Chinese Journal of Tissue Engineering Research ›› 2026, Vol. 30 ›› Issue (34): 9041-9047.doi: 10.12307/2026.819
Previous Articles Next Articles
Key role of m6A methylation in sarcopenia
Li Jiatong1, Liang Songlin1, Liu Runjia2, Li Nianhu1, 3
- 1Shandong University of Traditional Chinese Medicine, Jinan 250014, Shandong Province, China; 2Guang’anmen Hospital, Chinese Academy of Chinese Medical Sciences, Beijing 100032, China; 3Affiliated Hospital of Shandong University of Traditional Chinese Medicine, Jinan 250014, Shandong Province, China
-
Received:2025-08-01Revised:2025-12-08Online:2026-12-08Published:2026-04-15 -
Contact:Li Nianhu, MD, Professor, Doctoral supervisor, Chief physician, Shandong University of Traditional Chinese Medicine, Jinan 250014, Shandong Province, China; Affiliated Hospital of Shandong University of Traditional Chinese Medicine, Jinan 250014, Shandong Province, China -
About author:Li Jiatong, Shandong University of Traditional Chinese Medicine, Jinan 250014, Shandong Province, China -
Supported by:Shandong Provincial Natural Science Foundation Project, No. ZR2023MH063 (to LNH); Xu Zhanwang National Famous Veteran Traditional Chinese Medicine Experts Inheritance Studio Construction Project, No. [2022]75 (to LNH)
CLC Number:
Cite this article
Li Jiatong, Liang Songlin, Liu Runjia, Li Nianhu. Key role of m6A methylation in sarcopenia[J]. Chinese Journal of Tissue Engineering Research, 2026, 30(34): 9041-9047.
share this article
Add to citation manager EndNote|Reference Manager|ProCite|BibTeX|RefWorks
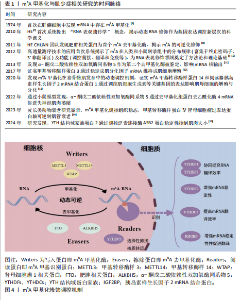
2.1 m6A甲基化与肌少症相关研究的时间脉络 见表1。 2.2 m6A甲基化概述 m6A是真核生物RNA中最普遍且保守的内部甲基化修饰,自20世纪70年代首次在肝癌细胞中被发现以来,m6A逐渐成为表观转录组学的研究热点[6]。m6A甲基化广泛分布于多种RNA类型中,其修饰位点呈现显著的序列偏好性,主要富集于mRNA的终止密码子附近、3’非翻译区及编码序列中,常见于保守基序DRACH(D=G/A/U,R=G/A,H=A/U/C)[16],m6A甲基化通过动态可逆的表观调控网络参与RNA生命周期的全阶段调控,包括剪接、翻译、亚细胞定位以及降解过程。在生理功能层面,m6A甲基化调控造血干细胞分化、神经发育及配子成熟,还可以影响免疫应答、脂肪生成等生物过程[17]。病理状态下,m6A异常与急性髓系白血病[18]、胶质瘤[19]、肝细胞癌等癌症密切相关[20],m6A 还参与糖尿病等代谢疾病[21]、阿尔茨海默症等神经退行性疾病[22]、心力衰竭等心血管疾病的发生发展[23]。 近年来,基于高通量测序法RNA甲基化免疫共沉淀(methylated RNA immunoprecipitation sequencing,MeRIP-seq)技术的突破,m6A甲基化的分子机制研究取得重大进展。这一过程受到多种蛋白质复合体的调控,主要有3种关键的分子:甲基化酶(writers)、去甲基化酶(erasers)和阅读蛋白(readers)参与了m6A甲基化的进程,分别起到添加、去除和识别m6A位点的作用[5],见图4。"
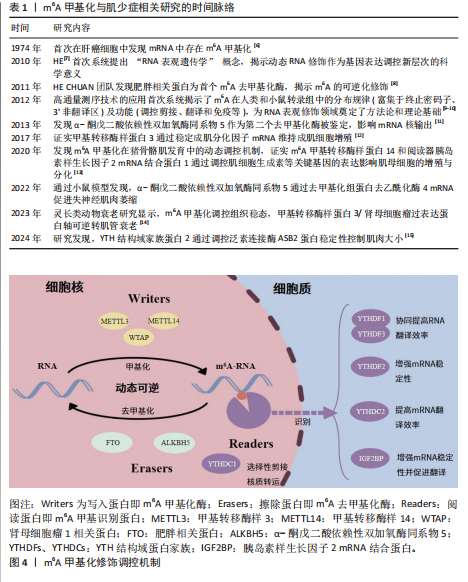
2.2.1 m6A甲基化酶 甲基化酶,也称为甲基转移酶,通过将供体中的甲基转移到RNA腺苷第6位氮原子上,添加甲基化修饰,从而启动RNA甲基化修饰过程。甲基转移酶主要包括甲基转移酶样蛋白3、甲基转移酶样蛋白14和肾母细胞瘤1相关蛋,其中仅有甲基转移酶样蛋白3是该甲基转移酶复合物中具有催化活性的亚基,负责甲基转移,而甲基转移酶样蛋白14提供结构支持,激活并增强甲基转移酶样蛋白3的催化活性[24];肾母细胞瘤1相关蛋白没有催化活性,但能与甲基转移酶样蛋白3和甲基转移酶样蛋白14构成的异二聚体相互作用,辅助对目标位点的准确修饰,从而调节RNA转录的m6A水平[24]。甲基转移酶样蛋白3与甲基转移酶样蛋白14、肾母细胞瘤1相关蛋白一起构成的复合体将m6A修饰稳定在RNA上[25]。 2.2.2 m6A去甲基化酶 去甲基化酶,可以擦除RNA甲基化修饰达到RNA修饰的去甲基化结果。m6A的甲基化酶执行在mRNA上“写入”甲基基团的功能,而去甲基化酶则执行“擦除”这一沉积物,这就表明m6A甲基化过程是动态可逆的。去甲基化酶主要是肥胖相关蛋白和α-酮戊二酸依赖性双加氧酶同系物5,肥胖相关蛋白主要通过连续的氧化移除m6A的甲基基团[26],而α-酮戊二酸依赖性双加氧酶同系物5则直接从mRNA上消除m6A修饰[11]。 2.2.3 m6A甲基识别蛋白 阅读蛋白可以特异性地识别m6A并与之结合,读取修饰上的信息,从而指导RNA在下游序列的翻译和降解。m6A阅读蛋白主要包括3个蛋白家族:YTH结构域蛋白家族,如YTH结构域包含蛋白1-2和YTH结构域家族蛋白1-3;胰岛素样生长因子2 mRNA 结合蛋白,如胰岛素样生长因子2 mRNA结合蛋白 1-3;异质核核糖核蛋白,如异质核核糖核蛋白A2B1、异质核核糖核蛋白C和异质核核糖核蛋白G[27]。 