Chinese Journal of Tissue Engineering Research ›› 2026, Vol. 30 ›› Issue (24): 6421-6432.doi: 10.12307/2026.242
m6A-related ferroptosis gene expression and its association with immune infiltration in Alzheimer’s disease: machine learning and molecular biology validation
Xu Dongfang1, Zhao Kun2, Lu Changzhu2, Wang Yuge2, Bai Lianjie3, Meng Fanmou2, Wang Yang2, 4, Yao Hongbo5
- 1Qiqihar Medical University, Qiqihar 161000, Heilongjiang Province, China; 2Department of Physiology, 5Department of Histology and Embryology, School of Basic Medicine, Qiqihar Medical University, Qiqihar 161000, Heilongjiang Province, China; 3Department of Ultrasound, the Second Affiliated Hospital of Qiqihar Medical University, Qiqihar 161000, Heilongjiang Province, China; 4Heilongjiang Provincial Key Laboratory of Food & Medicine Homology and Metabolic Disease Prevention, Qiqihar 161000, Heilongjiang Province, China
-
Received:2025-10-15Revised:2025-11-14Online:2026-08-28Published:2026-02-05 -
Contact:Wang Yang, PhD, Master's supervisor, Department of Physiology, School of Basic Medicine, Qiqihar Medical University, Qiqihar 161000, Heilongjiang Province, China; Heilongjiang Provincial Key Laboratory of Food & Medicine Homology and Metabolic Disease Prevention, Qiqihar 161000, Heilongjiang Province, China Co-corresponding author: Yao Hongbo, PhD, Master's supervisor, Department of Histology and Embryology, School of Basic Medicine, Qiqihar Medical University, Qiqihar 161000, Heilongjiang Province, China -
About author:Xu Dongfang, Qiqihar Medical University, Qiqihar 161000, Heilongjiang Province, China -
Supported by:Heilongjiang Natural Science Foundation of China, No. LH2021H122 (to YHB); Heilongjiang Postdoctoral Funding Project, No. LBH-Z23294 (to WY); Qiqihar Science and Technology Plan Joint Guidance Project, No. LSFGG-2023035 (to WY); Qiqihar Academy of Medical Sciences Project, No. QMSI2021M-11 (to WY)
CLC Number:
Cite this article
Xu Dongfang, Zhao Kun, Lu Changzhu, Wang Yuge, Bai Lianjie, Meng Fanmou, Wang Yang, , Yao Hongbo. m6A-related ferroptosis gene expression and its association with immune infiltration in Alzheimer’s disease: machine learning and molecular biology validation[J]. Chinese Journal of Tissue Engineering Research, 2026, 30(24): 6421-6432.
