Chinese Journal of Tissue Engineering Research ›› 2025, Vol. 29 ›› Issue (32): 7004-7014.doi: 10.12307/2025.916
Previous Articles Next Articles
Relationship between seven serum lipid traits and osteoarthritis: a large sample analysis of European population in IEU OPEN GWAS database
Wu Zhenhua1, Zhang Xiwei2, Wang Yipin2, Li Qianqian1
- 1Liaoning University of Traditional Chinese Medicine, Shenyang 110847, Liaoning Province, China; 2Affiliated Hospital, Liaoning University of Traditional Chinese Medicine, Shenyang 116600, Liaoning Province, China
-
Received:2024-10-08Accepted:2024-11-30Online:2025-11-18Published:2025-04-29 -
Contact:Zhang Xiwei, Professor, Affiliated Hospital, Liaoning University of Traditional Chinese Medicine, Shenyang 116600, Liaoning Province, China -
About author:Wu Zhenhua, Master candidate, Liaoning University of Traditional Chinese Medicine, Shenyang 110847, Liaoning Province, China -
Supported by:Key Reform Clinical Department Diagnosis and Treatment Capacity Building Project of Liaoning Province Provincial Hospitals, No. LNCCC-D53-2015 (to ZXW); Shenyang Science and Technology Planning Project, No. 22-321-34-10 (to WYP)
CLC Number:
Cite this article
Wu Zhenhua, Zhang Xiwei, Wang Yipin, Li Qianqian. Relationship between seven serum lipid traits and osteoarthritis: a large sample analysis of European population in IEU OPEN GWAS database[J]. Chinese Journal of Tissue Engineering Research, 2025, 29(32): 7004-7014.
share this article
Add to citation manager EndNote|Reference Manager|ProCite|BibTeX|RefWorks
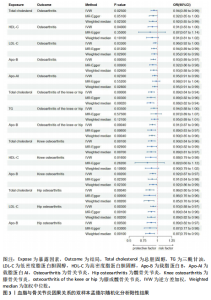
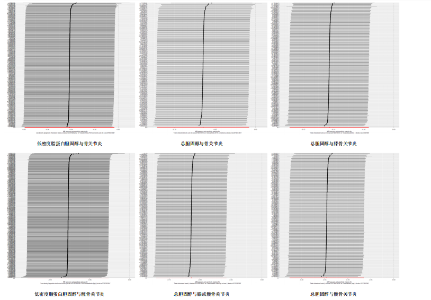
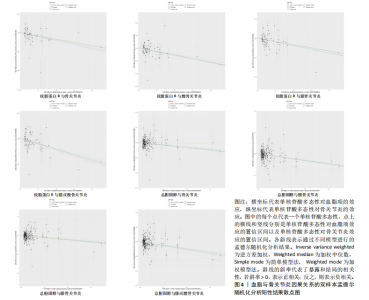
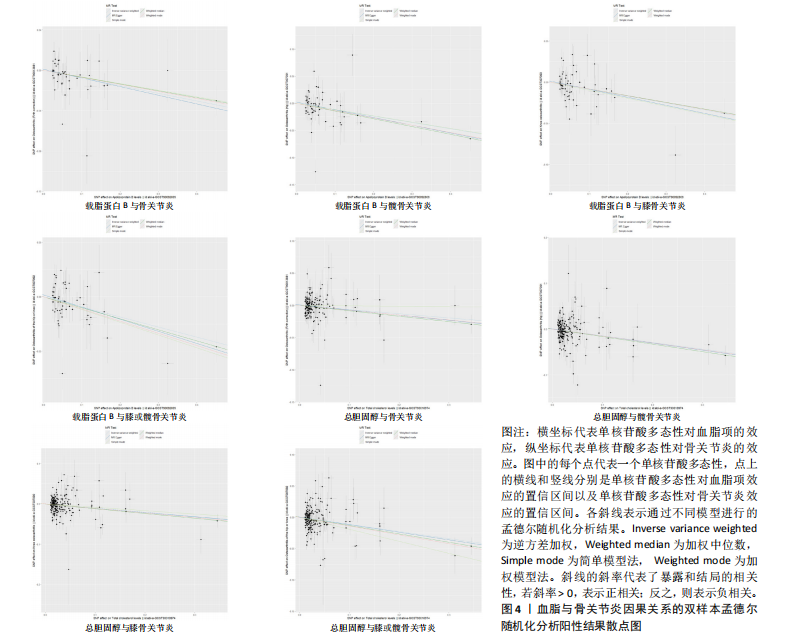
2.1 工具变量的筛选 在双样本孟德尔随机化分析中,根据阈值5×10-8的筛选条件,得出血脂7项共774个单核苷酸多态性,与暴露合并后得出血脂7项与骨关节炎、膝或髋骨关节炎和髋骨关节炎共767个单核苷酸多态性,与膝骨关节炎共768个单核苷酸多态性,作为工具变量,F统计量平均值范围为21.952-3 638.567,符合F统计量> 10。根据多效性检测,低密度脂蛋白胆固醇与膝或髋骨关节炎和膝骨关节炎存在水平多效性(P < 0.05),因此该2项结果不予采纳,其余结果无水平多效性(P > 0.05)。在多变量孟德尔随机化分析中,4个分析根据阈值5×10-8的筛选条件,得出暴露与结局工具变量的单核苷酸多态性数量分别是:1 420与284,576与192,468与156,1 002与334。 2.2 血脂7项与骨关节炎的因果效应 逆方差加权分析结果显示:①共有5种血脂项表型与骨关节炎具有显著因果关联,即总胆固醇(OR=0.937 2,95%CI=0.885 6-0.991 9,P=0.025)、低密度脂蛋白胆固醇(OR=0.959 4,95%CI=0.923 6-0.996 6,P=0.033)、高密度脂蛋白胆固醇(OR=0.911 2,95%CI=0.833 5-0.996 2,P=0.04)、载脂蛋白B(OR=0.926 7,95%CI=0.887 7-0.967 4,P=0.000 5)和载脂蛋白AI (OR=0.951 2,95%CI=0.911 0-0.993 1,P=0.023)与骨关节炎呈负相关;②共有3种血脂项表型与膝或髋骨关节炎具有显著因果关联,即总胆固醇(OR=0.892 3,95%CI=0.843 