Chinese Journal of Tissue Engineering Research ›› 2021, Vol. 25 ›› Issue (10): 1592-1598.doi: 10.3969/j.issn.2095-4344.3070
Previous Articles Next Articles
Research progress of circular RNAs in osteoarthritis: from known to unknown
Xu Yiyang1, 2, Mao Guping1, Zhang Ziji1, Kang Yan1
- 1Department of Joint Surgery, First Affiliated Hospital of Sun Yat-sen University, Guangzhou 510080, Guangdong Province, China; 2Provincial School of Clinical Medicine of Fujian Medical University/Second Department of Orthopedics of Fujian Provincial Hospital, Fuzhou 350001, Fujian Province, China
-
Received:2020-03-30Revised:2020-04-03Accepted:2020-05-09Online:2021-04-08Published:2020-12-18 -
Contact:Kang Yan, MD, Professor, Chief physician, Department of Joint Surgery, First Affiliated Hospital of Sun Yat-sen University, Guangzhou 510080, Guangdong Province, China -
About author:Xu Yiyang, Doctoral candidate, Attending physician, Department of Joint Surgery, First Affiliated Hospital of Sun Yat-sen University, Guangzhou 510080, Guangdong Province, China; Provincial School of Clinical Medicine of Fujian Medical University/Second Department of Orthopedics of Fujian Provincial Hospital, Fuzhou 350001, Fujian Province, China -
Supported by:the National Natural Science Foundation of China, No. 81972051
CLC Number:
Cite this article
Xu Yiyang, Mao Guping, Zhang Ziji, Kang Yan. Research progress of circular RNAs in osteoarthritis: from known to unknown[J]. Chinese Journal of Tissue Engineering Research, 2021, 25(10): 1592-1598.
share this article
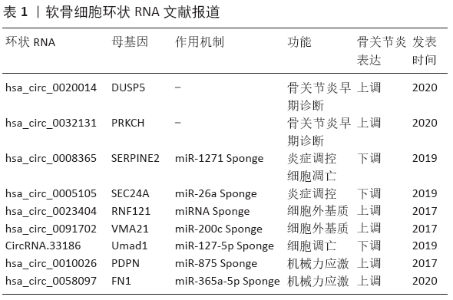
2.1 环状RNA的形成和调控 以环状RNA的来源序列进行分类,包括外显子环状RNA、内含子环状RNA以及外显子-内含子环状RNA 3种。环状结构的形成依赖特殊的反向剪接(Back-splicing),即下游剪接供体反向连接至上游剪接受体,并以共价键结合。JECK等[10]在2013年提出了2种经典的含外显子RNA环化机制:内含子配对模式(Intron-pairing)和套索模式(Lariat-driven)。而内含子环状RNA主要依赖靠近5’剪切位点的GU富集元件和3’分支位点的C富集元件互作形成类似套索结构,再由剪接体加工和3’尾端降解成环[14]。此外,QKI蛋白和盲肌样蛋白1等RNA结合蛋白也能够与侧翼内含子的特异元件结合并互相形成二聚体[15-19]。 环状RNA的生成受到顺式剪接(Cis-splicing)和反式剪接(Trans-splicing)中多种剪接因子的调控[17,20]。U2核糖核蛋白剪接体(U2 snRNP)和信使RNA前体剪接因子2等线性剪切组件的下调直接抑制了剪接体功能,进而通过可变剪接途径(Alternative Splice)促使外显子环化的发生[20-21];可稳定内含子配对结构[22];腺苷脱氨酶1则使腺苷转化成肌苷,富集在环状RNA附近位点[23],它们是环状RNA生成的负调控因子。 