功能上,YTH结构域家族蛋白1和YTH结构域家族蛋白3协同作用可以提高目标 RNA的翻译效率[28],YTH结构域家族蛋白2在对mRNA稳定性的调控上发挥着作用[29]。YTH结构域包含蛋白1可以通过募集和调节前体mRNA剪接因子来调节mRNA选择性剪接[30],能够促进细胞核内mRNA向胞浆内转移[31]。而YTH结构域包含蛋白2可提高目标靶点的翻译效率,同时会降低其靶点的丰度,目前发现YTH结构域包含蛋白2在调节哺乳动物的精子形成中起重要作用[32]。胰岛素样生长因子2 mRNA结合蛋白在正常和应激条件下以依赖m6A的方式促进其靶mRNA的稳定性和储存,从而影响基因表达输出[33]。此外,其他类型的“阅读器”研究也在不断深入,例如异质核核糖核蛋白A2B1也有选择性剪接效应,还可以促进microRNA(miRNA)的加工[34];异质核核糖核蛋白C通过丝裂原活化蛋白激酶通路增强白细胞介素1受体相关激酶1 mRNA的稳定性[35];脆性X智力障碍蛋白通过募集异质核核糖核蛋白M影响溶质载体家族7成员11的剪接以减少脂质活性氧的产生,对维持铁对变态反应的抗性至关重要,可能是改善乳腺癌患者预后的一种潜在治疗方案[36]。与m6A甲基化酶和m6A去甲基化酶相比,阅读器家族具有更高的功能特异性和调控复杂性。值得关注的是,阅读器家族成员数目最为庞大且调控机制复杂,其功能多样性与分子机制尚未完全阐明,因此,在疾病发生发展机制研究及新型治疗靶点开发等领域具有广阔的探索空间。 2.3 m6A在肌少症中的作用机制 m6A甲基化在肌少症中的作用机制,见图5。 2.3.1 调控卫星细胞 在探讨m6A甲基化与肌少症之间的关联时,目前研究焦点主要集中在肌肉干细胞的增殖分化能力上。骨骼肌卫星细胞作为一类具有自我更新和多向分化潜能的成体干细胞,通常处于静止状态,位于肌纤维基底膜和肌膜之间,在肌肉受损修复或受到运动、药物刺激后,它们会进入激活状态,开始增殖和分化[37]。配对盒7和卫星细胞在受伤时被激活,增殖为前体细胞肌母细胞,分化为肌管,并成熟为肌肉。在肌肉修复和再生的过程中,肌生成调节因子家族成员的表达起着核心调控作用,包括成肌分化因子、肌细胞生成素、生肌调节因子4、生肌因子5和肌球蛋白重链,这些肌生成调节因子家族成员的表达对于卫星细胞的增殖和分化至关重要,进而影响肌肉损伤的修复过程[38]。 KUDOU等[12]通过甲基转移酶样蛋白3敲低实验证实,在增殖性肌源细胞中,甲基转移酶样蛋白3可通过在成肌分化因子mRNA的5’非翻译区引入m6A修饰,维持成肌分化因子 mRNA的加工成熟及稳定表达;功能实验显示,成肌细胞中甲基转移酶样蛋白3缺失会显著下调成肌分化因子 mRNA水平,进而阻碍骨骼肌分化进程,提示甲基转移酶样蛋白3介导的m6A修饰对肌源细胞分化潜能的维持至关重要。GHELLER等[39]的研究发现,在小鼠成肌细胞(小鼠C2C12细胞)和卫星细胞中敲低甲基转移酶样蛋白3可以降低m6A修饰水平,并导致成年肌细胞过早分化,破坏骨骼肌再生所需的增殖-分化动态平衡,不利于骨骼肌修复,同样说明了甲基转移酶样蛋白3对骨骼肌分化再生的重要性;值得注意的是,移植前降低卫星细胞的甲基转移酶样蛋白3表达虽可提升初次移植时的细胞定植效率,但会导致卫星细胞丧失连续移植能力,提示甲基转移酶样蛋白3参与维持干细胞自我更新特性,它的功能异常可能引发干细胞干性的过早耗竭。在分子机制层面,LIANG等[40]通过高通量测序法RNA甲"
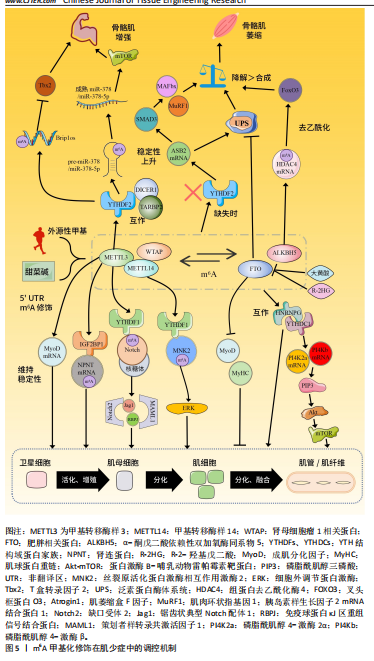

基化免疫共沉淀及功能验证发现,甲基转移酶样蛋白3介导的m6A修饰通过调控Notch信号通路关键组分(如缺口受体2、锯齿状典型Notch配体1、免疫球蛋白κJ区重组信号结合蛋白)的翻译效率,在卫星细胞的自我更新与分化抉择中发挥核心调控作用;进一步研究揭示,甲基转移酶样蛋白3/甲基转移酶样蛋白14复合体可直接靶向丝裂原活化蛋白激酶信号通路中的关键节点分子丝裂原活化蛋白激酶相互作用激酶2,通过m6A修饰依赖机制招募阅读器YTH结构域家族蛋白1,特异性提升丝裂原活化蛋白激酶相互作用激酶2表达水平,进而激活细胞外调节蛋白激酶/丝裂原活化蛋白激酶信号传导。作为调控细胞增殖的核心通路,细胞外调节蛋白激酶信号的激活可促进肌源细胞增殖,加速损伤后骨骼肌修复[41]。 LI等[42]体外研究显示,成肌细胞分化过程中m6A去甲基化酶肥胖相关蛋白与α-酮戊二酸依赖性双加氧酶同系物5表达升高,伴随m6A水平下降;施加肥胖相关蛋白特异性抑制剂大黄酸干预后提升了m6A水平,降低了成肌关键基因肌球蛋白重链/肌细胞生成素mRNA及蛋白表达,阻滞了成肌细胞向肌管的分化过程。在氯化钡诱导的动物骨骼肌损伤模型中,大黄酸干预后损伤肌肉m6A水平升高,肌纤维横截面积减小,成熟标志肌球蛋白重链表达下调,并且未成熟肌细胞核滞留于纤维中央,提示抑制肥胖相关蛋白会延缓骨骼肌再生与重塑。该研究印证了抑制肥胖相关蛋白介导的m6A去甲基化会增加RNA甲基化修饰,抑制成肌分化并延缓骨骼肌修复。 YANG 等[43]系统探讨了5个关键m6A甲基化调控因子对牛骨骼肌成肌细胞凋亡的调控作用,发现敲低肥胖相关蛋白在mRNA/蛋白水平增强了抗凋亡因子B细胞超大淋巴瘤和B细胞淋巴瘤2蛋白的表达,但流式检测显示促进了细胞凋亡;敲低甲基转移酶样蛋白3显著抑制了细胞凋亡;敲低α-酮戊二酸依赖性双加氧酶同系物5和 肾母细胞瘤1相关蛋白均显著促进细胞凋亡,并上调促凋亡因子B淋巴细胞瘤-2基因相关启动子、半胱天冬酶3和半胱天冬酶6的表达(mRNA 和/或蛋白水平),其中肾母细胞瘤1相关蛋白敲低还抑制了抗凋亡因子B细胞超大淋巴瘤和B细胞淋巴瘤2蛋白的表达;敲低甲基转移酶样蛋白14则对细胞凋亡无显著影响。结果表明,m6A甲基化酶通过差异性调控凋亡相关基因的表达网络参与牛骨骼肌成肌细胞的凋亡进程,为解析m6A甲基化在骨骼肌发育中的复杂调控网络提供了重要实验依据。 QIAO等[44]通过体内外功能实验与线性扩增cDNA末端测序技术发现,m6A阅读蛋白YTH结构域包含蛋白1是卫星细胞激活与增殖的关键调控因子;转录组分析显示,YTH结构域包含蛋白1通过识别m6A修饰的靶标mRNA,参与调控前体mRNA剪接及成熟mRNA核输出过程;研究进一步发现,m6A阅读蛋白异质核核糖核蛋白G与YTH结构域包含蛋白1存在相互作用并协同调控卫星细胞增殖,提示YTH结构域包含蛋白1可通过多维度RNA加工机制介导卫星细胞功能。LIU等[45]基于条件性基因敲除小鼠模型证实,YTH结构域包含蛋白1通过结合m6A修饰的磷脂酰肌醇4-激酶2α和磷脂酰肌醇4-激酶β mRNA调控磷脂酰肌醇4-激酶的可变剪切,进而激活蛋白激酶B-哺乳动物雷帕霉素靶蛋白信号通路,驱动卫星细胞从静息状态退出并进入细胞周期,最终促进骨骼肌再生。蛋白激酶B-哺乳动物雷帕霉素靶蛋白信号通路已经被证明是维持肌肉的稳态和大小的关键通路,该信号通路持续激活会增加肌纤维横截面积,以往研究多集中体育锻炼、中药活性成分作用等干预在胰岛素样生长因子结合蛋白7-蛋白激酶B-哺乳动物雷帕霉素靶蛋白轴与磷脂酰肌醇激酶-蛋白激酶B-哺乳动物雷帕霉素靶蛋白轴在骨骼肌细胞中的作用[46-49],为解析表观转录组与细胞信号网络的交互作用提供了新视角,作为骨骼肌生长与再生的核心调控通路,蛋白激酶B-哺乳动物雷帕霉素靶蛋白仍有重要的研究价值。