Chinese Journal of Tissue Engineering Research ›› 2026, Vol. 30 ›› Issue (34): 8889-8898.doi: 10.12307/2026.894
Previous Articles Next Articles
Weighted gene co-expression network analysis combined with machine learning identifies autophagy and senescence signature genes in osteoarthritis chondrocytes
Yang Huaqun1, Abudouainijiang·Abulimiti1, Wang Fazheng1, Maimaitishawutiaji·Maimaiti2, Li Simi1, Muhetaer·Maimaitirexiati1
- 1Department of Sports Medicine, 2Department of Spinal Orthopedics, The First People’s Hospital of Kashgar Region, Kashgar 844000, Xinjiang Uygur Autonomous Region, China
-
Received:2025-09-17Revised:2026-02-13Online:2026-12-08Published:2026-04-13 -
Contact:Muhetaer·Maimaitirexiati, Associate chief physician, Department of Sports Medicine, The First People’s Hospital of Kashgar Region, Kashgar 844000, Xinjiang Uygur Autonomous Region, China -
About author:Yang Huaqun, Associate chief physician, Department of Sports Medicine, The First People’s Hospital of Kashgar Region, Kashgar 844000, Xinjiang Uygur Autonomous Region, China Abudouainijiang·Abulimiti, MS, Attending physician, Department of Sports Medicine, The First People’s Hospital of Kashgar Region, Kashgar 844000, Xinjiang Uygur Autonomous Region, China Yang Huaqun and Abudouainijiang·Abulimiti contributed equally to this work. -
Supported by:Xinjiang Special Training Program for Minority Scientific and Technological Talents, No. 2022D03040 (to YHQ); “Pearl River Scholar·Tianshan Talent” Cooperative Expert Studio Innovation Team Program of the First People’s Hospital of Kashgar Region, No. KDYY202111 (to WFZ)
CLC Number:
Cite this article
Yang Huaqun, Abudouainijiang·Abulimiti, Wang Fazheng, Maimaitishawutiaji·Maimaiti, Li Simi, Muhetaer·Maimaitirexiati. Weighted gene co-expression network analysis combined with machine learning identifies autophagy and senescence signature genes in osteoarthritis chondrocytes[J]. Chinese Journal of Tissue Engineering Research, 2026, 30(34): 8889-8898.
share this article
Add to citation manager EndNote|Reference Manager|ProCite|BibTeX|RefWorks
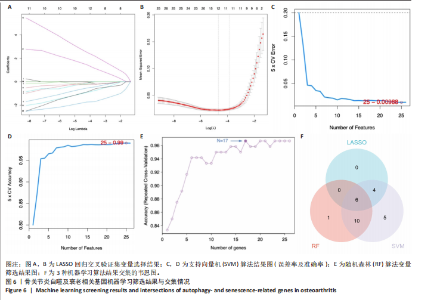
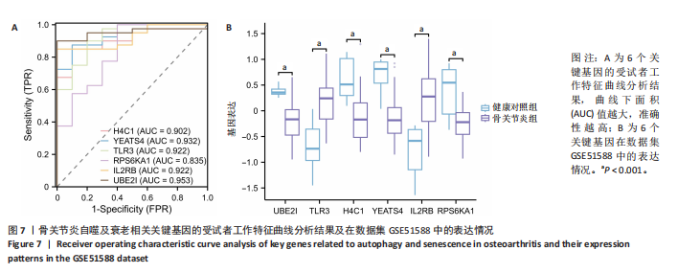
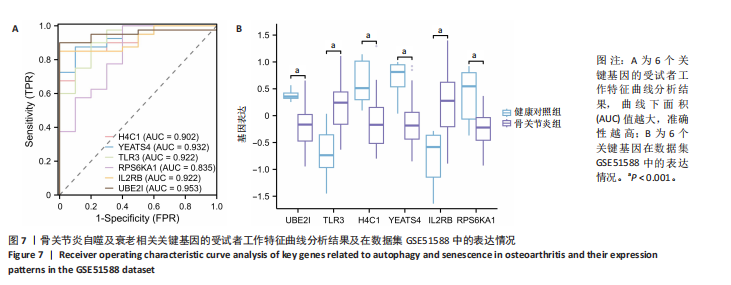