1-0.944 3,P=0.000 08)、三酰甘油(OR=0.938 5,95%CI=0.884 7-0.995 6,P=0.035)和载脂蛋白B(OR=0.911 6,95%CI=0.865 9-0.959 7,P=0.000 4)与膝或髋骨关节炎呈负相关;③共有3种血脂项表型与膝骨关节炎具有显著因果关联,即总胆固醇(OR=0.898 3,95%CI=0.841 2-0.959 3,P=0.001)、高密度脂蛋白胆固醇(OR=0.881 2,95%CI=0.794 7-0.977 0,P=0.016)和载脂蛋白B(OR= 0.919 0,95%CI=0.869 8-0.971 0,P= 0.002)与膝骨关节炎呈负相关;④共有3种血脂项表型与髋骨关节炎具有显著因果关联,即总胆固醇(OR=0.864 5,95%CI=0.797 5-0.937 3,P=0.000 4)、低密度脂蛋白胆固醇(OR=0.925 6,95%CI=0.879 5-0.974 1,P=0.003)和载脂蛋白B(OR=0.888 8,95%CI=0.817 6-0.966 3,P=0.005)与髋骨关节炎呈负相关,其余逆方差加权分析结果无统计学差异(P > 0.05);同时使用MR-Egger和加权中位数对结果进行补充说明,见图3,4。反向孟德尔随机化分析结果均无统计学差异(P > 0.05)。 逆方差加权的Bonferroni校正结果显示,载脂蛋白B(P=0.007)与骨关节炎,总胆固醇(P=0.001)、载脂蛋白B (P=0.005)与膝或髋骨关节炎,总胆固醇(P=0.019)、载脂蛋白B(P=0.036)与膝骨关节炎,总胆固醇(P=0.005)、低密度脂蛋白胆固醇(P=0.042)依然具有显著关联(P < 0.05);总胆固醇(P=0.350)、高密度脂蛋白胆固醇(P=0.573)、低密度脂蛋白胆固醇(P=0.463)、载脂蛋白AI(P=0.322)与骨关节炎,三酰甘油(P=0.494)与膝或髋骨关节炎,高密度脂蛋白胆固醇(P=0.229)与膝骨关节炎,载脂蛋白B (P=0.008)与髋骨关节炎,均不具有显"
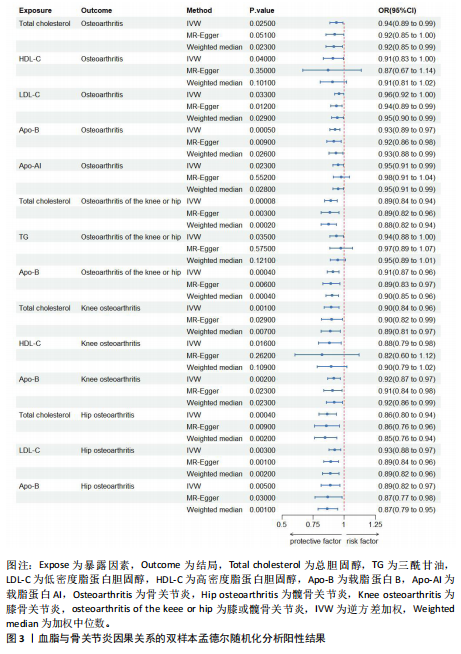
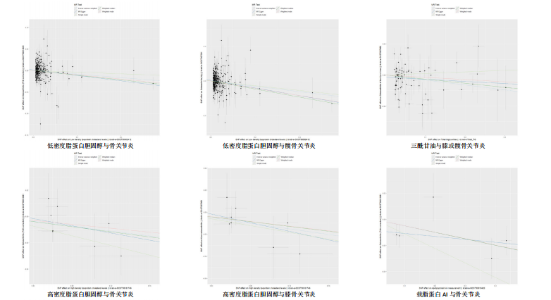
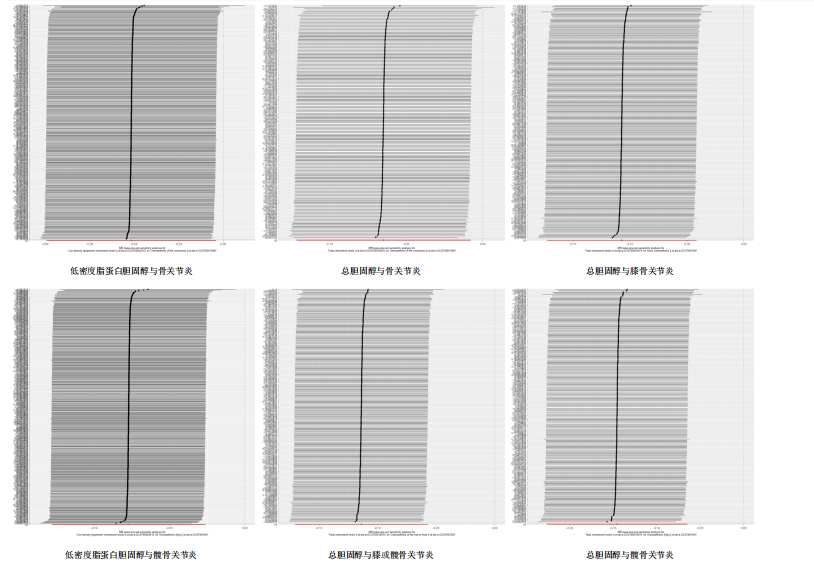
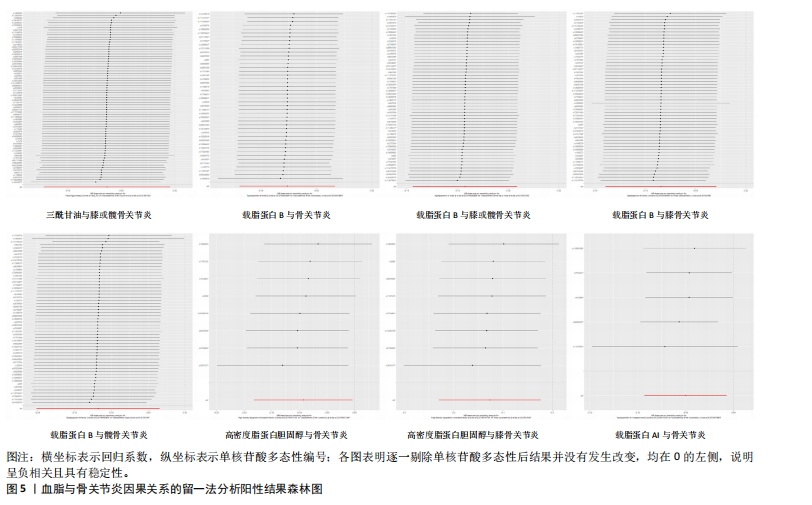
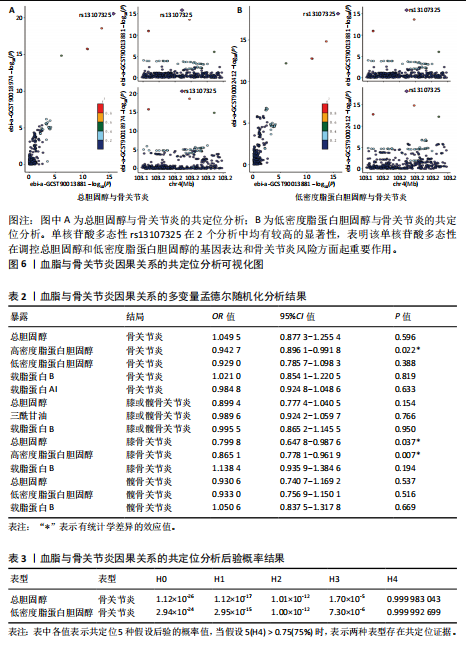
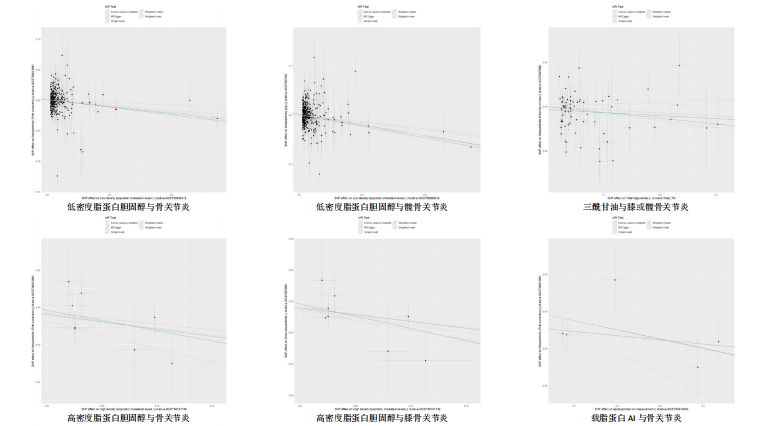
著性关联(P > 0.05)。 2.3 敏感性分析 采用Cochran Q检验对结果进行异质性检测,高密度脂蛋白胆固醇与骨关节炎(Q_P val=0.58)和膝骨关节炎(Q_P val=0.79),载脂蛋白B与骨关节炎(Q_P val=0.28),载脂蛋白AI与骨关节炎(Q_P val=0.79),均无异质性(P > 0.05),即采用固定效应模型。总胆固醇与骨关节炎(Q_P val=3.9×10-10)、膝或髋骨关节炎(Q_P val=3.1×10-20)、膝骨关节炎(Q_P val=3.4×10-15)和髋骨关节炎(Q_P val=2.8×10-13),低密度脂蛋白胆固醇与骨关节炎(Q_P val=5.7×10-15)和髋骨关节炎(Q_P val=2.3×10-12),载脂蛋白B与膝或髋骨关节炎(Q_P val=0.000 12)、膝骨关节炎(Q_P val= 0.021)和髋骨关节炎(Q_P val=7.7×10-6),三酰甘油与膝或髋骨关节炎(Q_P val= 3.6×10-9),均存在异质性(P < 0.05),因此这些采用随机效应模型。 多效性分析结果显示,根据MR-Egger截距,低密度脂蛋白胆固醇与膝或髋骨关节炎(P=0.002)和膝骨关节炎存在水平多效性(P < 0.05),因此不采纳这2项结果;总胆固醇(P=0.60)、高密度脂蛋白胆固醇(P=0.74)、低密度脂蛋白胆固醇(P=0.16)、载脂蛋白B(P=0.65)、载脂蛋白AI(P=0.37)与骨关节炎,总胆固醇(P=0.80)、三酰甘油(P=0.32)、载脂蛋白B(P=0.51)与膝或髋骨关节炎,总胆固醇(P=0.96)、高密度脂蛋白胆固醇(P=0.66)、载脂蛋白B(P=0.66)与膝骨关节炎,总胆固醇(P=0.82)、低密度脂蛋白胆固醇(P=0.13)、载脂蛋白B(P=0.60)与髋骨关节炎,均无水平多效性(P > 0.05)。 对结果进行留一法分析,对每个单核苷酸多态性消除后未对结果造成显著影响,所有结果均呈稳定负相关,见图5。 2.4 多变量孟德尔随机化分析的因果效应 逆方差加权分析结果显示:①共1个血脂项表型与骨关节炎具有显著因果关联,即高密度脂蛋白胆固醇(OR=0.942 7,95%CI=0.896 1-0.991 8,P=0.022)与骨关节炎呈负相关;②共2个血脂项表型与膝骨关节炎具有显著因果关联,即总胆固醇(OR=0.799 8,95%CI=0.647 8-0.987 6,P=0.037)和高密度脂蛋白胆固醇(OR=0.865 1,95%CI=0.778 1-0.961 9,P=0.007)与膝骨关节炎呈负相关,见表2所示。 2.5 共定位分析 以第5假设H4为效应量分析结果表明,总胆固醇(0.999 98)和低密度脂蛋白胆固醇(0.999 99)与骨关节炎具有共定位分析证据,显著的单核苷酸多态性是rs13107325,说明总胆固醇和低密度脂蛋白胆固醇与骨关节炎显著相关,并由同一个因果变异位点rs13107325所驱动,见表3、图6。"