ZHAO等[50]的研究发现,YTH结构域家族蛋白1的敲低减少了肌生成调节因子家族(包括肌球蛋白重链、成肌分化因子和肌细胞生成素)的mRNA翻译效率,进而减弱了甲基转移酶样蛋白3过表达对肌源性分化的促进作用,而YTH结构域家族蛋白2的敲低减弱了甲基转移酶样蛋白3过表达对细胞增殖的促进作用,指出了阅读蛋白的在骨骼肌发育中的不同作用。HUANG等[51]的研究聚焦YTH结构域家族蛋白1在调控骨骼肌纤维类型组成中的关键作用,体内表型分析显示,YTH结构域家族蛋白1基因敲除可诱导显著的骨骼肌肥大(表现为肌纤维横截面积增大和肌肉质量增加)并显著提升小鼠的跑步耐力等有氧运动能力;深入的组织学分析发现,在YTH结构域家族蛋白1敲除小鼠的骨骼肌中,Ⅰ型肌球蛋白重链阳性纤维的数量显著增加,Ⅰ型肌球蛋白重链是慢收缩氧化型肌纤维的特异性分子标志物,它的增多直接表明慢肌纤维比例升高;此外,琥珀酸脱氢酶染色阳性纤维也显著增多,琥珀酸脱氢酶作为线粒体电子传递链复合体Ⅱ的关键组分,它的活性增强是慢肌纤维依赖氧化磷酸化供能的典型特征,进一步证实了慢肌纤维在YTH结构域家族蛋白1缺失骨骼肌中的富集。这些发现共同表明,YTH结构域家族蛋白1缺失促进了骨骼肌向更具氧化代谢特性、更耐疲劳的慢肌纤维表型转化。 综上所述,甲基转移酶样蛋白3/14通过m6A RNA甲基化YTH结构域家族蛋白1/2信号轴调控卫星细胞,促进肌细胞的增殖和肌源性分化,甲基转移酶样蛋白3对于调节卫星细胞至关重要,外源性甲基转移酶样蛋白3的递送可以诱导骨骼肌生长[52]。最新研究发现,以跑步为代表的有氧运动机械刺激显著增加了甲基转移酶样蛋白3介导的m6A甲基化,提高了成肌分化因子的表达水平;此外,甲基供体甜菜碱也能够通过增强m6A甲基化水平来增强成肌分化因子表达,这提示运动和营养补充在促进骨骼肌生长中的巨大潜力[53]。在骨骼肌衰老研究领域,灵长类动物模型研究进一步揭示了甲基转移酶样蛋白3的关键调控作用,研究显示,衰老骨骼肌中甲基转移酶样蛋白3水平较年轻对照组降低50%以上;功能验证表明,甲基转移酶样蛋白3缺失可导致人多能干细胞衍生肌管的衰老进程显著加速,伴随细胞凋亡水平显著升高;机制上,甲基转移酶样蛋白3通过催化mRNA的m6A修饰靶向下游维持骨骼肌稳态的关键因子肾连蛋白的 mRNA;进一步研究发现,m6A阅读蛋白胰岛素样生长因子2 mRNA 结合蛋白1可特异性识别肾连蛋白mRNA的m6A修饰位点,通过维持其 mRNA 稳定性来调控蛋白表达,进而对抗肌管萎缩[14]。作为首项系统探讨m6A表观转录组在灵长类动物骨骼肌衰老中作用的研究,该研究鉴定的甲基转移酶样蛋白3-m6A-肾连蛋白调控轴具有重要科学意义,该发现不仅揭示了骨骼肌衰老的新型表观调控机制,还为衰老相关肌少症的干预提供了潜在靶点。基于上述发现,未来研究可聚焦于m6A修饰谱作为衰老生物标志物的可行性,探索建立“m6A衰老时钟”的可能性,从衰老表观转录组层面为肌少症的早期诊断和治疗提供新思路。 m6A甲基化对卫星细胞的调控机制,见表2。 2.3.2 调控蛋白代谢 蛋白质的合成与分解处于动态平衡状态,这种状态被称为蛋白质稳态,它对于保持骨骼肌的体积至关重要。如果蛋白质的分解速率超过了合成速率,就会导致肌肉量的显著减少,最终可能发展成肌少症[54]。 调控泛素蛋白酶体系统:是骨骼肌蛋白代谢的主要调节机制,泛素连接酶(E3)在决定泛素蛋白酶体系统的选择性和特异性方面起着核心作用[55]。GILBERT等[15]的研究表明,YTH结构域家族蛋白2通过调控m6A修饰的mRNA稳定性,特别是对泛素连接酶ASB2的mRNA进行靶向调控,进而控制ASB2蛋白的表达水平,最终影响骨骼肌蛋白质稳态与肌纤维体积;机制上,YTH结构域家族蛋白2的缺失会导致ASB2 mRNA稳定性升高,促使ASB2蛋白水平显著增加。作为E3泛素连接酶复合体组分,ASB2通过与SMAD3蛋白相互作用,激活转化生长因子β/SMAD信号通路诱导抗肥大基因肌萎缩盒F因子/和肌肉环状指基因1的转录激活,最终增强泛素蛋白酶体系统介导的蛋白质降解过程,诱发骨骼肌萎缩[15,56]。相较于促进合成的蛋白激酶B-哺乳动物雷帕霉素靶蛋白信号通路,转化生长因子β/SMAD信号通路作为负调节途径在骨骼肌质量调控中发挥关键作用。研究显示,转化生长因子β/SMAD信号通路的核心效应分子SMAD3,可通过上调泛素连接酶基因表达推动蛋白降解程序激活;值得注意的是,肌肉生长抑制素作为转化生长因子β家族成员,正是通过该信号通路负向调控肌肉生长并促进萎缩进程,提示靶向抑制转化生长因子β/SMAD 信号可能成为干预肌少症和恶病质的潜在策略[57]。在去神经诱导的肌肉萎缩模型中,泛素蛋白酶体系统被证实为m6A甲基化修饰的关键下游通路,实验数据显示,使用肥胖相关蛋白抑制剂R-2-羟基戊二酸提升m6A甲基化水平可显著缓解去神经骨骼肌的萎缩表型,表现为肌纤维横截面积和肌肉质量的改善,这一发现为以m6A甲基化调控为靶点的抗萎缩治疗提供了实验依据,有望推动相关干预策略的临床转化[58]。 调控相关信号通路:叉头框蛋白O是磷脂酰肌醇激酶/蛋白激酶B信号通路的下游靶点,是肌肉萎缩的重要调节因子[59]。基于去神经骨骼肌萎缩模型的研究表明,在神经源性肌肉萎缩进程中,m6A去甲基化酶α-酮戊二酸依赖性双加氧酶同系物5的异常高表达,可通过特异性靶向组蛋白去乙酰化酶4的mRNA发挥作用;机制上,α-酮戊二酸依赖性双加氧酶同系物5通过催化组蛋白去乙酰化酶4 mRNA的3’-非翻译区发生去甲基化修饰,有效延长该mRNA的半衰期并增强其稳定性,进而显著提升组蛋白去乙酰化酶4 的表达水平,作为关键的分子桥梁,组蛋白去乙酰化酶4通过与叉头框蛋白O3直接结合介导后者的去乙酰化修饰,该过程可抑制叉头框蛋白O3的泛素化降解并促进其核内积累,最终导致叉头框蛋白O3丰度及其转录活性的双重增强;功能上,这一信号轴的激活可显著诱导肌萎缩盒F因子、肌肉环状指基因1等萎缩相关基因的表达,从而加速去神经肌肉的萎缩进程[60]。研究数据显示,靶向干预α-酮戊二酸依赖性双加氧酶同系物5可通过下调组蛋白去乙酰化酶4/叉头框蛋白O3信号通路的激活程度,有效改善去神经肌肉的质量丢失及肌纤维横截面积减少,提示α-酮戊二酸依赖性双加氧酶同系物5有望成为神经源性肌肉萎缩防治的潜在分子靶点。 m6A甲基化调控蛋白代谢的机制,见表3。 2.3.3 调控非编码RNA miRNA作为一类内源性非编码小分子RNA,在基因表达调控中发挥关键作用。在骨骼肌发育与分化的研究体系中,甲基转移酶样蛋白3过表达可显著抑制骨骼肌特异性miRNA(如miR-1a、miR-133a、"

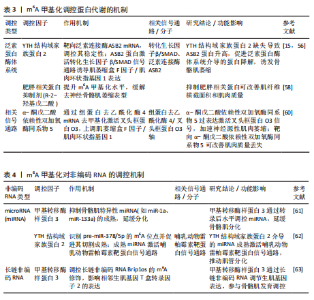
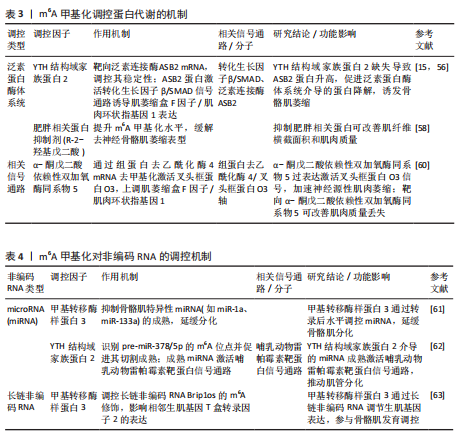