2.4 自噬及衰老相关差异表达基因的功能富集分析结果 基于clusterProfiler包对自噬及衰老相关差异表达基因进行GO富集分析,以探讨其生物学功能。GO富集分析显示,生物过程中,自噬及衰老相关差异表达基因显著富集于与细胞数量稳态和免疫细胞分化相关的生物学过程,包括细胞数量稳态、髓系细胞分化、红细胞分化;细胞分析中,自噬及衰老相关差异表达基因显著富集于与染色体功能和蛋白-DNA复合物相关的细胞组件,包括核染色体、蛋白质-DNA复合物、富含Ficolin-1颗粒腔;分子功能中,自噬及衰老相关差异表达基因显著富集于与生长因子结合和细胞因子受体活性相关的分子功能,包括生长因子结合、细胞因子结合及受体活性。KEGG通路富集分析显示,这些差异基因主要富集于坏死性凋亡、造血细胞谱系有关。采用柱状图来展示自噬及衰老相关差异表达基因GO富集分析及KEGG通路富集情况,见图5。 2.5 机器学习筛选与受试者工作特征曲线分析结果 通过LASSO、随机森林和支持向量机机器学习算法进一步筛选26个骨关节炎自噬及衰老相关基因基因。LASSO算法筛选出10个基因(图6A,B)。支持向量机算法筛选出25个基因,模型误差率为0.009(图6C),准确率为0.99(图6D)。随机森林算法筛选出17个基因(图6E)。韦恩图显示三者交集得到6个基因,分别为:UBE21、TLR3、H4C1、YEATS4、IL2RB、RPS6KA1(图6F)。使用pROC包绘制受试者工作特征曲线并计算AUC值,验证诊断性能,结果"
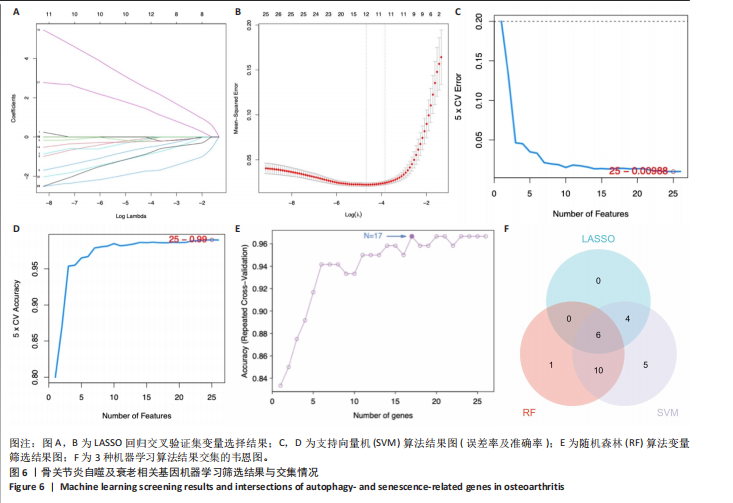
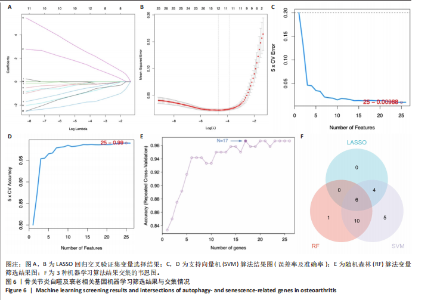
显示6个关键基因的AUC值均大于0.8,分别为UBE21(0.953)、TLR3(0.922)、H4C1(0.902)、YEATS4(0.932)、IL2RB(0.922)、RPS6KA1(0.835),见图7A。这些关键基因在数据集GSE51588中的表达情况见图7B。 2.6 关键基因的外部数据集验证结果 将GSE114007数据集作为外部验证集分析6个自噬及衰老相关关键基因的表达情况,结果显示骨关节炎组UBE2I、TLR3、IL2RB表达高于健康对照组(P < 0.05,P < 0.01),两组间H4C1、YEATS4、RPS6KA1表达比较差异无显著性意义(P > 0.05),见图8。"
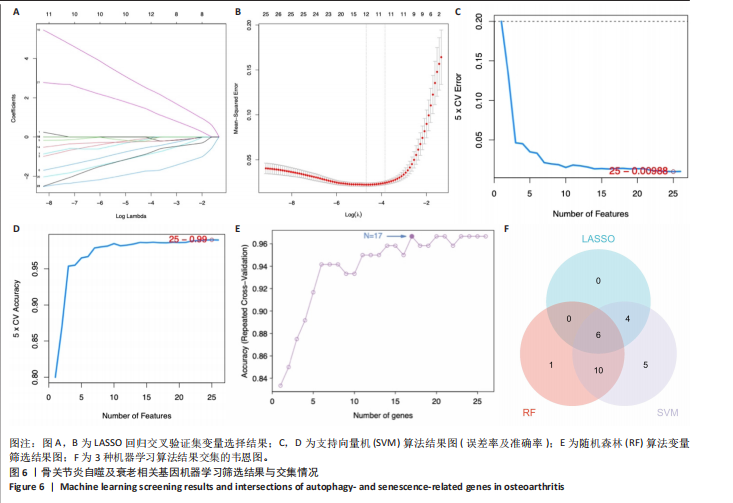
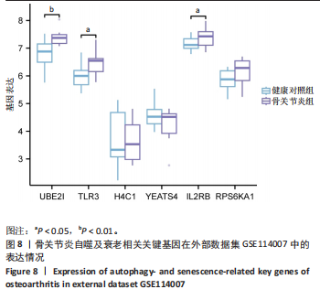
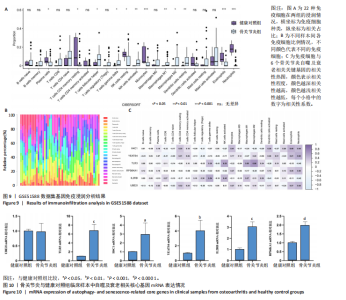
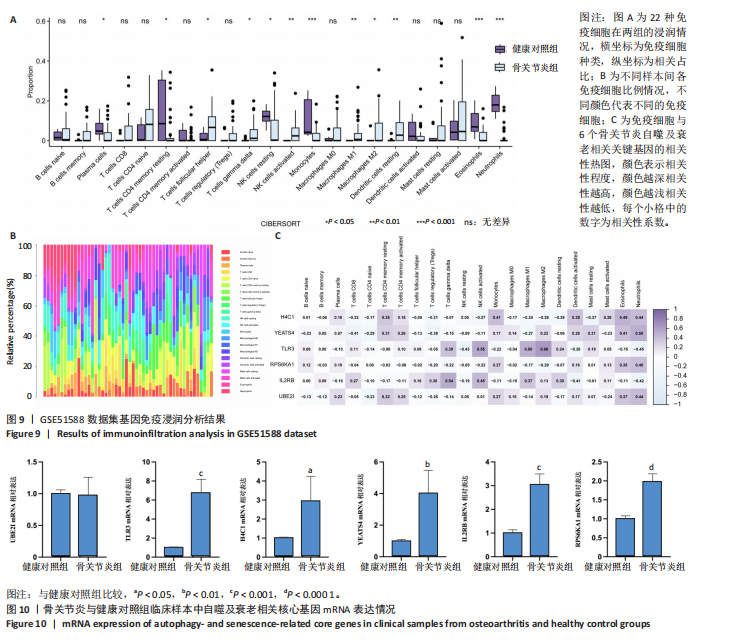
2.7 免疫浸润分析结果 基于CIBERSORT算法进行免疫浸润分析骨关节炎组与健康对照组在22种免疫细胞浸润方面的差异,结果显示多个免疫细胞类型的比例发生了显著变化。箱线图(图9A)表明骨关节炎组软骨组织中浆细胞、静息CD4记忆T细胞、静息NK细胞、单核细胞、M2型巨噬细胞、嗜酸性粒细胞和中性粒细胞的浸润明显降低,滤泡辅助T细胞、γδT细胞、激活的NK细胞、M1型巨噬细胞和静止树突状细胞浸润显著增加,提示多种免疫细胞可能参与调控关节炎的病理过程。利用柱状图表示不同样本间各免疫细胞的占比,结果显示不同样本间免疫细胞组成不同(图9B)。6个关键基因与免疫细胞的相关性分析(图9C)显示,中性粒细胞与多个基因呈正相关,因此推测其可能在关节炎自噬及衰老相关机制中发挥重要作用。 2.8 临床样本关键基因验证结果 RT-qPCR检测显示,与健康对照组相比,骨关节炎组TLR3、YEATS4、H4C1、IL2RB和RPS6KA1 mRNA表达升高(P < 0.05,P < 0.01,P < 0.001),两组间UBE2I mRNA表达比较差异无显著性意义(P > 0.05),见图10。"
| [1] XU T, LIU K, FAN J, et al. Metformin mitigates osteoarthritis progression by modulating the PI3K/AKT/mTOR signaling pathway and enhancing chondrocyte autophagy. Open Life Sci. 2024;19(1):20220922. [2] KONG P, AHMAD RE, ZULKIFLI A, et al. The role of autophagy in mitigating osteoarthritis progression via regulation of chondrocyte apoptosis: A review. Joint Bone Spine. 2024;91(3):105642. [3] LI J, YU G, HE Z, et al. Kojic acid alleviates osteoarthritis by attenuating inflammation and restoring impaired autophagy through regulating NF-κB and PI3K/AKT/mTOR signaling pathways: An in vivo and in vitro study. Int Immunopharmacol. 2025;147:114022. [4] LU R, HE Z, ZHANG W, et al. Oroxin B alleviates osteoarthritis through anti-inflammation and inhibition of PI3K/AKT/mTOR signaling pathway and enhancement of autophagy. Front Endocrinol (Lausanne). 