Chinese Journal of Tissue Engineering Research ›› 2026, Vol. 30 ›› Issue (34): 8878-8888.doi: 10.12307/2026.890
Previous Articles Next Articles
Therapeutic targets for knee osteoarthritis: identification via a bioinformatics approach
Chen Cai1, Hong Zhongyuan1, Deng Huaidong1, Zeng Qin1, Chen Jiancong2
- 1Department of Orthopedics I, 2Department of Rehabilitation Medicine, Dongguan Hospital of Traditional Chinese Medicine Affiliated to Guangzhou University of Chinese Medicine, Dongguan 523000, Guangdong Province, China
-
Received:2025-09-17Revised:2026-02-12Online:2026-12-08Published:2026-04-11 -
Contact:Hong Zhongyuan, MS, Associate chief physician, Department of Orthopedics I, Dongguan Hospital of Traditional Chinese Medicine Affiliated to Guangzhou University of Chinese Medicine, Dongguan 523000, Guangdong Province, China -
About author:Chen Cai, MS, Department of Orthopedics I, Dongguan Hospital of Traditional Chinese Medicine Affiliated to Guangzhou University of Chinese Medicine, Dongguan 523000, Guangdong Province, China -
Supported by:Guangdong Provincial Administration of Traditional Chinese Medicine Research Project, No. 20231365 (to HZY); Dongguan Municipal Science and Technology Bureau Social Development Science and Technology Project, No. 20221800900102 (to DHD)
CLC Number:
Cite this article
Chen Cai, Hong Zhongyuan, Deng Huaidong, Zeng Qin, Chen Jiancong. Therapeutic targets for knee osteoarthritis: identification via a bioinformatics approach[J]. Chinese Journal of Tissue Engineering Research, 2026, 30(34): 8878-8888.
share this article
Add to citation manager EndNote|Reference Manager|ProCite|BibTeX|RefWorks
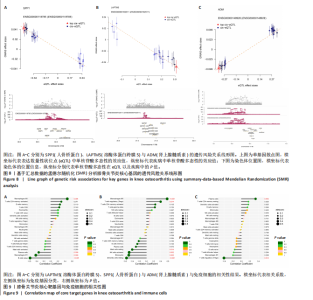
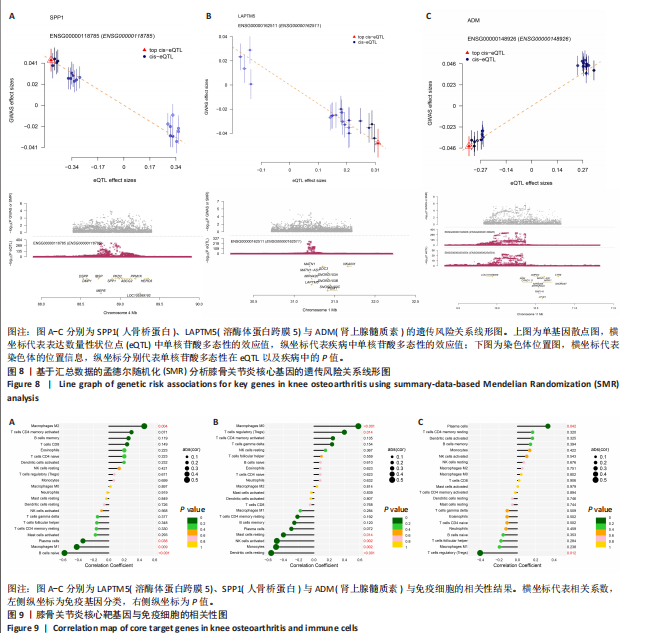
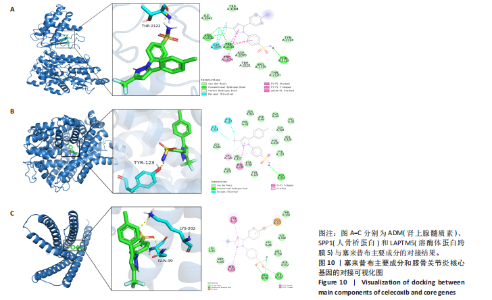
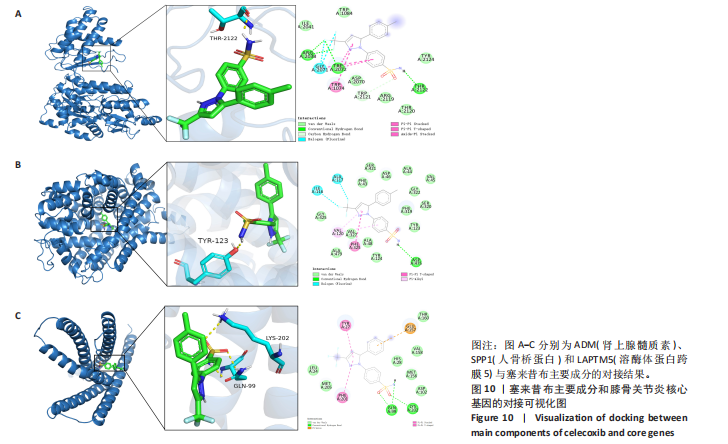
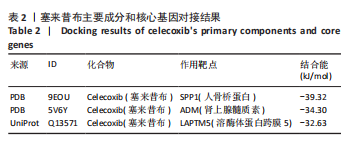
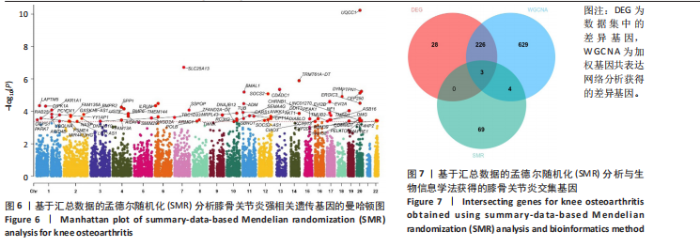
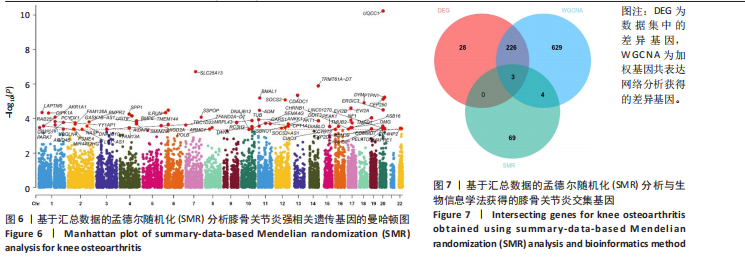
2.5 SMR分析获得膝骨关节炎强相关遗传基因 基于P < 0.05、FDR < 0.05、HEIDI 检验P > 0.05为条件,SMR分析共获得了76个基因与eQTL变异显著相关,利用R语言软件构建可视化曼哈顿图,如图6所示。 2.6 生物信息学法与SMR分析的交集基因 将生物信息学法与SMR分析获得的基因通过韦恩分析法取交集,SPP1(人骨桥蛋白?)、LAPTM5(溶酶体蛋白跨膜5)与ADM(肾上腺髓质素)是此次研究得到的关键基因,如图7所示。 2.7 核心基因与膝骨关节炎的遗传风险关系 构建SPP1、LAPTM5与ADM与膝骨关节炎的遗传共定位分析,并构建风险关系线形图,如图8所示。SMR分析结果表明,SPP1、LAPTM5 eQTL中单核苷酸多态性的效应值与GWAS数据库中膝骨关节炎的效应值呈负相关,ADM eQTL中的单核苷酸多态性效应值与GWAS数据库中膝骨关节炎的效应值呈正相关,表明遗传决定的SPP1与LAPTM5是膝骨关节炎的“保护”基因,而ADM是膝骨关节炎的致病基因。 