Chinese Journal of Tissue Engineering Research ›› 2026, Vol. 30 ›› Issue (25): 6544-6553.doi: 10.12307/2026.471
Previous Articles Next Articles
Combined proteomics and metabolomics analysis of pathological mechanisms in mouse models of coronary heart disease
Liu Jinwei1, Zhang Dan2, Guo Hongli2, Chen Huan2, Li Jingjing2, Cao Weiguo1
- 1College of Traditional Chinese Medicine, Chongqing Medical University, Chongqing 400016, China; 2College of Chinese Materia Medica, Chongqing University of Chinese Medicine, Chongqing 402760, China
-
Received:2025-10-15Revised:2026-03-05Online:2026-09-08Published:2026-04-22 -
Contact:Cao Weiguo, MS, Professor, Doctoral supervisor, College of Traditional Chinese Medicine, Chongqing Medical University, Chongqing 400016, China Co-corresponding author: Li Jingjing, PhD, Associate researcher, College of Chinese Materia Medica, Chongqing University of Chinese Medicine, Chongqing 402760, China -
About author:Liu Jinwei, MS candidate, College of Traditional Chinese Medicine, Chongqing Medical University, Chongqing 400016, China -
Supported by:National Natural Science Foundation of China, No. 82304770 (to LJJ); Chongqing Postdoctoral Special Funding, No. 2023CQBSHTB2014 (to LJJ); Chongqing Natural Science Foundation (General Program), Nos. CSTB2024NSCQ-MSX0697 (to LJJ) and CSTB2022NSCQ-MSX1553 (to GHL); Science and Technology Research Program of Chongqing Education Commission, No. KJQN202215132 (to CH)
CLC Number:
Cite this article
Liu Jinwei, Zhang Dan, Guo Hongli, Chen Huan, Li Jingjing, Cao Weiguo. Combined proteomics and metabolomics analysis of pathological mechanisms in mouse models of coronary heart disease[J]. Chinese Journal of Tissue Engineering Research, 2026, 30(25): 6544-6553.
share this article
Add to citation manager EndNote|Reference Manager|ProCite|BibTeX|RefWorks
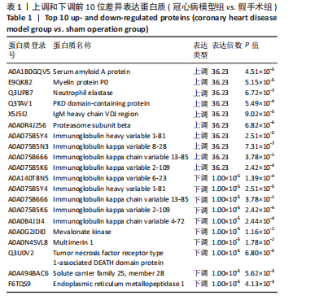
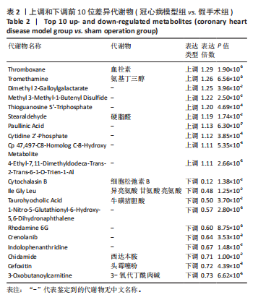
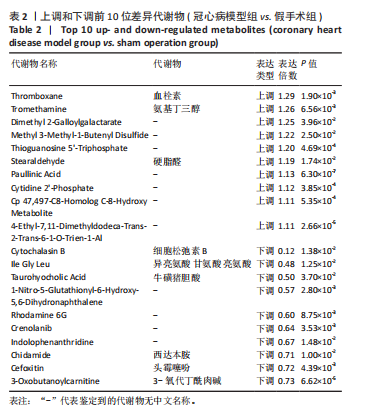
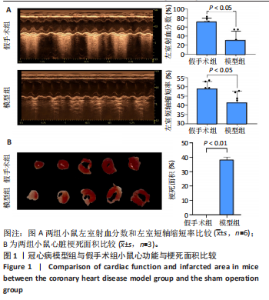
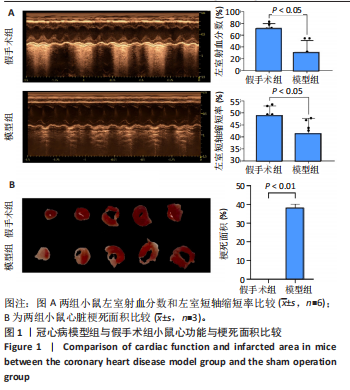
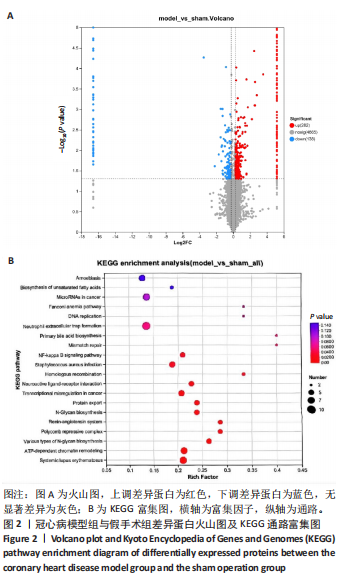
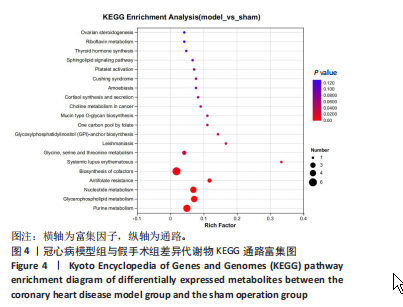
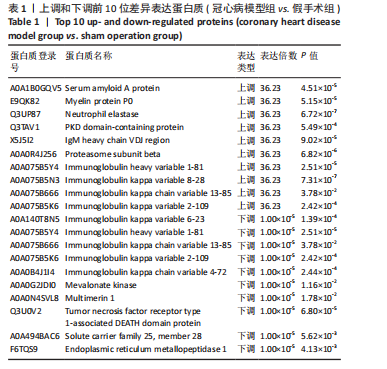
2.1 实验动物数量分析 实验使用20只小鼠,模型组14只造模中死亡8只,共进入结果分析12只。 2.2 超声心动图与TTC染色的分析结果 术后28 d超声心动图显示,与假手术组比较,模型组小鼠左心室前壁变薄、左室射血分数与左室短轴缩短率下降(P < 0.05),且M超声心动图中左心室前壁收缩幅度明显变小,见图1A。用TTC染色法检测两组小鼠的心肌梗死面积,结果如图1B所示,与假手术组相比,模型组小鼠的心肌梗死面积达到40%(P < 0.01)。 2.3 蛋白质组学分析结果 模型组与假手术组两组比较,共筛选出420个差异蛋白质,包括282个上调蛋白质和138个下调蛋白质,见图2A。表1列出了显著变化的前20位差异表达蛋白质。KEGG富集分析结果显示主要涉及ATP依赖性染色质重塑、肾素-血管紧张素系统、核因子κB信号通路等,见图2B。"