2022;13:1060721. [5] 薛庆云,王坤正,裴福兴,等.中国40岁以上人群原发性骨关节炎患病状况调查[J].中华骨科杂志,2015,35(12):1206-1212. [6] CHEN Y, PAN X, ZHAO J, et al. Icariin alleviates osteoarthritis through PI3K/Akt/mTOR/ULK1 signaling pathway. Eur J Med Res. 2022;27(1):204. [7] LIU Y, ZHANG Z, LI T, et al. Senescence in osteoarthritis: from mechanism to potential treatment. Arthritis Res Ther. 2022;24(1):174. [8] JEON OH, KIM C, LABERGE RM, et al. Local clearance of senescent cells attenuates the development of post-traumatic osteoarthritis and creates a pro-regenerative environment. Nat Med. 2017;23(6):775-781. [9] SHAO Y, ZHANG H, GUAN H, et al. PDZK1 protects against mechanical overload-induced chondrocyte senescence and osteoarthritis by targeting mitochondrial function. Bone Res. 2024;12(1):41. [10] CHEN Z, TANG M, WU Z, et al. Increased Rab1a accelerates osteoarthritis by inhibiting autophagy via activation of the mTORC1-S6K pathway. J Adv Res. 2025;75:607-619. [11] LIU Z, WANG T, SUN X, et al. Autophagy and apoptosis: regulatory factors of chondrocyte phenotype transition in osteoarthritis. Hum Cell. 2023;36(4):1326-1235. [12] PAN Y, YANG Y, FAN M, et al. Progranulin regulation of autophagy contributes to its chondroprotective effect in osteoarthritis. Genes Dis. 2023;10(4):1582-1595. [13] 李文铭,李永航,严才平,等.增强自噬的可注射水凝胶微球改善软骨微环境抗软骨细胞衰老[J]. 中国组织工程研究,2025,29(22):4710-4719. [14] FENG K, YE T, XIE X, et al. ESC-sEVs alleviate non-early-stage osteoarthritis progression by rejuvenating senescent chondrocytes via FOXO1A-autophagy axis but not inducing apoptosis. Pharmacol Res. 2024;209:107474. [15] BINVIGNAT M, PEDOIA V, BUTTE A J, et al. Use of machine learning in osteoarthritis research: a systematic literature review. RMD Open. 2022;8(1): e001998. [16] XU W, SUN Y, ZHAO S, et al. Identification and validation of autophagy-related genes in primary open-angle glaucoma. BMC Med Genomics. 2023;16(1):287. [17] LI ZH, LI SP, LI YH, et al. Identification of aging-related biomarkers for intervertebral disc degeneration in whole blood samples based on bioinformatics and machine learning. Front Immunol. 