2.8 膝骨关节炎核心基因的免疫浸润分析结果 采用CIBERSORT算法获得了膝骨关节炎中22种免疫细胞类型的比例,结果表明,SPP1与巨噬细胞M0、调节性T细胞呈正相关,与静息树突状细胞、单核细胞、活化的NK细胞和静息肥大细胞呈负相关;LAPTM5与巨噬细胞M2呈正相关,与幼稚B细胞、巨噬细胞M1和浆细胞呈负相关;ADM与浆细胞呈正相关,与调节性T细胞呈负相关。结果如图9所示。 2.9 分子对接与分子动力学模拟结果 2.9.1 分子对接结果 分子对接结果表明核心基因与塞来昔布活性成分均能稳定结合,一般来说当结合能小于-20.92 kJ/mol时表明二者具有较好的结合活性,小于-29.29 kJ/mol表明二者有强烈的结合活性[12],结合能越低说明两者结合活性越强、亲和力越高、构象也越稳定,表明这些配体与相应的受体靶蛋白具有良好的结合亲和力,详细结果见表2,可视化结果见图10。"
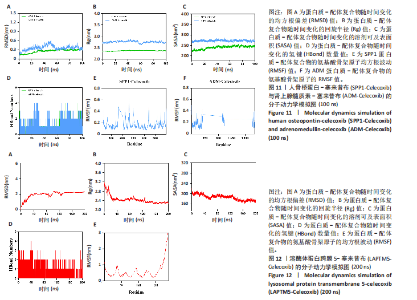
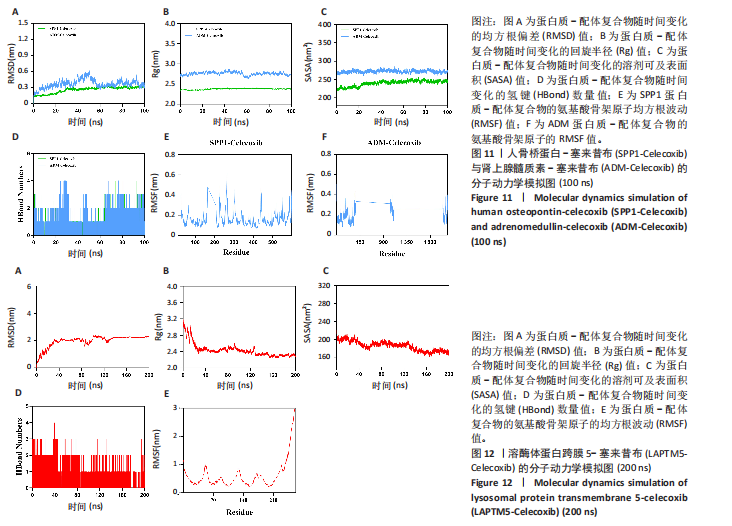
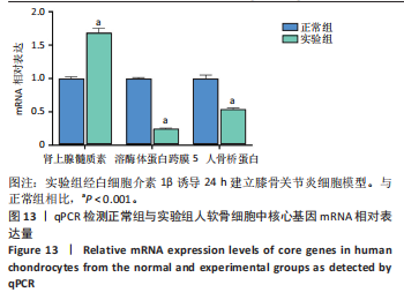
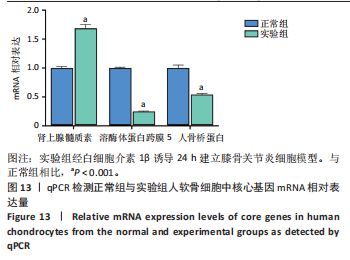
2.9.2 分子动力学模拟结果 核心基因与塞来昔布的分子动力学模拟图,见图11,12。 均方根偏差是衡量蛋白质和配体构象稳定性的良好指标,也是衡量原子位置与起始位置偏差程度的指标,均方根偏差越小,构象的稳定性越好。因此,利用均方根偏差对仿真系统的平衡性进行了评估。结果显示,SPP1-塞来昔布复合物体系的均方根偏差在30 ns后达到平衡,最终在0.33 nm上下波动。ADM-塞来昔布复合物体系的均方根偏差在98 ns后达到平衡,最终在0.31 nm上下波动。LAPTM5-塞来昔布复合物体系的均方根偏差在40 ns后达到平衡,最终在2.27 nm上下波动。 回转半径可以用来描述整体结构的变化情况,可用于表征蛋白质结构的紧密程度,回转半径变化越大表明体系越膨胀。结果显示,SPP1-塞来昔布、ADM-塞来昔布和LAPTM5-塞来昔布复合物体系在运动过程中呈现轻微波动,说明小分子-靶蛋白复合物在运动过程中发生了构象变化。 溶剂可及表面积是评估蛋白质表面积的指标,此次模拟计算靶蛋白和小分子之间的溶剂可及表面积,结果显示,SPP1-塞来昔布、ADM-塞来昔布和LAPTM5-塞来昔布复合物体系呈现轻微波动,证明结合小分子会影响结合微环境,并导致一定程度上溶剂可及表面积的变化。 氢键在配体与蛋白质的结合中起着重要的作用。结果显示,SPP1-塞来昔布小分子与靶蛋白之间的氢键数量为0-5,在大多数情况下复合物大约有2个氢键;ADM-塞来昔布小分子与靶蛋白之间的氢键数量为0-4,在大多数情况下复合物大约有2个氢键;LAPTM5-塞来昔布小分子与靶蛋白之间的氢键数量为0-3,在大多数情况下复合物大约有1个氢键,表明这种配体与靶蛋白具有良好的氢键相互作用。 均方根涨落值可以表示蛋白质中氨基酸残基的柔性大小。SPP1-塞来昔布和ADM-塞来昔布复合物的均方根涨落值相对较低(大多在0.5 nm以下),因此灵活性更低,稳定性较高。LAPTM5-塞来昔布复合物的均方根涨落值较高(大多在0.7 nm以下),因此复合物具有较高的柔性。 2.10 核心基因的细胞实验验证结果 qPCR检测结果表明,实验组细胞中ADM mRNA表达高于正常组(P < 0.001),SPP1与LAPTM5 mRNA表达低于正常组(P < 0.001),结果如图13所示。"
| [1] ZHANG Y, JI Q. Current advances of photobiomodulation therapy in treating knee osteoarthritis. Front Cell Dev Biol. 2023;11:1286025. [2] 马兆臣,肖自青,张楚,等.从空间异质性视角研究全杜仲胶囊治疗膝骨关节炎的作用机制[J].中国中药杂志,2025,50(8):2209-2216. [3] LIU Y, DA W, XU MJ, et al. Single-cell transcriptomics reveals novel chondrocyte and osteoblast subtypes and their role in knee osteoarthritis pathogenesis. Signal Transduct Target Ther. 2025;10(1):40. [4] 李裕柳,谷天,熊倩,等.基于网络药理学和体外实验验证探讨膝痹康方治疗膝骨关节炎的作用机制 [J].中国老年学杂志,2025,45(12):2882-2890. [5] TANG K, SUN L, CHEN L, et al. Bioinformatics Analysis and Experimental Validation of Mitochondrial Autophagy Genes in Knee Osteoarthritis. Int J Gen Med. 2024;17:639-650. [6] DIAO Z, GUO D, ZHANG J, et al. Causal relationship between modifiable risk factors and knee osteoarthritis: a Mendelian randomization study. Front Med (Lausanne). 