share this article
Add to citation manager EndNote|Reference Manager|ProCite|BibTeX|RefWorks
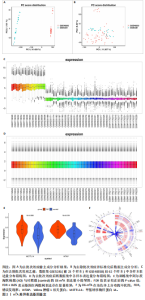
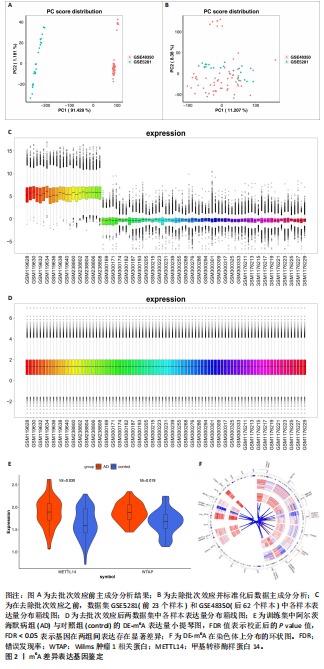
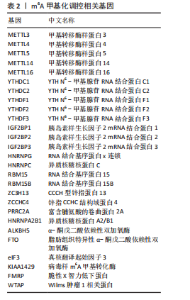
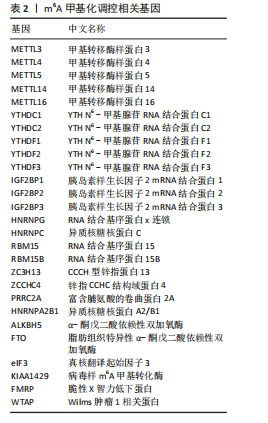
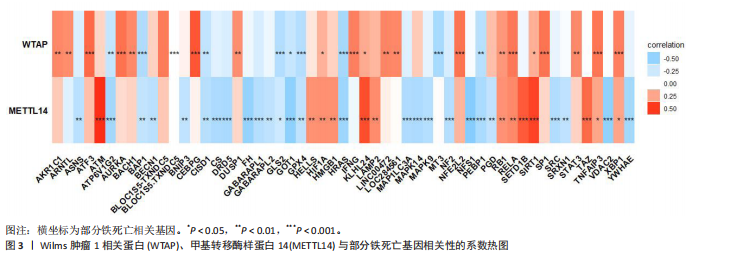
2.1 m6A差异表达基因 数据集GSE5281和GSE48350合并后训练集(56例正常样本vs. 29例阿尔茨海默病样本),经数据预处理发现原始数据存在显著批次偏倚(图2A,C),进行跨平台批次校正与标准化处理后,主成分分析显示样本均匀分布(图2B,D),验证批次效应有效消除。m6A甲基化调控相关基因见表2。在27个m6A调控基因中[剔除真核翻译起始因子3、脆性X智力低下蛋白、甲基转移酶样蛋白16( methyltransferase-like protein 16,METTL16)、RNA结合基序蛋白x连锁4个未检测到的基因],鉴定出2个显著差异表达基因,分别为Wilms肿瘤1相关蛋白(Wilms Tumor 1-Associated Protein,WTAP)、甲基转移酶样蛋白14( methyltransferase-like protein 14,METTL14)基因,分别位于6号、4号染色体上(图2E,F)。 2.2 m6A相关铁死亡基因 基于259个铁死亡核心基因,在训练集中筛选出240个相关基因。选取铁死亡相关基因与WTAP、METTL14相关性较高的部分基因以热图展示(图3)。获得16"
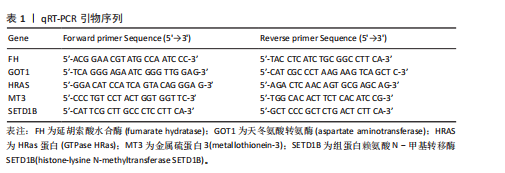
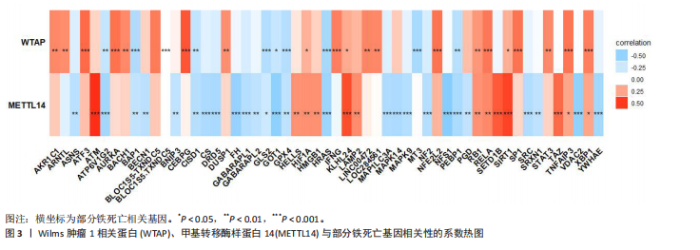
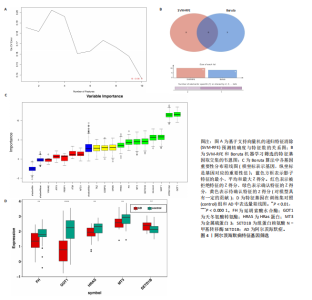
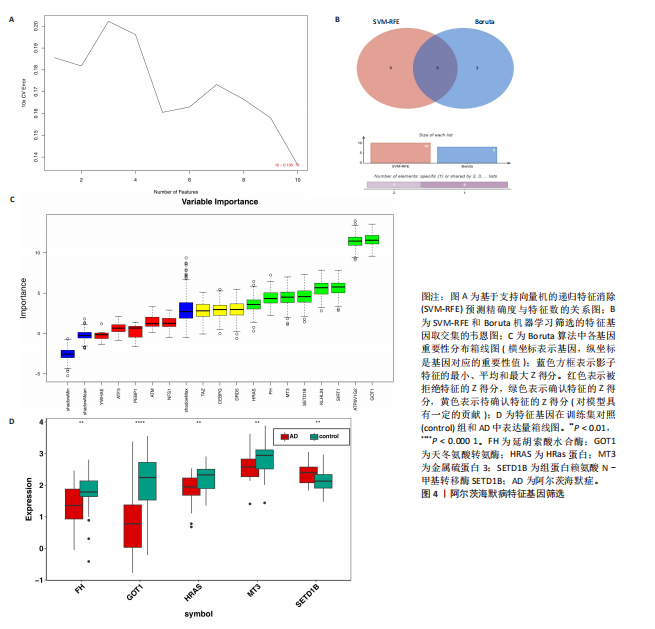
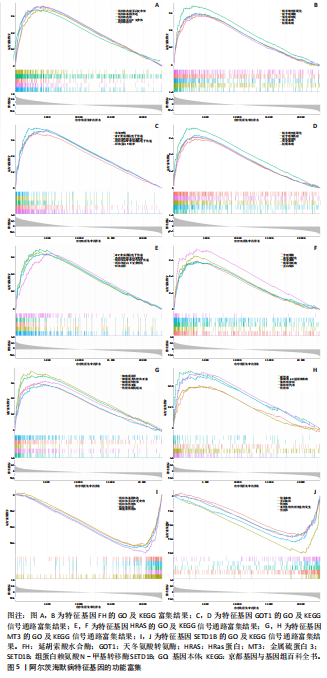
个与WTAP、METTL14高度相关的铁死亡基因,分别为半胱氨酸脱硫酶(cysteine desulfurase,NFS1)、GOT1、FH、HRAS、YWHAE、 ATP6V1G2、DRD5、磷脂酰乙醇胺结合蛋白(phosphatidylethanolamine-binding protein,PEBP1)、MT3、CCAAT增强子结合蛋白γ (CCAAT/enhancer-binding protein gamma,CEBPG)、ATF3、Tafazzin (TAZ)、SETD1B、KLHL24、SIRT1、丝氨酸蛋白激酶ATM(Serine-protein kinase ATM,ATM)。 2.3 阿尔茨海默病特征基因 基于支持向量机的递归特征消除方法筛选10个阿尔茨海默病特征基因:GOT1、MT3、NFS1、SETD1B、CEBPG、PEBP1、FH、HRAS、TAZ、ATM。Boruta方法筛选得到8个阿尔茨海默病特征基因:GOT1、FH、HRAS、ATP6V1G2、MT3、SETD1B、KLHL24、SIRT1。两种机器学习法筛选共有特征基因5个,分别为FH、GOT1、HRAS、MT3、SETD1B基因。5个特征基因的mRNA表达量在训练集阿尔茨海默病样本与正常样本组间均存在显著性差异(P < 0.05或P < 0.01),见图4。 2.4 特征基因功能富集 基因集富集分析按照显著性排序,选取基因本体和京都基因与基因组百科全书中前5项进行可视化展示。京都基因与基因组百科全书信号通路主要富集在氧化磷酸化信号通路、亨廷顿病、帕金森病、脂肪酸降解及代谢通路、蛋白酶体信号通路等。基因本体功能定位于细胞中线粒体内膜、胞质核糖体结构成分等,参与的生物过程主要富集在线粒体ATP合成耦合电子传递过程、线粒体内膜蛋白复合物形成、线粒体基因表达等(图5)。"
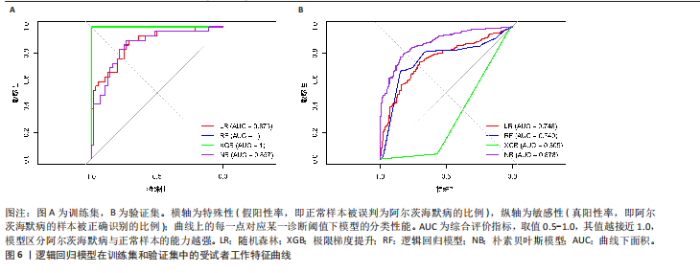
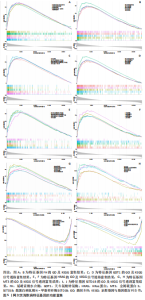
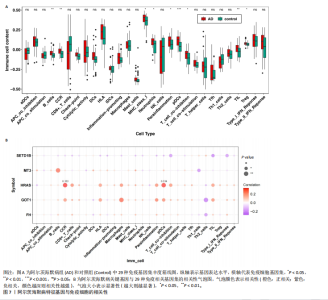
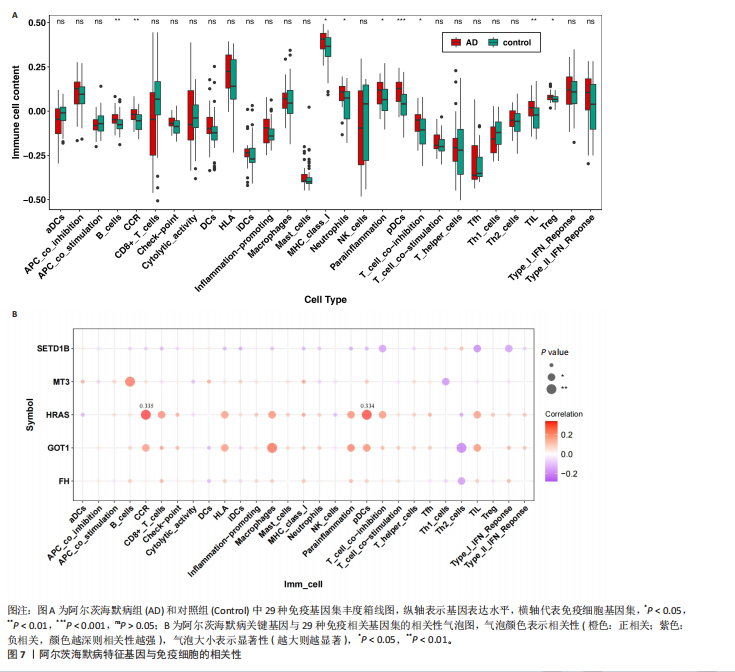
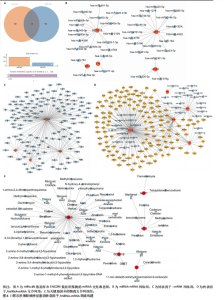
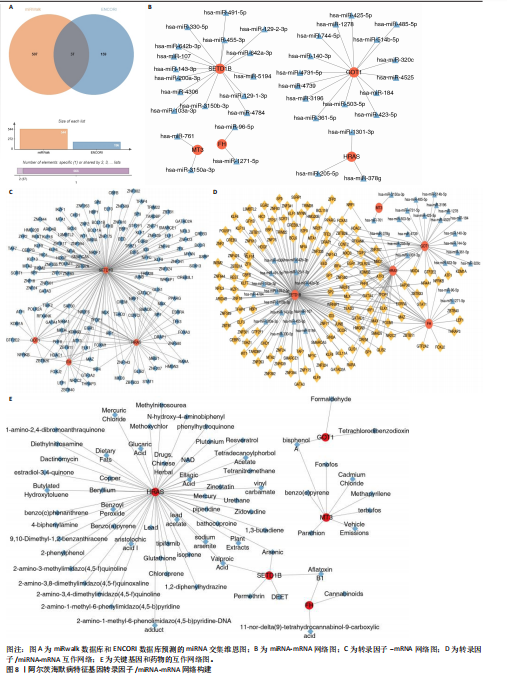
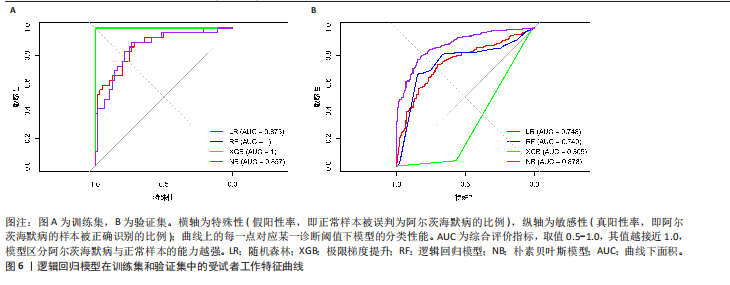