2025;16:1565945. [18] AVELAR RA, ORTEGA JG, TACUTU R, et al. A multidimensional systems biology analysis of cellular senescence in aging and disease. Genome Biol. 2020;21(1):91. [19] SUN M, YANG Q, HU C, et al. Identification and Validation of Autophagy-Related Genes in Sepsis-Induced Acute Respiratory Distress Syndrome and Immune Infiltration. J Inflamm Res. 2022;15:2199-2212. [20] DING JT, HONG FF, YANG SL. Roles of autophagy in rheumatoid arthritis. Clin Exp Rheumatol. 2022;40(11):2179-2187. [21] CHENG L, CHANG S, TAN Y, et al. Platelet-rich plasma combined with isometric quadriceps contraction regulates autophagy in chondrocytes via the PI3K/AKT/mTOR pathway to promote cartilage repair in knee osteoarthritis. Regen Ther. 2025;28:81-89. [22] TANG L, DING J, YANG K, et al. New insights into the mechanisms and therapeutic strategies of chondrocyte autophagy in osteoarthritis. J Mol Med (Berl). 2024; 102(10):1229-1244. [23] LEE DY, BAHAR ME, KIM CW, et al. Autophagy in Osteoarthritis: A Double-Edged Sword in Cartilage Aging and Mechanical Stress Response: A Systematic Review. J Clin Med. 2024;13(10):3005. [24] MUSUMECI G, CASTROGIOVANNI P, TROVATO FM, et al. Biomarkers of Chondrocyte Apoptosis and Autophagy in Osteoarthritis. Int J Mol Sci. 2015; 16(9):20560-20575. [25] ANDREI C, MIHAI DP, NITULESCU GM, et al. Modulating Autophagy in Osteoarthritis: Exploring Emerging Therapeutic Drug Targets. Int J Mol Sci. 2024; 25(24):13695. [26] LV X, ZHAO T, DAI Y, et al. New insights into the interplay between autophagy and cartilage degeneration in osteoarthritis. Front Cell Dev Biol. 2022;10:1089668. [27] WU J, HUANG S, YU Y, et al. Human adipose and synovial-derived MSCs synergistically attenuate osteoarthritis by promoting chondrocyte autophagy through FoxO1 signaling. Stem Cell Res Ther. 2024;15(1):261. [28] LORENZO-GÓMEZ I, NOGUEIRA-RECALDE U, GARCÍA-DOMÍNGUEZ C, et al. Defective chaperone-mediated autophagy is a hallmark of joint disease in patients with knee osteoarthritis. Osteoarthritis Cartilage. 2023;31(7):919-933. [29] KHALEQUE MA, KIM JH, TANVIR MAH, et al. Significance of Necroptosis in Cartilage Degeneration. Biomolecules. 2024;14(9):1192. [30] WANG H, ZHOU J, LU Y, et al. The interplay between autophagy and programmed cell death in osteoarthritis: insights into mechanisms and therapeutic targets. Mol Cell Biochem. 2025;480(8):4627-4646. [31] DIEKMAN BO, LOESER RF. Aging and the emerging role of cellular senescence in osteoarthritis. Osteoarthritis Cartilage. 