2024;11:1405188. [7] LIN T, ZHOU F, MAO H, et al. Vitamin D and idiopathic pulmonary fibrosis: a two-sample mendelian randomization study. BMC Pulm Med. 2023;23(1):309. [8] WANG J, YANG M, XU K, et al. The causal associations between growth factors and constipation: a two-sample Mendelian randomization study. Front Physiol. 2023;14:1204146. [9] LI Y, SUNDQUIST K, ZHANG N, et al. Mitochondrial related genome-wide Mendelian randomization identifies putatively causal genes for multiple cancer types. EBioMedicine. 2023;88:104432. [10] JO S, KIM T, IYER VG, et al. CHARMM-GUI: a web-based graphical user interface for CHARMM. J Comput Chem. 2008;29(11):1859-1865. [11] SUN Y, SU S, LI M, et al. Inhibition of miR-182-5p Targets FGF9 to Alleviate Osteoarthritis. Anal Cell Pathol (Amst). 2023;2023:5911546. [12] CHEN L, CHEN S, RONG Y, et al. Identification and evaluation of antioxidant peptides from highland barley distiller’s grains protein hydrolysate assisted by molecular docking. Food Chem. 2024;434:137441. [13] HASIN Y, SELDIN M, LUSIS A. Multi-omics approaches to disease. Genome Biol. 2017;18(1): 83. [14] XU Z, YANG J, MA Y, et al. Exploring of bladder cancer immune-related genes and potential therapeutic targets based on transcriptomic data and Mendelian randomization analysis. Front Immunol. 2025;16:1607098. [15] 吴昌强,韦敏,郑一鸣,等.干扰miR-483-5p对骨关节炎软骨细胞增殖及炎症因子的影响[J].中国老年学杂志,2023,43(12):3040-3042. [16] TANG W, YIN JB, LIN RG, et al. Rapgef3 modulates macrophage reprogramming and exacerbates synovitis and osteoarthritis under excessive mechanical loading. iScience. 2025;28(5):112131. [17] XIONG G, YANG Y, GUO M. Effect of resveratrol on abnormal bone remodeling and angiogenesis of subchondral bone in osteoarthritis. Int J Clin Exp Pathol. 2021;14(4):417-425. [18] FAUST HJ, ZHANG H, HAN J, et al. IL-17 and immunologically induced senescence regulate response to injury in osteoarthritis. J Clin Invest. 2020;130(10): 5493-5507. [19] YUE B, XIONG D, CHEN J, et al. SPP1 induces idiopathic pulmonary fibrosis and NSCLC progression via the PI3K/Akt/mTOR pathway. Respir Res. 2024;25(1):362. [20] LIN Z, TIAN XY, HUANG XX, et al. microRNA-186 inhibition of PI3K-AKT pathway via SPP1 inhibits chondrocyte apoptosis in mice with osteoarthritis. J Cell Physiol. 