miR-133b及miR-206)的成熟水平,延缓骨骼肌的分化[61]。作为m6A甲基化的关键“阅读器”蛋白,YTH结构域家族蛋白2可通过调控前体miR-378/miR-378-5p的成熟过程参与肌原分化调控:功能上,miR-378/miR-378-5p在肌管形成及肌原分化标志物(如肌球蛋白重链、成肌分化因子)的表达调控中发挥核心作用;机制层面,YTH结构域家族蛋白2通过识别前体miRNA上的特定m6A修饰位点,与DICER1、反式激活元件RNA结合蛋白2等miRNA加工复合体核心组分形成功能性互作,促进pre-miR-378/miR-378-5p的切割成熟,生成的成熟miRNA进一步通过激活哺乳动物雷帕霉素信号通路,推动肌原细胞向肌管分化,从而构建了“m6A修饰-YTH结构域家族蛋白2-miRNA-哺乳动物雷帕霉素靶蛋白”的层级调控轴[62]。此外,m6A甲基化在骨骼肌生成过程中对长链非编码RNA的表达具有显著影响,甲基转移酶样蛋白3/m6A/Brip1os轴调控Brip1os这一长链非编码RNA,Brip1os的下调表达会导致相邻生肌基因T盒转录因子2的表达上调,总之,甲基转移酶样蛋白3通过甲基转移酶样蛋白3/ Brip1os /T盒转录因子2轴调节骨骼肌发育[63]。这些发现表明,甲基转移酶样蛋白3在转录后水平影响肌肉特异性非编码RNA,在骨骼肌分化和发育过程中起着至关重要的作用。 m6A甲基化对非编码RNA的调控机制,见表4。"
| [1] CHEN LK, WOO J, ASSANTACHAI P, et al. Asian working group for sarcopenia: 2019 consensus update on sarcopenia diagnosis and treatment. J Am Med Dir Assoc. 2020;21(3):300-307.e2. [2] YUAN S, LARSSON SC. Epidemiology of sarcopenia: Prevalence, risk factors, and consequences. Metabolism. 2023;144:155533. [3] PETERMANN-ROCHA F, BALNTZI V, GRAY SR, et al. Global prevalence of sarcopenia and severe sarcopenia: A systematic review and meta‐analysis. J Cachexia Sarcopenia Muscle. 2022; 13(1):86-99. [4] LIANG W, XU F, LI L, et al. Epigenetic control of skeletal muscle atrophy. Cell Mol Biol Lett. 2024; 29(1):99. [5] ZACCARA S, RIES RJ, JAFFREY SR. Reading, writing and erasing mRNA methylation. Nat Rev Mol Cell Biol. 2019;20(10):608-624. [6] DESROSIERS R, FRIDERICI K, ROTTMAN F. Identification of methylated nucleosides in messenger RNA from novikoff hepatoma cells. Proc Natl Acad Sci U S A. 1974;71(10):3971-3975. [7] HE C. Grand challenge commentary: RNA epigenetics? Nat Chem Biol. 2010;6(12):863-865. [8] JIA G, FU Y, ZHAO X, et al. N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat Chem Biol. 2011;7(12):885-887. [9] DOMINISSINI D, MOSHITCH-MOSHKOVITZ S, SCHWARTZ S, et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature. 2012;485(7397):201-206. [10] MEYER KD, SALETORE Y, ZUMBO P, et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell. 2012;149(7):1635-1646. [11] ZHENG G, DAHL J A, NIU Y, et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol Cell. 2013;49(1):18-29. [12] KUDOU K, KOMATSU T, NOGAMI J, et al. The requirement of Mettl3-promoted MyoD mRNA maintenance in proliferative myoblasts for skeletal muscle differentiation. Open Biol. 2017;7(9): 170119. [13] ZHANG X, YAO Y, HAN J, et al. Longitudinal epitranscriptome profiling reveals the crucial role of N6-methyladenosine methylation in porcine prenatal skeletal muscle development. J Genet Genomics. 2020;47(8):466-476. [14] WU Z, LU M, LIU D, et al. m6A epitranscriptomic regulation of tissue homeostasis during primate aging. Nat Aging. 2023;3(6):705-721. [15] GILBERT CJ, RABOLLI CP, GOLUBEVA VA, et al. YTHDF2 governs muscle size through a targeted modulation of proteostasis. Nat Commun. 2024; 15(1):2176. [16] JIANG X, LIU B, NIE Z, et al. The role of m6A modification in the biological functions and diseases. Signal Transduct Target Ther. 2021; 6(1):74. [17] ZHAO Y, SHI Y, SHEN H, et al. m6A-binding proteins: The emerging crucial performers in epigenetics. J Hematol Oncol. 