2.5 特征基因诊断性能 在训练集中,随机森林分析和极限梯度提升模型的曲线下面积均为1.0,出现过拟合现象;逻辑回归模型模型曲线下面积为0.873,朴素贝叶斯模型为0.857,二者性能稳定且未出现过拟合。在验证集中,随机森林模型曲线下面积降至0.749,极限梯度提升模型骤降至0.309,逻辑回归模型曲线下面积为0.748,朴素贝叶斯模型为0.878。综合来看,逻辑回归模型在训练集与验证集的性能衰减程度最小,泛化能力显著优于随机森林和极限梯度提升;而朴素贝叶斯模型虽验证集曲线下面积较高,但在训练集的拟合程度弱于逻辑回归模型。因此,选取逻辑回归模型作为最优模型,结果表明基于这5个关键基因构建的逻辑回归模型在训练与验证中均具有较高的分类性能,是阿尔茨海默病发生发展中的关键基因(图6)。 2.6 特征基因与免疫细胞相关性 通过单样本基因集富集分析,发现训练集中具有明显差异表达量的9个免疫相关基因集分别为B细胞(B_cells)、趋化因子受体(CC chemokine receptor, CCR)、主要组织相容性复合体Ⅰ类(Major histocompatibility complex class I-related protein 1,MHC_class_I)、 中性粒细胞(Neutrophils)、副炎症(Parainflammation)、浆细胞样树突状细胞(Plasmacytoid dendritic cells, pDCs)、T细胞共抑制物(T_cell_co-inhibition)、肿瘤浸润淋巴细胞(tumor-infiltrating lymphocyte,TIL)、调节性T细胞(Regulatory T cells,Tregs)。在阿尔茨海默病关键基因与29种免疫相关基因集的相关性分析中,关键基因HRAS与趋化因子受体基因集和浆细胞样树突状细胞基因集均表现出较高的相关性,相关系数分别为0.335和0.334(图7)。 2.7 特征基因转录因子/miRNA-mRNA网络构建 基于5个关键基因,分别从miRwalk3.0和ENCORI数据库筛选出544个和196个相关miRNA,取交集获得37个共同miRNA,其中2个miRNA对应FH基因,15个miRNA对应GOT1,3个miRNA对应HRAS,2个miRNA对应MT3,15个miRNA对应SETD1B。通过NetworkAnalys数据库获取5个关键基因的142个转录因子,构建200对转录因子-mRNA互作网络。利用Cytoscape软件对5个mRNA、37个miRNA及142个转录因子进行联合可视化,构建转录因子/ miRNA -mRNA互作网络。基于CTD数据库中预测关键基因治疗阿尔茨海默病的潜在药物,共获得71种靶向治疗药物(图8)。 2.8 特征基因验证 为验证特征基因在阿尔茨海默病中的特异性调控作用,此次研究通过qRT-PCR以及Western Blotting实验检测APP/PS1转基因阿尔茨海默病模型小鼠海马组织关键基因转录及蛋白表达水平。结果显示,相对于正常野生型小鼠,阿尔茨海默病模型小鼠GOT1、HRAS 的mRNA以及蛋白表达量均降低(P < 0.05),而SETD1B的mRNA以及蛋白表达量则显著上调(P < 0.05),而FH及MT3表达水平在转录以及蛋白表达方面均无明显改变,无统计学差异(图9)。"
| [1] LIU Y, TAN Y, ZHANG Z, et al. The interaction between ageing and Alzheimer’s disease: insights from the hallmarks of ageing. Transl Neurodegener. 2024;13(1):7. [2] 王刚,齐金蕾,刘馨雅,等.中国阿尔茨海默病报告2024[J].诊断学理论与实践,2024, 23(3):219-256. [3] JIA J, NING Y, CHEN M, et al. Biomarker changes during 20 years preceding Alzheimer’s disease. N Engl J Med. 2024;390(8):712-722. [4] GRAFF-RADFORD J, YONG KXX, APOSTOLOVA LG, et al. New insights into atypical Alzheimer’s disease in the era of biomarkers. Lancet Neurol. 2021;20(3):222-234. [5] LIU E, ZHANG Y, WANG J. Updates in Alzheimer’s disease: from basic research to diagnosis and therapies. Transl Neurodegener. 2024;13(1):45. [6] XU Z, LV B, QIN Y, et al. Emerging roles and mechanism of m6A methylation in cardiometabolic diseases. Cells. 