem Cell. 2018;23(6):898-899. [31] TIAN C, HUANG Y, LI Q, et al. Mettl3 Regulates Osteogenic Differentiation and Alternative Splicing of Vegfa in Bone Marrow Mesenchymal Stem Cells. Int J Mol Sci. 2019;20(3):551. [32] YAN G, YUAN Y, HE M, et al. m(6)A Methylation of Precursor-miR-320/RUNX2 Controls Osteogenic Potential of Bone Marrow-Derived Mesenchymal Stem Cells. Mol Ther Nucleic Acids. 2020;19:421-436. [33] LI D, CAI L, MENG R, et al. METTL3 Modulates Osteoclast Differentiation and Function by Controlling RNA Stability and Nuclear Export. Int J Mol Sci. 2020;21(5):1660. [34] YAO Y, BI Z, WU R, et al. METTL3 inhibits BMSC adipogenic differentiation by targeting the JAK1/STAT5/C/EBPβ pathway via an m(6)A-YTHDF2-dependent manner. FASEB J. 2019;33(6):7529-7544. [35] CARMELIET P, NG YS, NUYENS D, et al. Impaired myocardial angiogenesis and ischemic cardiomyopathy in mice lacking the vascular endothelial growth factor isoforms VEGF164 and VEGF188. Nat Med. 1999;5(5):495-502. [36] QIAO H, ZHANG R, GAO L, et al. Molecular Imaging for Comparison of Different Growth Factors on Bone Marrow-Derived Mesenchymal Stromal Cells’ Survival and Proliferation In Vivo. Biomed Res Int. 2016; 2016:1363902. [37] ZHANG J, GUAN J, QI X, et al. Dimethyloxaloylglycine Promotes the Angiogenic Activity of Mesenchymal Stem Cells Derived from iPSCs via Activation of the PI3K/Akt Pathway for Bone Regeneration. Int J Biol Sci. 2016;12(6):639-652. [38] ZHANG J, LIU X, LI H, et al. Exosomes/tricalcium phosphate combination scaffolds can enhance bone regeneration by activating the PI3K/Akt signaling pathway. Stem Cell Res Ther. 2016;7(1):136. [39] WU Y, XIE L, WANG M, et al. Mettl3-mediated m(6)A RNA methylation regulates the fate of bone marrow mesenchymal stem cells and osteoporosis. Nat Commun. 2018;9(1):4772. [40] JIN M, SONG S, GUO L, et al. Increased serum GDF11 concentration is associated with a high prevalence of osteoporosis in elderly native Chinese women. Clin Exp Pharmacol Physiol. 2016;43(11):1145-1147. [41] LU Q, TU ML, LI CJ, et al. GDF11 Inhibits Bone Formation by Activating Smad2/3 in Bone Marrow Mesenchymal Stem Cells. Calcif Tissue Int. 2016;99(5):500-509. [42] LIU W, ZHOU L, ZHOU C, et al. GDF11 decreases bone mass by stimulating osteoclastogenesis and inhibiting osteoblast differentiation. Nat Commun. 2016;7:12794. [43] SHEN GS, ZHOU HB, ZHANG H, et al. The GDF11-FTO-PPARγ axis controls the shift of osteoporotic MSC fate to adipocyte and inhibits bone formation during osteoporosis. Biochim Biophys Acta Mol Basis Dis. 2018;1864(12):3644-3654. [44] DIEPPE PA, LOHMANDER LS. Pathogenesis and management of pain in osteoarthritis. Lancet. 2005;365(9463):965-973. [45] HWANG HS, KIM HA. Chondrocyte Apoptosis in the Pathogenesis of Osteoarthritis. Int J Mol Sci. 2015;16(11):26035-26054. [46] LIU Q, LI M, JIANG L, et al. METTL3 promotes experimental osteoarthritis development by regulating inflammatory response and apoptosis in chondrocyte. Biochem Biophys Res Commun. 2019; 516(1):22-27. [47] ZHANG Y, GU X, LI D, et al. METTL3 Regulates Osteoblast Differentiation and Inflammatory Response via Smad Signaling and MAPK Signaling. Int J Mol Sci. 2019;21(1):199. [48] KASSEM A, HENNING P, LUNDBERG P, et al. Porphyromonas gingivalis Stimulates Bone Resorption by Enhancing RANKL (Receptor Activator of NF-κB Ligand) through Activation of Toll-like Receptor 2 in Osteoblasts. J Biol Chem. 2015;290(33):20147-20158. [49] SAINT-PASTOU TC, GASQUE P. Bone responses in health and infectious diseases: A focus on osteoblasts. J Infect. 2017;75(4):281-292. [50] ZHANG Y, XU J, RUAN YC, et al. Implant-derived magnesium induces local neuronal production of CGRP to improve bone-fracture healing in rats. Nat Med. 2016;22(10):1160-1169. [51] SON JH, CHO YC, SUNG IY, et al. Melatonin promotes osteoblast differentiation and mineralization of MC3T3-E1 cells under hypoxic conditions through activation of PKD/p38 pathways. J Pineal Res. 2014;57(4):385-392. [52] LI D, LIU J, GUO B, et al. Osteoclast-derived exosomal miR-214-3p inhibits osteoblastic bone formation. Nat Commun. 2016;7:10872. [53] 薛徽, 孙瑶. 影响骨折愈合的生物因素研究新进展[J]. 口腔医学, 2018,38(11):1043-1047. [54] MI B, XIONG Y, YAN C, et al. Methyltransferase-like 3-mediated N6-methyladenosine modification of miR-7212-5p drives osteoblast differentiation and fracture healing. J Cell Mol Med. 2020;24(11): 6385-6396. [55] VIJAYAMURUGAN N, BAKHSHI S. Review of management issues in relapsed osteosarcoma. Expert Rev Anticancer Ther. 2014;14(2):151-161. [56] HUANG Y, SU R, SHENG Y, et al. Small-Molecule Targeting of Oncogenic FTO Demethylase in Acute Myeloid Leukemia. Cancer Cell. 2019;35(4):677-691. [57] ZHANG C, SAMANTA D, LU H, et al. Hypoxia induces the breast cancer stem cell phenotype by HIF-dependent and ALKBH5-mediated m⁶A-demethylation of NANOG mRNA. Proc Natl Acad Sci U S A. 2016; 113(14):E2047-E2056. [58] BARBIERI I, TZELEPIS K, PANDOLFINI L, et al. Promoter-bound METTL3 maintains myeloid leukaemia by m(6)A-dependent translation control. Nature. 