2019;234(5):6042-6053. [21] LV C, LI Y, XU J, et al. Association of SPP1 promoter variants with hip osteoarthritis susceptibility in Chinese population. Gene. 2015;564(1):9-13. [22] QU Y, WANG Y, WANG S, et al. A comprehensive analysis of single-cell RNA transcriptome reveals unique SPP1+ chondrocytes in human osteoarthritis. Comput Biol Med. 2023;160:106926. [23] 白洁,石鹏飞,周胜虎,等.骨质疏松症患者血清CYR61、CX3CL1和OPN水平及与骨代谢水平的相关性分析[J].检验医学与临床,2025,22(14):1938-1942. [24] LIU Y, FU L, LIU Z. The Role and Clinical Relevance of Osteopontin in Allergic Airway Diseases. J Clin Med. 2023;12(6):2433. [25] BAI RJ, LIU D, LI YS, et al. OPN inhibits autophagy through CD44, integrin and the MAPK pathway in osteoarthritic chondrocytes. Front Endocrinol. 2022;13:919366. [26] GONG S, XIANG K, CHEN L, et al. Integrated bioinformatics analysis identified leucine rich repeat containing 15 and secreted phosphoprotein 1 as hub genes for calcific aortic valve disease and osteoarthritis. IET Syst Biol. 2024;18(3):77-91. [27] YANG L, YU X, LIU M, et al. A comprehensive analysis of biomarkers associated with synovitis and chondrocyte apoptosis in osteoarthritis. Front Immunol. 2023;14:1149686. [28] LUO W, LIN Z, YUAN Y, et al. Osteopontin (OPN) alleviates the progression of osteoarthritis by promoting the anabolism of chondrocytes. Genes Dis. 2023; 10(4):1714-1725. [29] LIU Q, ZENG H, YUAN Y, et al. Osteopontin inhibits osteoarthritis progression via the OPN/CD44/PI3K signal axis. Genes Dis. 2022;9(1):128-139. [30] SHANG H, HAO Y, HU W, et al. OPN gene locus is associated with the risk of knee osteoarthritis: a case-control study. Biosci Rep. 2019;39(2):BSR20182023. [31] ZHANG MM, LIANG MJ, ZHANG DM, et al. The function and mechanism of LAPTM5 in diseases. Biomed Pharmacother. 2024;178:117237. [32] XING L, LI Y, LI W, et al. Expression of RUNX2/LAPTM5 in the Induction of MC3T3-e1 Mineralization and Its Possible Relationship with Autophagy. Tissue Eng Regen Med. 2022;19(6):1223-1235. [33] TIAN K, HE X, LIN X, et al. Unveiling the Role of Sik1 in Osteoblast Differentiation: Implications for Osteoarthritis. Mol Cell Biol. 2024;44(10):411-428. [34] HAN S, JIN X, HU T, et al. LAPTM5 regulated by FOXP3 promotes the malignant phenotypes of breast cancer through activating the Wnt/β‑catenin pathway. Oncol Rep. 2023;49(3):60. [35] LI B, JIANG T, WANG J, et al. Cuprorivaite microspheres inhibit cuproptosis