| [1] MOTTA F, BARONE E, SICA A, et al. Inflammaging and Osteoarthritis. Clin Rev Allergy Immunol. 2023;64(2):222-238. [2] ABRAMOFF B, CALDERA FE. Osteoarthritis: Pathology, Diagnosis, and Treatment Options. Med Clin North Am. 2020;104(2): 293-311. [3] MOLNAR V, MATIŠIĆ V, KODVANJ I, et al. Cytokines and Chemokines Involved in Osteoarthritis Pathogenesis. Int J Mol Sci. 2021;22(17):9208. [4] HINES MR, GOMEZ-CONTRERAS PC, LIMAN S, et al. A reciprocal relationship between mitochondria and lipid peroxidation determines the chondrocyte intracellular redox environment. Redox Biol. 2024;75:103306. [5] NEDUNCHEZHIYAN U, VARUGHESE I, SUN AR, et al. Obesity, Inflammation, and Immune System in Osteoarthritis. Front Immunol. 2022;4(13):907750. [6] ZHENG L, ZHANG Z, SHENG P, et al. The role of metabolism in chondrocyte dysfunction and the progression of osteoarthritis. Ageing Res Rev. 2021;3(66):101249. [7] DE ROOVER A, ESCRIBANO-NÚÑEZ A, MONTEAGUDO S, et al. Fundamentals of osteoarthritis: Inflammatory mediators in osteoarthritis. Osteoarthritis Cartilage. 2023;31(10):1303-1311. [8] GU Y, JIN Q, HU J, et al. Causality of genetically determined metabolites and metabolic pathways on osteoarthritis: a two-sample mendelian randomization study. J Transl Med. 2023;21(1):357. [9] CAO C, SHI Y, ZHANG X, et al. Cholesterol-induced LRP3 downregulation promotes cartilage degeneration in osteoarthritis by targeting Syndecan-4 . Nat Commun. 2022;13(1):7139. [10] ANSARI MY, AHMAD N, HAQQI TM. Oxidative stress and inflammation in osteoarthritis pathogenesis: Role of polyphenols. Biomed Pharmacother. 2020;9(129):110452. [11] GOICOECHEA L, CONDE DE LA ROSA L, TORRES S, et al. Mitochondrial cholesterol: Metabolism and impact on redox biology and disease. Redox Biol. 2023;3(61):102643. [12] SAMPATH SJP, VENKATESAN V, GHOSH S, et al. Obesity, Metabolic Syndrome, and Osteoarthritis-An Updated Review. Curr Obes Rep. 2023;12(3):308-331. [13] ANDERSSON M, HAGLUND E, AILI K, et al. Associations between metabolic factors and radiographic knee osteoarthritis in early disease - a cross-sectional study of individuals with knee pain. BMC Musculoskelet Disord. 2022;23(1):938. [14] RICHMOND RC, DAVEY SMITH G. Mendelian Randomization: Concepts and Scope. Cold Spring Harb Perspect Med. 2022;12(1):a040501. [15] FABO T, KHAVARI P. Functional characterization of human genomic variation linked to polygenic diseases. Trends Genet. 