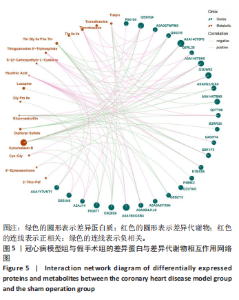
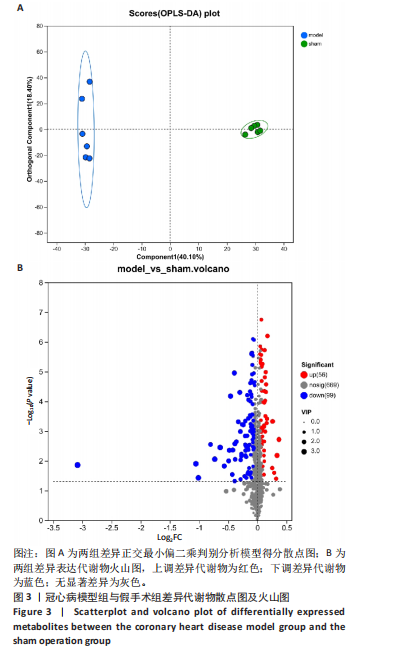
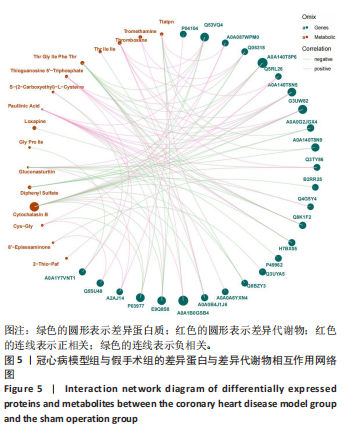
2.5 蛋白质组学与代谢组学联合分析 相关性网络分析是指利用相关性系数建立网络互作关系进行可视化。相关性分析是指对两个或多个具备相关性的变量元素进行分析,从而衡量两个变量因素的相关密切程度。相关性的元素之间需要存在一定的联系或者概率才可以进行相关性分析。基于Pearson相关系数,可以度量样本中蛋白与代谢物之间的关联程度。相关系数的取值范围为(-1,+1)。当相关系数< 0时,称为负相关;> 0时,称正相关;=0时,称为零相关,即没有相关性。此项研究保留至少一组相关性系数绝对值|corr| > 0.95 & P < 0.05 的节点;各取每个组学 log2FC绝对值Top30的节点作图,其中蛋白节点中不包括最大和最小FC值的节点,这种是对照或实验组计算的均值为0的物质,分析软件为python,Igraph。基于Pearson相关系数对155个差异代谢物和420个差异蛋白质进行相关性分析,发现26个差异蛋白与16个差异代谢物存在显著相关性(P < 0.05),见图5。26个差异蛋白集合详情见表3。"
| [1] YANKAI D, ZENGXIAO Z, PIN D, et al. Circulating metabolites and depression: a bidirectional Mendelian randomization.Front Neurosci. 2023;17:1146613. [2] KUISHUAI X, LIANG Z, TIANRUI W, et al. Transcriptome sequencing and bioinformatics analysis of gastrocnemius muscle in type 2 diabetes mellitus rats. BMC Musculoskelet Disord. 2024;25(1):457. [3] 王明韵,刘祎,杨漾,等.基于NPNT/JAK2/STAT3信号通路探讨血府逐瘀汤促冠心病血瘀证模型大鼠心肌修复的作用机制[J].湖南中医药大学学报,2025,45(1):6-14. [4] 陈杏,于子璇,张书萌,等.养心通脉方不同时间干预对早发冠心病血瘀证模型大鼠DNA甲基化的影响[J].中医杂志,2025,66(11):1165-1173. [5] 栾依含,刘先利,宗文静,等.心血管病中医证候蛋白质组学研究概述[J].中国中医基础医学杂志,2016,22(9):1277-1280. [6] 杨光,高嘉良,王阶.冠心病中医证候的蛋白质组学研究进展[J].中华中医药杂志,2022,37(11):6601-6606. [7] 王传池,吴珊,江丽杰,等.1990~2020年我国冠心病中医证的流行病学调查研究概况[J].中国中医基础医学杂志,2020,26(12):1883-1893. [8] 唐武生,万玉莹,李亚强.血清淀粉样蛋白A对急性心肌梗死患者PCI后发生抑郁的预测价值研究[J].中文科技期刊数据库(全文版)医药卫生,2025(1):115-120. [9] GONÇALVES CA, SESTERHEIM P. Serum amyloid A protein has been undervalued as a biomarker of COVID-19. Diabetes Metab Res Rev. 2021; 37(1):e3368. [10] OHOUD M, ENAYAT H, ALI MA, et al. Study of Some Inflammatory Mediators in the Serum of Patients With Atherosclerosis and Acute Myocardial Infarction. Cureus. 2021;13(10):e18450. [11] 杨彩琴,马淑媗,陈颖,等.ST段抬高型心肌梗死病人血清淀粉样蛋白A和一氧化氮水平与预后的关系[J].安徽医药,2021,25(4):809-813. [12] LIN H, ZHANG M, HU M, et al. Emerging applications of single-cell profiling in precision medicine of atherosclerosis. J Transl Med. 2024;22(1):97. [13] PETER D, HAJIME K, JK R, et al. NLRP3 inflammasomes are required for atherogenesis and activated by cholesterol crystals. Nature. 2010;464(7293): 1357-1361. [14] 李淑萍,刘新春,卢娟.益气活血汤联合氢氯吡格雷及阿托伐他汀治疗老年冠心病心绞痛的效果[J].