2024;32(4):365-371. [32] TAMATTA R, PAI V, JAISWAL C, et al. Neuroinflammaging and the Immune Landscape: The Role of Autophagy and Senescence in Aging Brain. Biogerontology. 2025;26(2):52. [33] XU F, LI Z, LIU T, et al. The role of cellular senescence in the pathogenesis of Rheumatoid Arthritis: Focus on IL-6 as a target gene. Cytokine. 2024;184:156762. [34] DALMAO-FERNÁNDEZ A, HERMIDA-GÓMEZ T, NOGUEIRA-RECALDE U, et al. Mitochondrial Role on Cellular Apoptosis, Autophagy, and Senescence during Osteoarthritis Pathogenesis. Cells. 2024;13(11):976. [35] SILVA R. The dichotomic role of cytokines in aging. Biogerontology. 2024;26(1):17. [36] KANDHAYA-PILLAI R, YANG X, TCHKONIA T, et al. TNF-α/IFN-γ synergy amplifies senescence-associated inflammation and SARS-CoV-2 receptor expression via hyper-activated JAK/STAT1. Aging Cell. 2022;21(6):e13646. [37] STOLBERG-STOLBERG J, BOETTCHER A, SAMBALE M, et al. Toll-like receptor 3 activation promotes joint degeneration in osteoarthritis. Cell Death Dis. 2022; 13(3):224. [38] QIAN Y, CHU G, ZHANG L, et al. M2 macrophage-derived exosomal miR-26b-5p regulates macrophage polarization and chondrocyte hypertrophy by targeting TLR3 and COL10A1 to alleviate osteoarthritis. J Nanobiotechnology. 2024;22(1):72. [39] WAN Z, WANG X, FU Z, et al. Toll-like receptor activation regulates the paracrine effect of adipose-derived mesenchymal stem cells on reversing osteoarthritic phenotype of chondrocytes. Mol Biol Rep. 2024;51(1):550. [40] ZHOU D, LUO Y, LI F, et al. Exploring the mechanisms of PANoptosis in osteoarthritis and the therapeutic potential of andrographolide through bioinformatics and single-cell analysis. Biol Direct. 2025;20(1):41. [41] HOU Z, WANG M, CAO S. Inhibition of Toll-Like Receptor 3 Relieves Osteoarthritis by Suppression of Cartilage Degradation, Nuclear Factor Kappa B-Mediated Inflammation, and Activation of Autophagy. Cartilage. 2025: 19476035251317713. doi: 10.1177/19476035251317713. [42] YANG Y, YUAN S, CHE M, et al. Genetic analysis of the relation between IL2RA/IL2RB and rheumatoid arthritis risk. Mol Genet Genomic Med. 2019;7(7):e00754. [43] HU M, DENG Y, BAI Y, et al. Identifying Key Biomarkers Related to Immune Response in the Progression of Diabetic Kidney Disease: Mendelian Randomization Combined With Comprehensive Transcriptomics and Single-Cell Sequencing Analysis. J Inflamm Res. 2025;18:949-972. [44] LIU M, WU C, WU C, et al. Immune cells differentiation in osteoarthritic cartilage damage: friends or foes? Front Immunol. 