2020;13(1):35. [18] WENG H, HUANG F, YU Z, et al. The m6A reader IGF2BP2 regulates glutamine metabolism and represents a therapeutic target in acute myeloid leukemia. Cancer Cell. 2022;40(12):1566-1582.e10. [19] YIN J, DING F, CHENG Z, et al. METTL3-mediated m6A modification of LINC00839 maintains glioma stem cells and radiation resistance by activating wnt/β-catenin signaling. Cell Death Dis. 2023;14(7):417. [20] CHEN A, ZHANG VX, ZHANG Q, et al. Targeting the oncogenic m6A demethylase FTO suppresses tumourigenesis and potentiates immune response in hepatocellular carcinoma. Gut. 2025; 74(1):90-102. [21] LI X, YANG Y, LI Z, et al. Deficiency of WTAP in islet beta cells results in beta cell failure and diabetes in mice. Diabetologia. 2023;66(6):1084-1096. [22] YIN H, JU Z, ZHENG M, et al. Loss of the m6A methyltransferase METTL3 in monocyte-derived macrophages ameliorates alzheimer’s disease pathology in mice. PLoS Biol. 2023;21(3):e3002017. [23] QIN Y, LI L, LUO E, et al. Role of m6A RNA methylation in cardiovascular disease (review). Int J Mol Med. 2020;46(6):1958-1972. [24] YANG Y, HSU PJ, CHEN YS, et al. Dynamic transcriptomic m6A decoration: Writers, erasers, readers and functions in RNA metabolism. Cell Res. 2018;28(6):616-624. [25] PING XL, SUN BF, WANG L, et al. Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Res. 2014;24(2):177-189. [26] FU Y, JIA G, PANG X, et al. FTO-mediated formation of N6-hydroxymethyladenosine and N6-formyladenosine in mammalian RNA. Nat Commun. 2013;4(1):1798. [27] REICHEL M, KÖSTER T, STAIGER D. Marking RNA: m6A writers, readers, and functions in arabidopsis. J Mol Cell Biol. 2019;11(10):899-910. [28] SHI H, WANG X, LU Z, et al. YTHDF3 facilitates translation and decay of N6-methyladenosine-modified RNA. Cell Res. 2017;27(3):315-328. [29] WANG X, LU Z, GOMEZ A, et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature. 2014;505(7481): 117-120. [30] XIAO W, ADHIKARI S, DAHAL U, et al. Nuclear m 6 a reader YTHDC1 regulates mRNA splicing. Mol Cell. 2016;61(4):507-519. [31] ROUNDTREE IA, LUO GZ, ZHANG Z, et al. YTHDC1 mediates nuclear export of N6-methyladenosine methylated mRNAs. eLife. 2017;6:e31311. [32] HSU PJ, ZHU Y, MA H, et al. Ythdc2 is an N6-methyladenosine binding protein that regulates mammalian spermatogenesis. Cell Res. 2017; 27(9):1115-1127. [33] HUANG H, WENG H, SUN W, et al. Recognition of RNA N6-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat Cell Biol. 2018;20(3):285-295. [34] ALARCÓN CR, GOODARZI H, LEE H, et al. HNRNPA2B1 is a mediator of m6A-dependent nuclear RNA processing events. Cell. 2015;162(6): 1299-1308. [35] CHEN JJ, LU TZ, WANG T, et al. The m6A reader HNRNPC promotes glioma progression by enhancing the stability of IRAK1 mRNA through the MAPK pathway. Cell Death Dis. 2024;15(6):390. [36] WANG N, SHI B, DING L, et al. FMRP protects breast cancer cells from ferroptosis by promoting SLC7A11 alternative splicing through interacting with hnRNPM. Redox Biol. 2024;77:103382. [37] LAI Y, RAMÍREZ-PARDO I, ISERN J, et al. Multimodal cell atlas of the ageing human skeletal muscle. Nature. 2024;629(8010):154-164. [38] NEDERVEEN JP, FORTINO SA, BAKER JM, et al. Consistent expression pattern of myogenic regulatory factors in whole muscle and isolated human muscle satellite cells after eccentric contractions in humans. J Appl Physiol. 