2017;552(7683):126-131. [59] MIAO W, CHEN J, JIA L, et al. The m6A methyltransferase METTL3 promotes osteosarcoma progression by regulating the m6A level of LEF1. Biochem Biophys Res Commun. 2019;516(3):719-725. [60] ZHOU L, YANG C, ZHANG N, et al. Silencing METTL3 inhibits the proliferation and invasion of osteosarcoma by regulating ATAD2. Biomed Pharmacother. 2020;125:109964. [61] LI J, RAO B, YANG J, et al. Dysregulated m6A-Related Regulators Are Associated With Tumor Metastasis and Poor Prognosis in Osteosarcoma. Front Oncol. 2020;10:769. |
| [1] | An Yang, Liao Yinan, Xie Chengxin, Li Qinglong, Huang Ge, Jin Xin, Yin Dong. Mechanism of Inulae flos in the treatment of osteoporosis: an analysis based on network pharmacology [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(在线): 1-8. |
| [2] | Huang Dengcheng, Wang Zhike, Cao Xuewei. Comparison of the short-term efficacy of extracorporeal shock wave therapy for middle-aged and elderly knee osteoarthritis: a meta-analysis [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(9): 1471-1476. |
| [3] | Peng Zhihao, Feng Zongquan, Zou Yonggen, Niu Guoqing, Wu Feng. Relationship of lower limb force line and the progression of lateral compartment arthritis after unicompartmental knee arthroplasty with mobile bearing [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(9): 1368-1374. |
| [4] | Huang Zexiao, Yang Mei, Lin Shiwei, He Heyu. Correlation between the level of serum n-3 polyunsaturated fatty acids and quadriceps weakness in the early stage after total knee arthroplasty [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(9): 1375-1380. |
| [5] | Li Jiacheng, Liang Xuezhen, Liu Jinbao, Xu Bo, Li Gang. Differential mRNA expression profile and competitive endogenous RNA regulatory network in osteoarthritis [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(8): 1212-1217. |
| [6] | Geng Qiudong, Ge Haiya, Wang Heming, Li Nan. Role and mechanism of Guilu Erxianjiao in treatment of osteoarthritis based on network pharmacology [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(8): 1229-1236. |
| [7] | Liu Xiangxiang, Huang Yunmei, Chen Wenlie, Lin Ruhui, Lu Xiaodong, Li Zuanfang, Xu Yaye, Huang Meiya, Li Xihai. Ultrastructural changes of the white zone cells of the meniscus in a rat model of early osteoarthritis [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(8): 1237-1242. |
| [8] | Li Zhongfeng, Chen Minghai, Fan Yinuo, Wei Qiushi, He Wei, Chen Zhenqiu. Mechanism of Yougui Yin for steroid-induced femoral head necrosis based on network pharmacology [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(8): 1256-1263. |
| [9] | Shu Wenbo, Chen Mengchi, Li Hua, Huang Liqian, Huang Binbin, Zhang Wenhai, Wu Yachen, Wang Zefeng, Li Qiaoli, Liu Peng. Correlation between body fat distribution and characteristics of daily physical activity in college students [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(8): 1277-1283. |
| [10] | Wu Xun, Meng Juanhong, Zhang Jianyun, Wang Liang. Concentrated growth factors in the repair of a full-thickness condylar cartilage defect in a rabbit [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(8): 1166-1171. |
| [11] | Tang Hui, Yao Zhihao, Luo Daowen, Peng Shuanglin, Yang Shuanglin, Wang Lang, Xiao Jingang. High fat and high sugar diet combined with streptozotocin to establish a rat model of type 2 diabetic osteoporosis [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(8): 1207-1211. |
| [12] | Hou Guangyuan, Zhang Jixue, Zhang Zhijun, Meng Xianghui, Duan Wen, Gao Weilu. Bone cement pedicle screw fixation and fusion in the treatment of degenerative spinal disease with osteoporosis: one-year follow-up [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(6): 878-883. |
| [13] | Li Shibin, Lai Yu, Zhou Yi, Liao Jianzhao, Zhang Xiaoyun, Zhang Xuan. Pathogenesis of hormonal osteonecrosis of the femoral head and the target effect of related signaling pathways [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(6): 935-941. |
| [14] | He Xiangzhong, Chen Haiyun, Liu Jun, Lü Yang, Pan Jianke, Yang Wenbin, He Jingwen, Huang Junhan. Platelet-rich plasma combined with microfracture versus microfracture in the treatment of knee cartilage lesions: a meta-analysis [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(6): 964-969. |
| [15] | Hua Haotian, Zhao Wenyu, Zhang Lei, Bai Wenbo, Wang Xinwei. Meta-analysis of clinical efficacy and safety of antibiotic artificial bone in the treatment of chronic osteomyelitis [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(6): 970-976. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||