and oxidative stress in osteoarthritis via Wnt/β-catenin pathway. Mater Today Bio. 2024;29:101300. [36] SONG Z, WANG X, HE L, et al. Suppression of lysosomal-associated protein transmembrane 5 ameliorates cardiac function and inflammatory response by inhibiting the nuclear factor-kappa B (NF-κB) pathway after myocardial infarction in mice. Exp Anim. 2022;71(4):415-425. [37] SUN W, YUE J, CUI Y, et al. Wedelolactone alleviates inflammation and cartilage degeneration by suppressing the NF-κB signaling pathway in osteoarthritis. Int Immunopharmacol. 2024;143(Pt 1):113359. [38] LIU L, HUANG R, MA D, et al. Correlation of Adrenomedullin Concentrations with Knee Osteoarthritis Grade. Med Sci Monit. 2016;22:2775-2778. [39] ABD ELAZEEM MI, AHMED AB, MOHAMED RA, et al. Serum level of Adrenomedullin in patients with primary knee osteoarthritis; relation to disease severity. Egypt Rheumatol Rehabil. 2021;48:1-7. [40] ZHANG Z, LIU W, XIONG J, et al. Candidate Marker Genes for Diagnosis of Osteoarthritis and Prediction of Their Regulatory Mechanisms. Folia Biol (Praha). 2023;69(1):22-33. [41] MOSSER DM, EDWARDS JP. Exploring the full spectrum of macrophage activation. Nat Rev Immunol. 2008;8(12):958-969. [42] KRAUS VB, MCDANIEL G, HUEBNER JL, et al. Direct in vivo evidence of activated macrophages in human osteoarthritis. Osteoarthritis Cartilage. 2016;24(9):1613-1621. [43] ZHANG J, CHENG F, RONG G, et al. Circular RNA hsa_circ_0005567 overexpression promotes M2 type macrophage polarization through miR-492/SOCS2 axis to inhibit osteoarthritis progression. Bioengineered. 2021;12(1):8920-8930. [44] MA Y, YANG H, ZONG X, et al. Artificial M2 macrophages for disease-modifying osteoarthritis therapeutics. Biomaterials. 2021;274:120865. [45] NEES TA, ZHANG JA, PLATZER H, et al. Infiltration Profile of Regulatory T Cells in Osteoarthritis-Related Pain and Disability. Biomedicines. 2022;10(9):2111. [46] KELLER LE, TAIT WOJNO ED, BEGUM L, et al. Regulatory T cells provide chondroprotection through increased TIMP1, IL-10 and IL-4, but cannot mitigate the catabolic effects of IL-1β and IL-6 in a tri-culture model of osteoarthritis. Osteoarthr Cartil Open. 2021;3(3):100193. [47] GUILLEM-LLOBAT P, MARÍN M, ROULEAU M, et al. New Insights into the Pro-Inflammatory and Osteoclastogenic Profile of Circulating Monocytes in Osteoarthritis Patients. Int