环状RNA的形成和调控是机体生理和病理过程的重要组成部分,无论是病因学研究,还是临床诊断、治疗手段扩展,它都极具启示作用;更加全面地剖析了环状RNA生成、调控机制,是干预其功能作用的前提,也是转录组学临床转化的“利剑”。 2.2 环状RNA的特征 ①保守性:环状RNA在不同种属间的序列有极高的保守性[10,24-25],表达量越高的环状RNA保守性更好,不仅由于基因序列的同源性,也有剪接位点的共享[26]。②普遍性:环状RNA广泛存在于生物体中,占母基因转录本的5%-10%,甚至比线性mRNA的丰度更高[27],细胞质、细胞核、血清外泌体、唾液和其他体液中都检出有相当丰度的环状RNA[11,28-30]。③稳定性:由于缺少线性RNA 3’末端的poly(A)尾,环状RNA更加耐受核糖核酸外切酶R的降解[31]。胞质中的环状RNA半衰期可超过48 h,远高于线性转录本的半衰期[10]。④组织特异性:NICOLET等[32]对骨髓造血细胞中环状RNA分布进行k-聚类分析,发现其分布具有高度规律性,并且这些环状RNA多数来自相同的基因,只是在剪接位点上存在差异。而同一环状RNA在不同组织中也可发挥不同的作用机制[33-34]。 环状RNA的生物学特征是发现、鉴定新的环状RNA以及验证其功能的主要参考。能否从某一类疾病为出发点,归纳环状RNA在其中所表现的特征,也许是未来该方向值得研究的问题,也有助于挖掘其更广泛的功能作用。 2.3 环状RNA的功能 2.3.1 ceRNA或miRNAs Sponge 环状RNA具备多种微小RNAs(miRNAs)的反应元件(MER),能够充当竞争性内源性RNA(ceRNA)或者以海绵吸附方式影响miRNA对靶基因的作用[35]。CiRS-7/CDR1as首先在神经组织中被发现能够充当miR-7的海绵,而70个结合位点也是迄今为止报道最高效的吸附能力[36]。 2.3.2 蛋白质Sponge 环状RNA与RNA结合蛋白的结合也被称为蛋白质Sponge,这类蛋白质常常与环状RNA生成、母基因转录等相关。circYAP能结合翻译起始因子4G,与多聚腺苷结合蛋白形成复合物抑制母基因YAP的翻译效率[37]。 2.3.3 蛋白质支架(Scaffold) 环状RNA与酶及其作用底物一同形成复合物,其本身并不参与催化反应,而是稳定的结合域。circAmotl1能同时结合磷酸肌醇依赖蛋白激酶1和蛋白激酶B,随后促进蛋白激酶B磷酸化并转变为激活状态而入核[38]。 2.3.4 调控转录和剪接 细胞核中,外显子-内含子环状RNA已被证实可参与调控母基因的转录。CircEI3FJ和circPAIP2共定位于细胞核中,能够在母基因启动子区域结合,而U1核糖核蛋白剪接体(U1 snRNP)也同样结合在该区域进而形成复合体,激活RNA聚合酶Ⅱ(RNA Polymerase Ⅱ),实现外显子-内含子环状RNA促进母基因的转录[39]。 2.3.5 翻译蛋白 2017年Yang等[40]首次在哺乳动物内源环状RNA中证实这一功能:环状RNA上的N6-甲基腺嘌呤(m6A)修饰位点募集YTH结构域家族蛋白3后利用翻译起始因子eIF4G2进行多肽翻译。ZHANG等[41]更是首次发现人内源性环状RNA能够以类拟病毒的重叠编码策略翻译功能蛋白。 环状RNA功能机制是是阐释其在各种生命活动中发挥作用的重要着力点,只有通过环状RNA功能的验证,才能构建更加完善的疾病相关表观遗传学网络,进而阐释和干预疾病的进程。 2.4 环状RNA与骨关节炎 国内外已有一系列的研究揭示了环状RNA在骨关节炎进展中的作用。这些报道涉及了骨关节炎发病机制的各个方面:早期骨关节炎的诊断、骨关节炎炎症反应的调控、关节软骨细胞外基质稳态、关节软骨细胞衰老凋亡以及关节软骨机械应力刺激等,见表1。 2.4.1 环状RNA与早期骨关节炎的诊断 早期骨关节炎的症状、体征并不典型,目前尚缺乏有效的诊断标志物进行快速鉴别。而血液、尿液、精液、关节液和唾液中都存在稳定、丰富的环状RNA,能够使用PCR进行快速筛查。在代谢综合征、2型糖尿病、动脉粥样硬化、阿尔茨海默症以及肿瘤等疾病患者中,也证实环状RNA可作为诊断、预后标志物[42-46]。 2020年WANG等[47]利用5对大骨节患者和骨关节炎患者的软骨样本进行微阵列分析,发现1 627个环状RNA在2组的表达有差异。将表达差异最显著的20个环状RNA进行KEGG和GO生物信息学通路分析,进一步筛选出circ_0020014_CBC1可能参与骨与软骨疾病相关通路。2组患者外周血样本中该环状RNA表达结果,也提示临床医生可能可以通过血液检查鉴别大骨节病和骨关节炎。该研究组另一篇报道中,测定5个备选环状RNA在25对膝关节骨关节炎患者和健康对照组的软骨与外周血中的表达量,发现circ_0032131_CBC1在骨关节炎组和健康对照组的软骨与外周血中表达量均有显著差异,而利用描述受试者特征曲线(ROC)分析,也获得有效的诊断敏感度(AUC=0.8455);并且在circ_0032131_CBC1=9.365CT时,Yoden指数达到最大值0.55(敏感度0.90,特异度0.65)[48]。 