2019; 127(5):1419-1426. [39] GHELLER BJ, BLUM JE, FONG EHH, et al. A defined N6-methyladenosine (m6A) profile conferred by METTL3 regulates muscle stem cell/myoblast state transitions. Cell Death Discov. 2020;6(1):95. [40] LIANG Y, HAN H, XIONG Q, et al. METTL3-mediated m6A methylation regulates muscle stem cells and muscle regeneration by notch signaling pathway. Stem Cells Int. 2021;2021:1-13. [41] XIE SJ, LEI H, YANG B, et al. Dynamic m6A mRNA methylation reveals the role of METTL3/14-m6A-MNK2-ERK signaling axis in skeletal muscle differentiation and regeneration. Front Cell Dev Biol. 2021;9:744171. [42] LI R, CAO Y, WU W, et al. Inhibitor of FTO, rhein, restrains the differentiation of myoblasts and delays skeletal muscle regeneration. Animals. 2024;14(16):2434. [43] YANG X, MEI C, MA X, et al. m6A methylases regulate myoblast proliferation, apoptosis and differentiation. Animals. 2022;12(6):773. [44] QIAO Y, SUN Q, CHEN X, et al. Nuclear m6A reader YTHDC1 promotes muscle stem cell activation/proliferation by regulating mRNA splicing and nuclear export. eLife. 2023;12: e82703. [45] LIU J, ZUO H, WANG Z, et al. The m6A reader YTHDC1 regulates muscle stem cell proliferation via PI4K–Akt–mTOR signalling. Cell Prolif. 2023; 56(8):e13410. [46] CHEN Z, LI L, WU W, et al. Exercise protects proliferative muscle satellite cells against exhaustion via the Igfbp7-akt-mTOR axis. Theranostics. 2020;10(14):6448-6466. [47] HWANG J, KANG S, JUNG H. Effects of American wild ginseng and korean cultivated wild ginseng pharmacopuncture extracts on the regulation of C2C12 myoblasts differentiation through AMPK and PI3K/akt/mTOR signaling pathway. Mol Med Rep. 2022;25(6):192. [48] LI Y, LIU Z, YAN H, et al. Polygonatum sibiricum polysaccharide ameliorates skeletal muscle aging and mitochondrial dysfunction via PI3K/akt/mTOR signaling pathway. Phytomedicine. 2025;136:156316. [49] YEH TS, LEI TH, LIU JF, et al. Astragalus membranaceus enhances myotube hypertrophy through PI3K-mediated akt/mTOR signaling phosphorylation. Nutrients. 2022;14(8):1670. [50] ZHAO T, ZHAO R, YI X, et al. METTL3 promotes proliferation and myogenic differentiation through m6A RNA methylation/YTHDF1/2 signaling axis in myoblasts. Life Sci. 2022;298:120496. [51] HUANG X, XU C, ZHANG J, et al. Endurance exercise remodels skeletal muscle by suppressing Ythdf1-mediated myostatin expression. Cell Death Dis. 2025;16(1):96. [52] PETROSINO JM, HINGER SA, GOLUBEVA VA, et al. The m6A methyltransferase METTL3 regulates muscle maintenance and growth in mice. Nat Commun. 2022;13(1):168. [53] FENG S, ZHOU H, LIN X, et al. Exercise promotes skeletal muscle growth in adolescents via modulating Mettl3-mediated m6A methylation of MyoD in muscle satellite cells. Cell Mol Biol Lett. 2024;29(1):150. [54] FERNANDO R, DRESCHER C, NOWOTNY K, et al. Impaired proteostasis during skeletal muscle aging. Free Radic Biol Med. 2019;132:58-66. [55] ROM O, REZNICK AZ. The role of E3 ubiquitin-ligases MuRF-1 and MAFbx in loss of skeletal muscle mass. Free Radic Biol Med. 2016;98:218-230. [56] HABERECHT-MÜLLER S, KRÜGER E, FIELITZ J. Out of control: The role of the ubiquitin proteasome system in skeletal muscle during inflammation. Biomolecules. 