2022;11(7): 1101. [7] WEI G. RNA m6A modification, signals for degradation or stabilisation? Biochem Soc Trans. 2024;52(2):707-717. [8] HAN Y, SUN J, YAO M, et al. Biological roles of enhancer RNA m6A modification and its implications in cancer. Cell Commun Signal. 2025;23(1):254. [9] SHAFIK AM, ZHANG F, GUO Z, et al. N6-methyladenosine dynamics in neurodevelopment and aging, and its potential role in Alzheimer’s disease. Genome Biol. 2021;22(1):17. [10] HAN M, LIU Z, XU Y, et al. Abnormality of m6A mRNA methylation is involved in Alzheimer’s disease. Front Neurosci. 2020;14:98. [11] ZHANG X, YANG S, HAN S, et al. Differential methylation of circRNA m6A in an APP/PS1 Alzheimer’s disease mouse model. Mol Med Rep. 2023;27(2):55. [12] HU B, SHI Y, XIONG F, et al. Rewired m6A of promoter antisense RNAs in Alzheimer’s disease regulates neuronal genes in 3D nucleome. Nat Commun. 2025;16(1):5251. [13] RU Q, LI Y, CHEN L, et al. Iron homeostasis and ferroptosis in human diseases: mechanisms and therapeutic prospects. Signal Transduct Target Ther. 2024;9(1):271. [14] DIXON SJ, OLZMANN JA. The cell biology of ferroptosis. Nat Rev Mol Cell Biol. 2024;25: 424-442. [15] WANG L, FANG X, LING B, et al. Research progress on ferroptosis in the pathogenesis and treatment of neurodegenerative diseases. Front Cell Neurosci. 2024;18: 1359453. [16] ABDUKARIMOV N, KOKABI K, KUNZ J. Ferroptosis and Iron Homeostasis: Molecular Mechanisms and Neurodegenerative Disease Implications. Antioxidants (Basel). 2025; 14(5):527. [17] LI X, CHEN J, FENG W, et al. Berberine ameliorates iron levels and ferroptosis in the brain of 3 x Tg-AD mice. Phytomedicine. 2023;118:154962. [18] SHEN M, LI Y, WANG Y, et al. N6-methyladenosine modification regulates ferroptosis through autophagy signaling pathway in hepatic stellate cells. Redox Biol. 2021;47:102151. [19] LIU Z, LI H, PAN S. Discovery and validation of key biomarkers based on immune infiltrates in Alzheimer’s Disease. Front Genet. 2021;12: 658323. [20] TAO X, KANG N, ZHENG Z, et al. The regulatory mechanisms of N6-methyladenosine modification in ferroptosis and its implications in disease pathogenesis. Life Sci. 2024;355: 123011. [21] JIANG X, YAN Q, XIE L, et al. Construction and validation of a Ferroptosis-related prognostic model for gastric cancer. J Oncol. 