2025;16:1545284. [45] RUYSSEN-WITRAND A, LUKAS C, NIGON D, et al. Association of IL-2RA and IL-2RB genes with erosive status in early rheumatoid arthritis patients (ESPOIR and RMP cohorts). Joint Bone Spine. 2014;81(3):228-234. [46] ARNOLD CR, WOLF J, BRUNNER S, et al. Gain and loss of T cell subsets in old age--age-related reshaping of the T cell repertoire. J Clin Immunol. 2011;31(2):137-146. [47] PUA HH, GUO J, KOMATSU M, et al. Autophagy is essential for mitochondrial clearance in mature T lymphocytes. J Immunol. 2009;182(7):4046-4055. [48] NIKOLICH-ŽUGICH J. The twilight of immunity: emerging concepts in aging of the immune system. Nat Immunol. 2018;19(1):10-19. |
| [1] | Zhang Haiwen, Zhang Xian, Xu Taichuan, Li Chao. Bibliometric and visual analysis of the research status and trends of senescence in osteoporosis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(6): 1580-1591. |
| [2] | Lyu Guoqing, Aizimaitijiang·Rouzi, Xiong Daohai. Irisin inhibits ferroptosis in human articular chondrocytes: roles and mechanisms [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(6): 1359-1367. |
| [3] | Jia Jinwen, Airefate·Ainiwaer, Zhang Juan. Effects of EP300 on autophagy and apoptosis related to allergic rhinitis in rats [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(6): 1439-1449. |
| [4] | Lai Jiaming, , Song Yuling, Chen Zixi, Wei Jinghuan, Cai Hao, , Li Guoquan, . Screening of diagnostic markers for endothelial cell Senescence in mice with radiation-induced heart disease and analysis of immune infiltration [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(6): 1450-1463. |
| [5] | You Huijuan, Wu Shuzhen, Rong Rong, Chen Liyuan, Zhao Yuqing, Wang Qinglu, Ou Xiaowei, Yang Fengying. Macrophage autophagy in lung diseases: two-sided effects [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(6): 1516-1526. |
| [6] | Liu Kexin, , Hao Kaimin, Zhuang Wenyue, , Li Zhengyi. Autophagy-related gene expression in pulmonary fibrosis models: bioinformatic analysis and experimental validation [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(5): 1129-1138. |