慢性病学杂志,2024,25(11):1668-1670+ 1674. [15] 蒙夏玲,张娇珍,柯海霞,等.冠心病心绞痛中医证型与血清SAA、 Lp-PLA2和FIB水平的关系[J].中国中医急症,2023,32(12):2175-2178. [16] D”URSO D, BROPHY PJ, STAUGAITIS SM, et al. Protein zero of peripheral nerve myelin: biosynthesis, membrane insertion, and evidence for homotypic interaction. Neuron. 1990;4(3):449-460. [17] 谭晗,王韵.蛋白激酶D的生物学功能研究进展[J].生理科学进展, 2011,42(5):359-362. [18] ZHANG L, MALIK S, PANG J, et al. Phospholipase Cε Hydrolyzes Perinuclear Phosphatidylinositol 4-Phosphate to Regulate Cardiac Hypertrophy. Cell. 2013;153(1):216-227. [19] 杨雷,刘暖,毛秉豫.蛋白激酶D1调控胶原表达逆转心肌梗死后心室重构的作用[J].安徽医科大学学报,2019,54(10):1535-1539. [20] 王小平,薛志鹏,杜少兵,等.基于PI3K/PDK1/Akt信号通路研究丹参-红花药对对寒凝血瘀型心肌缺血大鼠的保护作用及机制[J].中草药, 2022,53(16):5085-5092. [21] 张岚,武永新,张涛,等.藁本内酯调控PKD1/HIF-1α/VEGF通路对心力衰竭大鼠的改善作用[J].吉林大学学报(医学版),2024,50(1):42-49. [22] 杨雷,刘萍,刘暖,等.蛋白激酶D1对心肌梗死大鼠心肌组织炎症和凋亡的影响[J].中国药理学通报,2019,35(10):1437-1442. [23] MANSUY-AUBERT V, ZHOU LQ, XIE X, et al. Imbalance between Neutrophil Elastase and its Inhibitor α1-Antitrypsin in Obesity Alters Insulin Sensitivity, Inflammation, and Energy Expenditure. Cell Metab. 2013;17(4):534-548. [24] XU K, COONEY KA, SHIN EY, et al. Adenosine from a biologic source regulates neutrophil extracellular traps (NETs). J Leukoc Biol. 2019;105(6):1225-1234. [25] YUKINO O, KAZUKO T, NOBUYUKI M, et al. Neutrophil Elastase Deficiency Ameliorates Myocardial Injury Post Myocardial Infarction in Mice. Int J Mol Sci. 2021;22(2):722. [26] MASAHIRO F, RYUZO B. Neutrophil Elastase Inhibitor Sivelestat Attenuates Myocardial Injury after Cardioplegic Arrest in Rat Hearts. Ann Thorac Cardiovasc Surg. 2019;26(5):263-269. [27] 甘静,张克,杜函霖,等.中性粒细胞弹性蛋白酶抑制剂sivelestat对HFpEF小鼠心脏纤维化的作用[J].中国病理生理杂志,2025,41(1):11-18. [28] 姚璐,徐继前,杨小博,等.中性粒细胞弹性蛋白酶及其抑制剂在脓毒症中作用的研究进展[J].中华危重病急救医学,2022,34(11): 1209-1212. [29] 张彩凤,司立宁,贺荣章,等.中性粒细胞弹性蛋白酶抑制剂在脂多糖诱导的H9c2心肌细胞损伤中的作用及机制[J].中国高原医学与生物学杂志,2025,46(3):175-183. [30] 张颖函,乔博康,王媛,等.中性粒细胞相关蛋白酶减轻急性心肌梗死早期损伤的研究[J].心肺血管病杂志,2024,43(2):201-206. [31] JIANG Q, NIU R, JIANG J, et al. Association and predictive value of immunoglobulin and complement levels for incident coronary heart disease: a nested case-control study in Chinese adults. Eur J Prev Cardiol. 2025; 28:zwaf091. [32] 毛文慧,袁文肃.TRADD介导的信号通路及其与疾病的相关性[J].基因组学与应用生物学,2022,41(5):1148-1162. [33] YUHONG P, KE L, JUAN H, et al. Molecular cloning, functional characterization of duck TRADD and its effect on infection with duck Tembusu virus. Vet Microbiol. 