2021;2021:6635526. [22] HE Y, JIANG Z, CHEN C, et al. Classification of triple-negative breast cancers based on Immunogenomic profiling. J Exp Clin Cancer Res. 2018;37(1):327. [23] 张振涛. 散发性早发型阿尔茨海默病:独特特征、关键机制与未来展望[J]. 科学通报, 2025,70(1):6-7. [24] ZHANG R, ZHANG Y, GUO F, et al. RNA N6 -methyl‐ adenosine modifications and its roles in Alzheimer’s disease. Front Cell Neurosci. 2022;16:820378. [25] BARBIRI I, KOUZARIDES T. Role of RNA modifications in cancer. Nat Rev Cancer. 2020; 20(6):303-322. [26] HAN X, GUO J, FAN Z. Interactions between m6A modifi‐ cation and miRNAs in malignant tumors. Cell Death Dis.2021;12(6):598. [27] SENDINC E, SHI Y. RNA m6A methylation across the transcriptome. Mol Cell. 2023;83(3):428-441. [28] MI S, SHI Y, DARI G, et al. Function of m6A and its regu lation of domesticated animals’ complex traits. J Anim Sci. 2022; 100(3):skac034. [29] SANG A, ZHANG J, ZHANG M, et al. METTL4 mediated-N6 methyladenosine promotes acute lung injury by activating ferroptosis in alveolar epithelial cells. Free Rad Biol Med. 2024;213: 90-101. [30] ZHU Z, HUO F, ZHANG J, et al. Crosstalk between m6A modification and alternative splicing during cancer progression. Clin Transl Med. 2023;13(10):e1460. [31] CHEN Y, PENG C, CHEN J, et al. WTAP facilitates pro gression of hepatocellular carcinoma via m6A-HuR-depen dent epigenetic silencing of ETS1. Mol Cancer. 2019;18(1):127. [32] HUANG H, CAMATS-PERNA J, MEDEIROS R, et al. Altered expression of the m6A methyltransferase METTL3 in Alzheimer’s disease. eNeuro. 2020;7(5): ENEURO.0125-20.2020. [33] ZHAO F, XU Y, GAO S, et al. METTL3-dependent RNA m(6)A dysregulation contributes to neurodegeneration in Alzheimer’s disease through aberrant cell cycle events. Mol Neurodegener. 2021;16(1):70. [34] JIANG X, STOCKWELL BR, CONRAD M. Ferroptosis: mechanisms, biology and role in disease. Nat Rev Mol Cell Biol. 2021; 22(4):266-282. [35] YANG X, ZHANG S, HE C, et al. METTL14 suppresses proliferation and metastasis of colorectal cancer by down-regulating oncogenic long non-coding RNA XIST. Mol Cancer. 