J Mol Sci. 2024;25(3):1710. [48] LUO P, YUAN Q, WAN X, et al. Effects of Immune Cells and Cytokines on Different Cells in OA. J Inflamm Res. 2023;16:2329-2343. [49] WANG AA, GOMMERMAN JL, ROJAS OL. Plasma Cells: From Cytokine Production to Regulation in Experimental Autoimmune Encephalomyelitis. J Mol Biol. 2021;433(1):166655. [50] XIE X, DOODY GM, SHUWEIHDI F, et al. B-cell capacity for expansion and differentiation into plasma cells are altered in osteoarthritis. Osteoarthritis Cartilage. 2023;31(9):1176-1188. |
| [1] | Chen Qiuhan, Yang Long, Yuan Daizhu, Wu Zhanyu, Zou Zihao, Ye Chuan. Peri-knee osteotomy for treatment of knee osteoarthritis: optimization of treatment strategies [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(9): 2303-2312. |
| [2] | Guo Ying, Tian Feng, Wang Chunfang. Potential drug targets for the treatment of rheumatoid arthritis: large sample analysis from European databases [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(6): 1549-1557. |
| [3] | Li Linzhen, Jiao Hongzhuo, Chen Weinan, Zhang Mingzhe, Wang Jianlong, Zhang Juntao. Effect of icariin-containing serum on lipopolysaccharide-induced inflammatory damage in human chondrocytes [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(6): 1368-1374. |
| [4] | Gu Fucheng, Yang Meixin, Wu Weixin, Cai Weijun, Qin Yangyi, Sun Mingyi, Sun Jian, Geng Qiudong, Li Nan. Effects of Guilu Erxian Glue on gut microbiota in rats with knee osteoarthritis: machine learning and 16S rDNA analysis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(4): 1058-1072. |
| [5] | Zhao Canbin, Zeng Ping, Shi Weiqi, Liu Jinfu, Ding Qiang, Guo Liang, Wang Weiwei, Tao Hongcheng, Guo Yafeng, Qin Ying. Mechanism by which luteolin regulates macrophage polarization in the treatment of knee osteoarthritis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(34): 8868-8877. |
| [6] | Wang Yanfei, Jin Lianhai, Li Qingya, Fu Yuanfei, Tan Huangsheng, Deng Pengwei, Gao Kun. Synovial fluid exosome-mediated crosstalk between synoviocytes and chondrocytes in development and progression of knee osteoarthritis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(34): 9032-9040. |
| [7] | Wu Ruiqi, Dong Panfeng, Zhang Hongrui, Lyu Yongbin, Peng Qinglin, Zhuo Yinghong, Chen Yueping . Different physical factor therapies for knee osteoarthritis: a network meta-analysis of efficacy and safety [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(34): 9056-9066. |