2023;39(6):462-490. [16] PIERCE BL, BURGESS S. Efficient design for Mendelian randomization studies: subsample and 2-sample instrumental variable estimators. Am J Epidemiol. 2013; 178(7):1177-1184. [17] ZUBER V, GRINBERG NF, GILL D, et al. Combining evidence from Mendelian randomization and colocalization: Review and comparison of approaches. Am J Hum Genet. 2022;109(5):767-782. [18] ARVANITIS M, TAYEB K, STROBER BJ, et al. Redefining tissue specificity of genetic regulation of gene expression in the presence of allelic heterogeneity. Am J Hum Genet. 2022;109(2):223-239. [19] SAKAUE S, KANAI M, TANIGAWA Y, et al. A cross-population atlas of genetic associations for 220 human phenotypes. Nat Genet. 2021;53(10):1415-1424. [20] CHOUDHURY A, BRANDENBURG JT, CHIKOWORE T, et al. Meta-analysis of sub-Saharan African studies provides insights into genetic architecture of lipid traits. Nat Commun. 2022;13(1):2578. [21] KLIMENTIDIS YC, ARORA A, NEWELL M, et al. Phenotypic and Genetic Characterization of Lower LDL Cholesterol and Increased Type 2 Diabetes Risk in the UK Biobank. Diabetes. 2020;69(10):2194-2205. [22] KETTUNEN J, DEMIRKAN A, WÜRTZ P, et al. Genome-wide study for circulating metabolites identifies 62 loci and reveals novel systemic effects of LPA. Nat Commun, 2016;23(7):11122. [23] RICHARDSON TG, LEYDEN GM, WANG Q, et al. Characterising metabolomic signatures of lipid-modifying therapies through drug target mendelian randomisation. PLoS Biol. 2022;20(2):e3001547. [24] PIETZNER M, WHEELER E, CARRASCO-ZANINI J, et al. Genetic architecture of host proteins involved in SARS-CoV-2 infection. Nat Commun. 2020;11(1):6397. [25] MBATCHOU J, BARNARD L, BACKMAN J, et al. Computationally efficient whole-genome regression for quantitative and binary traits. Nat Genet. 2021;53(7):1097-1103. [26] TACHMAZIDOU I, HATZIKOTOULAS K, SOUTHAM L, et al. Identification of new therapeutic targets for osteoarthritis through genome-wide analyses of UK Biobank data. Nat Genet. 2019;51(2): 230-236. [27] BOWDEN J, HOLMES MV. Meta-analysis and Mendelian randomization: A review. Res Synth Methods. 2019;10(4):486-496. [28] LONG Y, TANG L, ZHOU Y, et al. Causal relationship between gut microbiota and cancers: a two-sample Mendelian randomisation study. BMC Med. 2023; 21(1):66. [29] BURGESS S, BOWDEN J, FALL T. Sensitivity Analyses for Robust Causal Inference from Mendelian Randomization Analyses with Multiple Genetic Variants. Epidemiology. 