| [7] | Hu Jing, Zhu Ling, Xie Juan, Kong Deying, Liu Doudou. Autophagy regulates early embryonic development in mice via affecting H3K4me3 modification [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(5): 1147-1155. |
| [8] | Bu Yangyang, Ning Xinli, Zhao Chen. Intra-articular injections for the treatment of osteoarthritis of the temporomandibular joint: different drugs with multiple combined treatment options [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(5): 1215-1224. |
| [9] | Zhang Qian, Huang Dongfeng. Weighted gene co-expression network analysis combined with machine learning to screen and validate biomarkers for osteoarthritis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(5): 1096-1105. |
| [10] | Gu Fucheng, Yang Meixin, Wu Weixin, Cai Weijun, Qin Yangyi, Sun Mingyi, Sun Jian, Geng Qiudong, Li Nan. Effects of Guilu Erxian Glue on gut microbiota in rats with knee osteoarthritis: machine learning and 16S rDNA analysis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(4): 1058-1072. |
| [11] | Ma Runqiu, Yang Huixia, Li Xuer, Bai Zhigang, Li Guizhong, Hao Yinju, Ma Shengchao, Jiang Yideng. Mechanism of glucocorticoid-induced mitochondrial dysfunction in osteoblasts in steroid-induced osteonecrosis of the femoral head [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(34): 8845-8851. |
| [12] | Yuan Jingjing, Zhang Xiaomin, Du Pengyang, Wang Weifeng. Mazdutide improves cognitive function in APP/PS1/Tau triple transgenic mice [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(34): 8962-8969. |
| [13] | Guan Yujie, Zhao Bin. Application and prospect of artificial intelligence in screening and diagnosis of scoliosis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(3): 721-730. |
| [14] | Wang Zhipeng, Zhang Xiaogang, Zhang Hongwei, Zhao Xiyun, Li Yuanzhen, Guo Chenglong, Qin Daping, Ren Zhen. A systematic review of application value of machine learning to prognostic prediction models for patients with lumbar disc herniation [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(3): 740-748. |
| [15] | Zhao Feifan, Cao Yujing. An artificial neural network model of ankylosing spondylitis and psoriasis shared genes and machine learning-based mining and validation [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(3): 770-784. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||