2021;11(9):1327. [57] LAN XQ, DENG CJ, WANG QQ, et al. The role of TGF-β signaling in muscle atrophy, sarcopenia and cancer cachexia. Gen Comp Endocrinol. 2024; 353:114513. [58] SUN J, ZHOU H, CHEN Z, et al. Altered m6A RNA methylation governs denervation-induced muscle atrophy by regulating ubiquitin proteasome pathway. J Transl Med. 2023;21(1):845. [59] YOSHIDA T, DELAFONTAINE P. Mechanisms of IGF-1-mediated regulation of skeletal muscle hypertrophy and atrophy. Cells. 2020;9(9):1970. [60] LIU Y, ZHOU T, WANG Q, et al. m6 a demethylase ALKBH5 drives denervation‐induced muscle atrophy by targeting HDAC4 to activate FoxO3 signalling. J Cachexia Sarcopenia Muscle. 2022; 13(2):1210-1223. [61] DIAO LT, XIE SJ, LEI H, et al. METTL3 regulates skeletal muscle specific miRNAs at both transcriptional and post-transcriptional levels. Biochem Biophys Res Commun. 2021;552:52-58. [62] DENG K, SU Y, LIU Z, et al. Ythdf2 facilitates precursor miR-378/miR-378-5p maturation to support myogenic differentiation. Cell Mol Life Sci. 2024;81(1):445. [63] XIE SJ, TAO S, DIAO LT, et al. Characterization of long non-coding RNAs modified by m6A RNA methylation in skeletal myogenesis. Front Cell Dev Biol. 2021;9:762669. [64] LIU J, LI K, CAI J, et al. Landscape and regulation of m6A and m6Am methylome across human and mouse tissues. Mol Cell. 2020;77(2):426-440.e6. [65] YANG X, MEI C, RAZA SHA, et al. Interactive regulation of DNA demethylase gene TET1 and m6A methyltransferase gene METTL3 in myoblast differentiation. Int J Biol Macromol. 2022;223: 916-930. [66] KHARRAZ Y, GUERRA J, MANN CJ, et al. Macrophage plasticity and the role of inflammation in skeletal muscle repair. Mediators Inflamm. 2013;2013:1-9. [67] TENG Y, YI J, CHEN J, et al. N6-methyladenosine (m6A) modification in natural immune cell-mediated inflammatory diseases. J Innate Immun. 2023;15(1):804-821. |
| [1] | Pan Dong, Yang Jialing, Tian Wei, Wang Dongji, Zhu Zheng, Ma Wenchao, Liu Na, Fu Changxi. Resistance exercise activates skeletal muscle satellite cells in aged rats: role of adiponectin receptor 1 pathway [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(7): 1736-1746. |
| [2] | Cao Yong, Teng Hongliang, Tai Pengfei, Li Junda, Zhu Tengqi, Li Zhaojin. Interactions between cytokines and satellite cells in muscle regeneration [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(7): 1808-1817. |
| [3] | Hou Chaowen, Li Zhaojin, Kong Jianda, Zhang Shuli. Main physiological changes in skeletal muscle aging and the multimechanism regulatory role of exercise [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(6): 1464-1475. |
| [4] | Yang Zhijie, Zhao Rui, Yang Haolin, Li Xiaoyun, Li Yangbo, Huang Jiachun, Lin Yanping, Wan Lei, HuangHongxing. Postmenopausal osteoporosis: predictive values of muscle mass, grip strength, and appendicular skeletal muscle index [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(5): 1073-1080. |