2017;28(1):30-42. [30] MOUNIER N, KUTALIK Z. Bias correction for inverse variance weighting Mendelian randomization. Genet Epidemiol. 2023; 47(4):314-331. [31] BURGESS S, BUTTERWORTH A, THOMPSON SG. Mendelian randomization analysis with multiple genetic variants using summarized data. Genet Epidemiol. 2013;37(7):658-665. [32] LI J, TANG M, GAO X, et al. Mendelian randomization analyses explore the relationship between cathepsins and lung cancer. Commun Biol. 2023;6(1):1019. [33] RASOOLY D, PELOSO GM. Two-Sample Multivariable Mendelian Randomization Analysis Using R. Curr Protoc. 2021;1(12): e335. [34] YAVORSKA OO, BURGESS S. MendelianRandomization: an R package for performing Mendelian randomization analyses using summarized data. Int J Epidemiol. 2017;46(6):1734-1739. [35] ZHENG J, BAIRD D, BORGES MC, et al. Recent developments in Mendelian randomization studies. Curr Epidemiol Rep. 2017;4(4):330-345. [36] BURGESS S, THOMPSON SG. Multivariable Mendelian randomization: the use of pleiotropic genetic variants to estimate causal effects. Am J Epidemiol. 2015;181(4): 251-260. [37] SANDERSON E. Multivariable Mendelian Randomization and Mediation. Cold Spring Harb Perspect Med. 2021;11(2):a038984. [38] BAKKER MK, VAN STRATEN T, CHONG M, et al. Anti-Epileptic Drug Target Perturbation and Intracranial Aneurysm Risk: Mendelian Randomization and Colocalization Study. Stroke. 2023;54(1):208-216. [39] GIAMBARTOLOMEI C, VUKCEVIC D, SCHADT EE, et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 2014;10(5):e1004.383. [40] DRIVAS TG, LUCAS A, RITCHIE MD. eQTpLot: a user-friendly R package for the visualization of colocalization between eQTL and GWAS signals. BioData Min. 2021; 14(1):32. [41] KATZ JN, ARANT KR, LOESER RF. Diagnosis and Treatment of Hip and Knee Osteoarthritis: A Review. JAMA. 2021; 325(6):568-578. [42] Bliddal H. Definition pathology and pathogenesis of osteoarthritis. Ugeskr Laeger. 2020;182(42):V06200477. [43] HEINEMEIER KM, SCHJERLING P, HEINEMEIER J, et al. Radiocarbon dating reveals minimal collagen turnover in both healthy and osteoarthritic human cartilage. Sci Transl Med. 2016;8(346):346ra90. [44] ALLEN KD, THOMA LM, GOLIGHTLY YM. Epidemiology of osteoarthritis. Osteoarthritis Cartilage. 2022;30(2): 184-195. [45] SEKULA P, DEL GRECO MF, PATTARO C, et al. Mendelian Randomization as an Approach to Assess Causality Using Observational Data. J Am Soc Nephrol. 