2020;19:46. [36] FAN Z, YANG G, ZHANG W, et al. Hypoxia blocks ferroptosis of hepatocellular carcinoma via suppression of METTL14 triggered YTHDF2-dependent silencing of SLC7A11. J Cell Mol Med. 2021;25(21):10197-10212. [37] WANG K, WANG G, LI G, et al. m6A writer WTAP targets NRF2 to accelerate bladder cancer malignancy via m6A-dependent ferroptosis regulation. Apoptosis. 2023; 28(3-4):627-638. [38] WANG W, CHEN J, LAI S, et al. METTL14 promotes ferroptosis in smooth muscle cells during thoracic aortic aneurysm by stabilizing the m6A modification of ACSL4. Am J Physiol Cell Physiol. 2025;328(2):C387-C399. [39] 陈特,肖权洲,孙杨,等.METTL14介导ACSL4的m6A甲基化在髓核细胞铁死亡和细胞衰老中的作用[J].中国现代医学杂志, 2024,34(20):31-39. [40] 宋凯.WTAP调控AR高甲基化修饰促进糖尿病心肌纤维化的分子机制[D].合肥:安徽医科大学,2024. [41] ZHU H, LEE OW, SHAH P, et al. Identification of activators of human fumarate hydratase by quantitative high-throughput screening. SLAS Discov. 2020;25(1):43-56. [42] 刘磊,贾少晗,于鹏.线粒体在铁死亡中的形态特征和作用[J].中国生物化学与分子生物学报,2023,39(6):769-777. [43] PENG H, DOU H, HE S, et al. The role of GOT1 in cancer metabolism. Front Oncol. 2024; 14:1519046. [44] ANDERSEN JV. The Glutamate/GABA-Glutamine Cycle: Insights, Updates, and Advances. J Neurochem. 2025;169(3): e70029. [45] 赵宇翔. c-Myc通过上调GOT1和Nrf2抵抗肝癌细胞中谷氨酰胺剥夺诱发的铁死亡[D].长春:吉林大学,2020. [46] KRMER DM, NELSON BS, LIN L, et al. GOT1 inhibition promotes pancreatic cancer cell death by ferroptosis. Nat Commun. 2021;12: 4860. [47] 徐箫. 综合分析铁死亡相关基因对肝细胞癌患者预后及免疫微环境的影响[D]. 青岛:青岛大学, 2023. [48] BARTOLACCI C, ANDREANI C, EL-GAMMAL Y, et al. Lipid Metabolism Regulates Oxidative Stress and Ferroptosis in RAS-Driven Cancers: A Perspective on Cancer Progression and Therapy. Front Mol Biosci. 2021;8:706650. [49] KOH JY, LEE SJ. Metallothionein-3 as a multifunctional player in the control of cellular processes and diseases. Mol Brain. 2020;13(1):116. [50] GUNN AP, MCLEAN CA, CROUCH PJ, et al. Quantification of metallothionein-III in brain tissues using liquid chromatography tandem mass spectrometry. Anal Biochem. 2021;630:114326. [51] MICHURINA A, SAKIB MS, KERIMOGLU C, et al. Postnatal expression of the lysine methyltransferase SETD1B is essential for learning and the regulation of neuron-enriched genes. EMBO J. 2022; 41(1):e106459. [52] ZENG Z, LAN T, WEI Y, et al. CCL5/CCR5 axis in human diseases and related treatments. Genes Dis. 2022;9(1):12-27. [53] BETTCHER BM, TANSEY MG, DOROTHÉE G, et al. Peripheral and central immune system crosstalk in Alzheimer disease - a research prospectus. Nat Rev Neurol. 