| [5] | Zhou Jian, Zhang Tao, Zhou Weili, Zhao Xingcheng, Wang Jun, Shen Jie, Qian Li, Lu Ming. Effects of resistance training on quadriceps mass and knee joint function in patients with osteoporosis and sarcopenia [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(5): 1081-1088. |
| [6] | Huang Liuyan, Zhang Wenxi, Chen Shuwen, Yu Shimei, Dai Zhong, Zuo Changqing. Forskolin promotes C2C12 myoblast differentiation via regulating the ERK and Akt signaling pathways [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(5): 1114-1121. |
| [7] | Li Guangzheng, Li Wei, Zhang Bochun, Ding Haoqin, Zhou Zhongqi, Li Gang, Liang Xuezhen. A prediction model for sarcopenia in postmenopausal women: information analysis based on the China Health and Retirement Longitudinal Study database [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(4): 849-857. |
| [8] | Sun Jiahe, Shi Jipeng, Zhu Tianrui, Quan Helong, Xu Hongqi. Effect of exercise intervention in elderly individuals with sarcopenia and its comorbidities: a meta-analysis [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(4): 997-1007. |
| [9] | He Qiwang, , , Chen Bo, Liang Fuchao, Kang Zewei, Zhou Yuan, Ji Anxu, Tang Xialin, . Relationship between Alzheimer’s disease and sarcopenia and body mass index: analysis of GWAS datasets for European populations [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(4): 1036-1046. |
| [10] | Chen Simin, Song Rui, Peng Xing, Wang Jie, Jing Zhaohang, Parhati Nasr, Abdunabi Wupu, He Qi, Yang Lei. Correlation between skeletal muscle mass and Chinese healthy eating index in older adults [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(34): 9075-9080. |
| [11] | Yin Xingxiao, Peng Hao, Song Yanping, Yao Na, Shen Zhen, Jiang Yang, Chen Hongbo, Huang Li, Song Yueyu, Li Yanqi, Chen Qigang. Sarcopenia and cognitive impairment: a data analysis based on European population databases [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(28): 7388-7395. |
| [12] | Chu Rui, Li Mingming, Xie Yeshou, Ni Tao, Du Yinuo. Traditional Chinese sports intervene in sarcopenia and its complications in the elderly: a meta-analysis on improving muscle strength, mass, and physical function [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(24): 6354-6364. |
| [13] | Zhang Zheng, Zhang Yibo, Xu Bin, Yan Shichao, Guo Hui. Sarcopenia and non-alcoholic fatty liver disease: analysis of the gut microbiota [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(23): 6000-6009. |
| [14] | Li Qian, Li Zhenxing, Qiao Pengyan, Wang Pingzhi. Visual analysis of shear wave elastography in skeletal muscle research [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(23): 6021-6029. |
| [15] | Yin Xingxiao, Jiang Yang, Song Yanping, Yao Na, Shen Zhen, Li Yanqi, Song Yueyu, Peng Hao, Chen Qigang. Association between sarcopenia and osteoporosis: a genome-wide data analysis in European populations [J]. Chinese Journal of Tissue Engineering Research, 2026, 30(23): 6030-6039. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||