2016;27(11): 3253-3265. [46] SCHADE DS, SHEY L, EATON RP. Cholesterol Review: A Metabolically Important Molecule. Endocr Pract. 2020;26(12):1514-1523. [47] SONG Y, LIU J, ZHAO K, et al. Cholesterol-induced toxicity: An integrated view of the role of cholesterol in multiple diseases. Cell Metab. 2021;33(10):1911-1925. [48] SCHWAGER JL, NEVITT MC, TORNER J, et al. Association of Serum Low-Density Lipoprotein, High-Density Lipoprotein, and Total Cholesterol With Development of Knee Osteoarthritis. Arthritis Care Res (Hoboken). 2022;74(2):274-280. [49] FUNCK-BRENTANO T, NETHANDER M, MOVÉRARE-SKRTIC S, et al. Causal Factors for Knee, Hip, and Hand Osteoarthritis: A Mendelian Randomization Study in the UK Biobank. Arthritis Rheumatol. 2019; 71(10):1634-1641. [50] CHEN J, TANG Z, SLOMINSKI AT, et al. Vitamin D and its analogs as anticancer and anti-inflammatory agents. Eur J Med Chem. 2020;1(207):112738. [51] ZHANG K, JI Y, DAI H, et al. High-Density Lipoprotein Cholesterol and Apolipoprotein A1 in Synovial Fluid: Potential Predictors of Disease Severity of Primary Knee Osteoarthritis. Cartilage. 2021;13(1_suppl): 1465S-1473S. [52] RYE KA, BARTER PJ. Regulation of high-density lipoprotein metabolism. Circ Res. 2014;114(1_suppl):143-156. [53] KOSTOPOULOU F, MALIZOS KN, PAPATHANASIOU I, et al. MicroRNA-33a regulates cholesterol synthesis and cholesterol efflux-related genes in osteoarthritic chondrocytes. Arthritis Res Ther. 2015;17(1):42. [54] 陈江水,杨华瑞,方志,等.脂质代谢异常与骨关节炎关系研究进展[J].海南医学,2018,29(5):682-684. [55] HINDY G, ÅKESSON KE, MELANDER O, et al. Cardiometabolic polygenic risk scores and osteoarthritis outcomes: a Mendelian randomization study using data from the Malmö diet and cancer study and the UK biobank. Arthritis Rheumatol. 2019; 71(6):925-934. [56] DIFFENDERFER MR, SCHAEFER EJ. The composition and metabolism of large and small LDL. Curr Opin Lipidol. 2014;25(3): 221-226. [57] 石银朋,奚阳,张志毅,等.血脂对骨关节炎影响研究进展[J].中国实用内科杂志,2020,40(1):67-69. [58] FELSON DT, MISRA D, LAVALLEY M, et al. Essential Fatty Acids and Osteoarthritis. Arthritis Care Res (Hoboken). 2024;76(6): 796-801. [59] HANSEN SB, WANG H. The shared role of cholesterol in neuronal and peripheral inflammation. Pharmacol Ther. 2023;9(249): 108486. [60] PHILLIPS MC. Molecular mechanisms of cellular cholesterol efflux. J Biol Chem. 2014;289(35):24020-24029. [61] LUO J, YANG H, SONG BL. Mechanisms and regulation of cholesterol homeostasis. Nature reviews. Nat Rev Mol Cell Biol. 2020; 21(4):225-245. [62] LI S, MA C, LI Y, et al. The schizophrenia-associated missense variant rs13107325 regulates dendritic spine density. Transl Psychiatry. 