2021; 17(11):689-701. [54] PIROLLA NFF, BATISTA VS, DIAS VIGAS FP, et al. Alzheimer’s disease: related targets, synthesis of available drugs, bioactive compounds under development and promising results obtained from multi-target approaches. Curr Drug Targets. 2021; 22(5):505-538. [55] REDDI SREE R, KALYAN M, ANAND N, et al. Newer Therapeutic Approaches in Treating Alzheimer’s Disease: A Comprehensive Review. ACS Omega. 2025;10(6):5148-5171. |
| [1] | Peng Zhiwei, Chen Lei, Tong Lei. Luteolin promotes wound healing in diabetic mice: roles and mechanisms [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(6): 1398-1406. |
| [2] | Lai Jiaming, , Song Yuling, Chen Zixi, Wei Jinghuan, Cai Hao, , Li Guoquan, . Screening of diagnostic markers for endothelial cell Senescence in mice with radiation-induced heart disease and analysis of immune infiltration [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(6): 1450-1463. |
| [3] | Lyu Guoqing, Aizimaitijiang·Rouzi, Xiong Daohai. Irisin inhibits ferroptosis in human articular chondrocytes: roles and mechanisms [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(6): 1359-1367. |
| [4] | Zhang Qian, Huang Dongfeng. Weighted gene co-expression network analysis combined with machine learning to screen and validate biomarkers for osteoarthritis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(5): 1096-1105. |
| [5] | Zou Rongji, Yu Fangfang, Wang Maolin, Jia Zhuopeng. Triptolide inhibits ferroptosis and improves cerebral ischemia-reperfusion injury in a rat model of cerebral artery occlusion/reperfusion [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(4): 873-881. |
| [6] | Yang Xiao, Bai Yuehui, Zhao Tiantian, Wang Donghao, Zhao Chen, Yuan Shuo. Cartilage degeneration in temporomandibular joint osteoarthritis: mechanisms and regenerative challenges [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(4): 926-935. |