2022;274:109557. [34] POSNER MG. Multimerin-1 and cancer: a review. Biosci Rep. 2022;42(2): BSR20211248. [35] ALEXANDER L, D’ANDRA P, SUBIA T, et al. Multimerin 1 supports platelet function in vivo and binds to specific GPAGPOGPX motifs in fibrillar collagens that enhance platelet adhesion. J Thromb Haemost. 2020;19(2):547-561. [36] GRYGLEWSKI RJ. Prostaglandins, Platelets, And Atherosclerosi. CRC Crit Rev Biochem. 1980;7(4):291-338. [37] FU LW, PHAN A, LONGHURST JC. Myocardial ischemia-mediated excitatory reflexes: a new function for thromboxane A2? Am J Physiol Heart Circ Physiol. 2008;295(6):H2530-H2540. [38] 范甲卯,张健,谢权.冠心病、急性心肌梗死患者血浆中血栓素、前列腺素的水平及临床意义[J].临床和实验医学杂志,2006,5(5):539. [39] 刘文豹,贾美枝,刘杰,等.血浆血栓素、前列环素在冠心病、急性心肌梗死中的应用[J].标记免疫分析与临床,2002,9(2):115-116. [40] 王紫艳,李磊,刘建勋,等.血瘀证血小板改变及中医药作用研究进展[J].中国中药杂志,2021,46(20):5201-5209. [41] 罗集,杨传华,李艳.原发性高血压血瘀证血小板活化对血管内皮功能的影响[J].山东中医药大学学报,2008,32(4):300-302. [42] 唐诗洋,李金霞,郑彩杏,等.五首活血化瘀方改善血瘀证家兔内皮细胞功能差异[J].中国中西医结合杂志,2023,43(10):1214-1220. [43] 俞佳峰,胡翠,吴先昊,等.温性活血化瘀中药及其活性成分抗血小板聚集的作用机制[J].中兽医医药杂志,2023,42(3):34-40. [44] LEHMANN GK, POPMA JJ, WERNER AJ, et al. Vascular remodeling and the local delivery of cytochalasin B after coronary angioplasty in humans. J Am Coll Cardiol. 2000;35(3):583-591. [45] SEGUIN A, JIA X, EARL AM, et al. The mitochondrial metal transporters mitoferrin1 and mitoferrin2 are required for liver regeneration and cell proliferation in mice. J Biol Chem. 2020;295(32):11002-11020. [46] RICHARDSON DR, LANE DJ, BECKER EM, et al. Mitochondrial iron trafficking and the integration of iron metabolism between the mitochondrion and cytosol. Proc Natl Acad Sci U S A. 2010;107(24):10775-10782. [47] DONGCHEN W, PENG Y, CHAOHUA K, et al. Mitoferrin 2 deficiency prevents mitochondrial iron overload-induced endothelial injury and alleviates atherosclerosis. Exp Cell Res. 2021;402(1):112552. [48] PALMGREN GM, NISSEN P. P-Type ATPases. Annu Rev Biophys. 2011;40: 243-266. [49] MOLDOVAN R, ICHIM AV, BELIȘ V. Immunohistochemical study of ATP1A3 and plakophilin 2 as new potential markers in the diagnosis of myocardial ischemia. Leg Med (Tokyo). 2025;72:102565. [50] 张露丹,孙艳君,刘凡琪,等.心力衰竭气虚血瘀证相关生物学标志物研究述评与展望[J].天津中医药,2024,41(2):258-263. [51] SHUXIAN C, YUANMING Z, CHUNYU S, et al. The role of glycolytic metabolic pathways in cardiovascular disease and potential therapeutic approaches.Basic Res Cardiol. 