2022;12(1):361. [63] FRANGOS T, MARET W. Zinc and Cadmium in the Aetiology and Pathogenesis of Osteoarthritis and Rheumatoid Arthritis. Nutrients. 2020;13(1):53. [64] WANG Z, LIU M, ZHOU Y, et al. Effect of blood lipids and lipid-lowering therapies on osteoarthritis risk: A Mendelian randomization study. Front Med (Lausanne). 2022;11(9):990569. [65] MENG H, JIANG L, SONG Z, et al. Causal Associations of Circulating Lipids with Osteoarthritis: A Bidirectional Mendelian Randomization Study. Nutrients. 2022; 14(7):1327. |
| [1] | Chen Jiayong, Tang Meiling, Lu Jianqi, Pang Yan, Yang Shangbing, Mao Meiling, Luo Wenkuan, Lu Wei, Zhou Jiatan. Based on Mendelian randomization, the causal relationship between 1400 metabolites and sarcopenia and the correlation analysis of cardiovascular disease were investigated [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(在线): 1-11. |
| [2] | Zhang Yibo, Lu Jianqi, Mao Meiling, Pang Yan, Dong Li, Yang Shangbing, Xiao Xiang. Exploring the causal relationship between rheumatoid arthritis and coronary atherosclerosis: a Mendel randomized study involving serum metabolites and inflammatory factors [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(在线): 1-9. |
| [3] | Dong Tingting, Chen Tianxin, Li Yan, Zhang Sheng, Zhang Lei. Causal relationship between modifiable factors and joint sports injuries [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(9): 1953-1962. |
| [4] | Yin Lu, Jiang Chuanfeng, Chen Junjie, Yi Ming, Wang Zihe, Shi Houyin, Wang Guoyou, Shen Huarui. Effect of Complanatoside A on the apoptosis of articular chondrocytes [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(8): 1541-1547. |
| [5] | Chen Shuai, Jin Jie, Han Huawei, Tian Ningsheng, Li Zhiwei . Causal relationship between circulating inflammatory cytokines and bone mineral density based on two-sample Mendelian randomization [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(8): 1556-1564. |
| [6] |
Zhao Wensheng, Li Xiaolin, Peng Changhua, Deng Jia, Sheng Hao, Chen Hongwei, Zhang Chaoju, He Chuan.
Gut microbiota and osteoporotic fractures #br#
#br#
[J]. Chinese Journal of Tissue Engineering Research, 2025, 29(6): 1296-1304.
|