| [7] | Gu Fucheng, Yang Meixin, Wu Weixin, Cai Weijun, Qin Yangyi, Sun Mingyi, Sun Jian, Geng Qiudong, Li Nan. Effects of Guilu Erxian Glue on gut microbiota in rats with knee osteoarthritis: machine learning and 16S rDNA analysis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(4): 1058-1072. |
| [8] | Guan Yujie, Zhao Bin. Application and prospect of artificial intelligence in screening and diagnosis of scoliosis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(3): 721-730. |
| [9] | Wang Zhipeng, Zhang Xiaogang, Zhang Hongwei, Zhao Xiyun, Li Yuanzhen, Guo Chenglong, Qin Daping, Ren Zhen. A systematic review of application value of machine learning to prognostic prediction models for patients with lumbar disc herniation [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(3): 740-748. |
| [10] | Zhao Feifan, Cao Yujing. An artificial neural network model of ankylosing spondylitis and psoriasis shared genes and machine learning-based mining and validation [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(3): 770-784. |
| [11] | Su Jiemao, Qi Yansong, Kong Keyu, Zhai Zanjing, Xu Yongsheng. Role of chondrocyte ferroptosis in the pathogenesis of osteoarthritis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(24): 6282-6288. |
| [12] | Zheng Peng, Jia Xiaoning, Tao Jingwei, Fan Xiao. Tetramethylpyrazine improves iron metabolism disorders in a rat model of spinal cord injury via the Keap-1/Nrf2 signaling pathway [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(23): 6134-6141. |
| [13] | Zhou Wen, Yang Hongwei. Molecular mechanism and natural drug screening for ferroptosis-targeted therapy in rheumatoid arthritis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(23): 6051-6061. |
| [14] | Hu Yalin, Huang Fengqin, Yang Boyin, Luo Xingmei. Transcription factor EB improves Alzheimer’s disease via the autophagy-lysosome pathway [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(22): 5844-5858. |
| [15] | Huang Yushan, Wang Rongrong, Li Xiangmiao, Bai Jinzhu. Prostaglandin E1 pretreatment inhibits ferroptosis in endothelial cells in a rat model of spinal cord ischemia-reperfusion injury [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(22): 5716-5727. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||