2023;118(1):48. [52] YEIH DF, YEH HI, LIN LY, et al. Enhanced activity and subcellular redistribution of myocardial hexokinase after acute myocardial infarction. Int J Cardiol. 2011;149(1):74-79. [53] CALMETTES G, JOHN SA, WEISS JN, et al. Hexokinase-mitochondrial interactions regulate glucose metabolism differentially in adult and neonatal cardiac myocytes. J Gen Physiol. 2013;142(4):425-436. [54] TATEKOSHI Y, MAHMOODZADEH A, SHAPIRO SJ, et al. Protein O-GlcNAcylation and hexokinase mitochondrial dissociation drive heart failure with preserved ejection fraction. Cell Metab. 2025;37(7):1584-1600.e10. [55] JIE S, JYOTSNA M, MEIYING Y, et al. Hypothermia Prevents Cardiac Dysfunction during Acute Ischemia Reperfusion by Maintaining Mitochondrial Bioenergetics and by Promoting Hexokinase II Binding to Mitochondria. Oxid Med Cell Longev. 2022;2022:4476448. [56] UTHMAN L, KUSCHMA M, RÖMER G, et al.Novel Anti-inflammatory Effects of Canagliflozin Involving Hexokinase II in Lipopolysaccharide-Stimulated Human Coronary Artery Endothelial Cells. Cardiovasc Drugs Ther. 2020; 35(6):1083-1094. [57] SHUXIAN C, YUANMING Z, CHUNYU S, et al. The role of glycolytic metabolic pathways in cardiovascular disease and potential therapeutic approaches.Basic Res Cardiol. 2023;118(1):48. [58] MCDANIEL ML, KING S, ANDERSON S, et al.Effect of cytochalasin B on hexose transport and glucose metabolism in pancreatic islets. Diabetologia. 1974;10(4):303-308. |
| [1] | Chen Yulin, He Yingying, Hu Kai, Chen Zhifan, Nie Sha Meng Yanhui, Li Runzhen, Zhang Xiaoduo , Li Yuxi, Tang Yaoping. Effect and mechanism of exosome-like vesicles derived from Trichosanthes kirilowii Maxim. in preventing and treating atherosclerosis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(7): 1768-1781. |
| [2] | Hu Yujie, Xie Ping, , Lu Weijie, Yang Kang, Deng Yaoting, Liu Mengyang. Meta-analysis of the clinical efficacy of high-intensity interval exercise and middle-intensity continuous training in patients with coronary heart disease [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(10): 2584-2593. |
| [3] | Liu Lin, Liu Shixuan, Lu Xinyue, Wang Kan. Metabolomic analysis of urine in a rat model of chronic myofascial trigger points [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(8): 1585-1592. |
| [4] |
Sun Guanghan, Xie Zhencong, Sun Mi, Xu Yang, Guo Dong.