| [7] | Ma Haoyu, Qiao Hongchao, Hao Qianqian, Shi Dongbo. Causal effects of different exercise intensities on the risk of osteoarthritis [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(6): 1305-1311. |
| [8] | Li Jiatong, Jin Yue, Liu Runjia, Song Bowen, Zhu Xiaoqian, Li Nianhu . Association between thyroid function levels and phenotypes associated with sarcopenia [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(6): 1312-1320. |
| [9] | Xu Tianjie, Fan Jiaxin, Guo Xiaoling, Jia Xiang, Zhao Xingwang, Liu kainan, Wang Qian. Metformin exerts a protective effect on articular cartilage in osteoarthritis rats by inhibiting the PI3K/AKT/mTOR pathway [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(5): 1003-1012. |
| [10] | Wu Guangtao, Qin Gang, He Kaiyi, Fan Yidong, Li Weicai, Zhu Baogang, Cao Ying . Causal relationship between immune cells and knee osteoarthritis: a two-sample bi-directional Mendelian randomization analysis [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(5): 1081-1090. |
| [11] | Ma Weibang, Xu Zhe, Yu Qiao, Ouyang Dong, Zhang Ruguo, Luo Wei, Xie Yangjiang, Liu Chen. Screening and cytological validation of cartilage degeneration-related genes in exosomes from osteoarthritis synovial fluid [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(36): 7783-7789. |
| [12] | Lu Xiuli, Xu Huazhen, Chen Yuxing, Yao Nan, Hu Zixuan, Huang Dane. Mechanism of Jiangu Formula in treating osteoporosis based on osteoclast-osteoblast coupling [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(32): 6828-6835. |
| [13] | Yan Laijun, Ge Haiya, Wang Zhengming, Yang Zongrui, Niu Lifeng, Zhan Hongsheng. Mechanism by which Tongdu Huoxue Decoction inhibits macrophage inflammation to delay intervertebral disc degeneration in rats [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(32): 6851-6857. |
| [14] | Nigeayi · Aihemaiti, Yilidanna · Dilixiati, An Wei, Maimaitituxun · Tuerdi. Expression of mitochondrial creatine kinase 2 in a rat model of temporomandibular joint osteoarthritis and its role in inflammation progression [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(32): 6877-6884. |
| [15] | Wang Ziheng, Wu Shuang. Oxidative stress-related genes and molecular mechanisms after spinal cord injury: data analysis and verification based on GEO database [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(32): 6893-6904. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||