| [8] | Li Xiaomin, Tian Xiangdong, Wang Chaolu. High tibial osteotomy on a single plane: femorofibular angle as a reference marker for mechanical axis correction [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(3): 570-576. |
| [9] | Li Zhengpeng, Shao Weigang, Zeng Hao, Xiang Kelin, Zhang Botao, Zou Shunyi, Chen Sheng, Qi Wen. Osteoarthritis characteristic genes and prediction of targeted food-medicine homology traditional Chinese medicine: bioinformatics analysis and kinetic simulation [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(29): 7739-7748. |
| [10] | Zhao Ruikai, Wang Yu, Guo Xiaohui, Sun Zehua, Wang Xu. Mendelian randomization analysis identifies potential drug targets for spinal osteoarthritis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(29): 7749-7754. |
| [11] | Yong Qiao, Sun Xin, Wang Guoyou, Zhang Lei, Shen Huarui, Liu Huan, Guan Taiyuan. Shaoyang Shenggu Fang inhibits oxidative stress and delays cartilage aging in rats with knee osteoarthritis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(28): 7251-7259. |
| [12] | Deng Qian, Peng Zining, Meng Fanyu, Huang Yuanbo, Liu Nian, Yan Weitian, Li Zhaofu, Peng Jiangyun. Molecular mechanisms of Toddalia asiatica against rheumatoid arthritis: bioinformatics and molecular dynamics simulation [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(28): 7267-7279. |
| [13] | Bai Xue, Tian Yukui, Guo Lei, Shi Mengni, Cui Xiaofeng, Wang Cheng, Li Jingxian, Zhu Qingguang, Liu Junchang. Construction of an early knee osteoarthritis rat model: CatWalk-based gait analysis and evaluation [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(28): 7280-7286. |
| [14] | Chen Ping, Lu Hongxu, Xilinbaoleri. Finite element analysis of effects of opening wedge high tibial osteotomy on knee joint and internal fixation stress [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(27): 6969-6977. |
| [15] | He Yanyan, Ge Jirong, Li Shengqiang, Chen Xuan, Huang Jingwen, Huang Xiaobin, Xue Lipeng. Transcriptomic analysis of expression and function of differential genes in traditional Chinese medicine syndromes of postmenopausal osteoporosis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(25): 6446-6454. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||