Therapeutic effect and mechanism by which Trichosanthis Fructus-Allii Macrostemonis Bulbus regulates gut microbiota in a rat model of coronary heart disease #br#
#br#
[J]. Chinese Journal of Tissue Engineering Research, 2025, 29(5): 917-927.
|
| [5] | Zhang Yixuan, Li Dongna, Liu Chunyan. Pathological processes, inflammatory responses, and related biomarkers of periodontitis: a multi-omics analysis [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(35): 7601-7610. |
| [6] | Liu Haowei, Tian Haodong, Huang Li, Yu Hanglin, Peng Li. Acute effects of blood flow restriction resistance exercise on serum metabolites in obese young men [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(29): 6249-6259. |
| [7] | Chai Jinlian, Sun Tiefeng, Li Wei, Zhang Bochun, Li Guangzheng, Zhou Zhongqi, Liang Xuezhen, Wang Ping. Therapeutic effect of Cornus Cervi Colla on steroid-induced osteonecrosis of the femoral head in rat models: fecal metabolomics analysis [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(29): 6187-6197. |
| [8] | Wang Zhifeng, Yang Jiao, Xi Yujiang, Xu Shuangfeng, Shi Ting, Lan Junfeng, Hao Zhihui, He Pengfen, Yang Aiming, Pan Pan, Wang Jian. Biomarkers affecting the progression of mild to moderate cognitive impairment after stroke: #br# a non-targeted metabolomics analysis [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(24): 5116-5126. |
| [9] | Liang Zhou, Zhang Chi, Pan Chengzhen, Yang Bo, Pu Zhanglin, Liu Hua, Peng Jinhui, Wen Lichun, Ling Guanhan, Chen Feng. Anti-osteoporotic mechanisms of kaempferol based on gut microbiota and comprehensive targeted metabolomics [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(20): 4190-4204. |
| [10] | Pei Yunxiang, Wu Hao. Hyperbaric oxygen intervention eliminates exercise-induced fatigue in a high-intensity interval training shock microcycle [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(14): 2979-2988. |
| [11] | Zhang Xiaoyun, Li Kunjian, Mo Jian, Chai Yuan, Huang Yourong . Current status and trends of research on animal models of postmenopausal osteoporosis: a bibliometric visualization analysis [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(14): 3070-3080. |
| [12] | Chen Xinfei, Dai Yahui, Xie Bingying, Huang Xiaobin, Huang Huimin, Huang Jingwen, Li Shengqiang, Ge Jirong. Metabolomics analysis of the lumbar spine after alendronate sodium intervention in ovariectomized rats with osteoporosis [J]. Chinese Journal of Tissue Engineering Research, 2025, 29(11): 2277-2284. |
| [13] | Song Jiating, Chen Jianmin, Wang Kewen, Huang Lanying, Xu Senming, Gui Yuchang, Xu Jianwen. Metabolomics analysis of serum and urine in patients with traumatic spinal cord injury [J]. Chinese Journal of Tissue Engineering Research, 2024, 28(在线): 1-6. |
| [14] | Zhou Chao, Yu Guangyu, Fan Jiaqi, Yu Chunmei, Wu Min, Chen Shibei. Establishment and validation of embryo high-quality prediction models based on the third-day 340 nm absorbance embryo culture [J]. Chinese Journal of Tissue Engineering Research, 2024, 28(7): 1050-1056. |
| [15] | Bu Xianzhong, Zhong Yuanming, Bu Baoxian, Li Jitian, Wang Lihe, Li Huiying, Yang Hanli, Xu Wei. Serum differential proteomic analysis of developmental cervical canal stenosis [J]. Chinese Journal of Tissue Engineering Research, 2024, 28(34): 5432-5439. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||