环状RNA独特的闭环结构、高稳定性以及组织、功能特异性都是作为早期骨关节炎诊断标志物的良好特征。 2.4.2 环状RNA参与骨关节炎炎症反应的调控 大量研究表明,骨关节炎与低度的炎症反应相关。白细胞介素1、肿瘤坏死因子α、核因子κB等多个促炎细胞因子在骨关节炎发生中起重要作用,其信号通路下游多为基质降解酶或细胞周期效应蛋白[49-50]。 利用白细胞介素1β和肿瘤坏死因子α建立动物骨关节炎模型是最经典的炎症模型。ZHOU等[51]报道了使用小鼠关节软骨细胞以不同浓度白细胞介素1β和不同时长处理,发现circRNA.33186随浓度、时间正相关的表达变化。而circ0008365/SERPINE2则是在10 μg/L 白细胞介素1β和 10 μg/L肿瘤坏死因子α分别处理24 h后都出现显著下降[52]。这提示在白细胞介素1β和肿瘤坏死因子α这2条经典炎症反应信号通路中,环状RNA都有相应的作用。 尽管骨关节炎与类风湿性关节炎的病理机制有区别,但是仍存在部分共同的炎症通路。如烟酰胺磷酸核糖基转移酶在骨关节炎和类风湿性关节炎患者外周血、关节液中都较健康人群明显升高,并且与疾病的严重程度呈正相关。这是由于关节滑膜外分泌形式烟酰胺磷酸核糖基转移酶作为促炎因素和趋化因子,抑制软骨细胞蛋白质合成、上调基质降解酶表达,而这一病理过程可能受到低氧诱导因子2α的调控[53]。2017年WU等[54]报道了hsa_circ_5105在使用白细胞介素1β刺激的软骨细胞中呈现时间依赖的上调:处理24 h后表达量增加至近5倍,这一趋势与具有潜在结合位点的miR-26a表达量相反。双荧光素酶报告基因检测验证了hsa_circ_5105与烟酰胺磷酸核糖基转移酶都能与miR-26a结合。然而,实时定量PCR提示转染hsa_circ_5105后,miR-26a的转录并未受到影响,但其对烟酰胺磷酸核糖基转移酶的抑制作用明显被拮抗;并且共转染hsa_circ_5105的小干扰RNA(siRNA)后,这种拮抗作用也被消减;进一步随着烟酰胺磷酸核糖基转移酶在软骨细胞中表达上调,促进了前列腺素E2、白细胞介素6和白细胞介素8等多种炎症因子的生成。值得注意的是,上述结果并未在hsa_circ_5105线性转录本过表达的实验中被重复,提示这一功能机制是该环状RNA所特有。 2.4.3 环状RNA与关节软骨细胞外基质稳态 软骨细胞外基质稳态能够确保关节软骨维持一定范围的承重能力,而骨关节炎软骨因蛋白聚糖的减少,无法提供足够的组织支撑。 LIU等[55]在2019年报道了对骨关节炎和正常软骨转录组学表达谱的分析,并构建存在显著差异的71个环状RNA和112 mRNA的共表达网络,其中circ0023404与基质金属蛋白酶13都具备与miR-136的结合位点;在circ0023404被敲降后,基质金属蛋白酶13表达呈现显著下调,而软骨细胞中Ⅱ型胶原则明显增加。在椎间盘髓核细胞外基质中也存在相似的研究结果,2018年CHENG等[56]发现circVMA21的转染明显减少了基质水解酶,如基质金属蛋白酶13、解聚素金属蛋白酶4、解聚素金属蛋白酶5,而增加了Ⅱ型胶原和蛋白聚糖的含量。 2.4.4 环状RNA与关节软骨细胞衰老、凋亡 关节周围组织衰老和骨关节炎微环境中累积的DNA损伤、氧化应激等改变都会软骨细胞的基因表达,并可能由于靶向DNA损伤而引起端粒耗损[50,57]。尤其是软骨细胞这类有丝分裂后细胞,长时间的复制间期使细胞损伤积累更为易感[58-59]。 细胞周期蛋白依赖性激酶抑制剂1(Cyclin-Dependent Kinase Inhibitor-1,CDKI1,也称为p21)的上调是细胞衰老最显著的变化之一,其通过抑制细胞周期蛋白依赖性激酶复合物活性调控细胞周期进程[59]。2017年DU等[60]在多个细胞系中都证实circFoxo3能够与p21、细胞周期蛋白依赖性激酶2形成三元复合物并且介导细胞周期停滞。磷脂酰肌醇3-激酶蛋白家族可作为锚定物促使蛋白激酶活化,抑制糖原合成酶激酶3β而调节细胞周期[61-62]。MI等[63]在2019年利用MC3T3-E1细胞系过表达circRNA AFF4,证实其影响下游miR-7223-5p/PI3KR1调控轴,抑制了细胞凋亡并增强增殖分化。 ZHOU等[51]在2019年报道了使用白细胞介素1β处理后的软骨细胞增殖功能受到明显抑制,CCK8试剂盒检测 A450 nm吸光度和EdU细胞成像实验均证实软骨细胞活性随处理时间延长而下降;但是转染了circRNA.33186小分子干扰RNA(siRNA)之后,软骨细胞活性有显著改善,细胞凋亡比例也随之下降,并且至少能够维持72 h。在CHENG等[52]2018年的研究中发现,转染circSERPINE2的人软骨细胞也同样部分改善了白细胞介素1β引起的细胞凋亡率增加。并且,如果将circSERPINE2和潜在靶点miR-1271一并下调,则细胞凋亡率比单独下调circSERPINE2下降更为显著。 这些证据表明环状RNA对关节软骨细胞衰老、凋亡有调控作用,骨关节炎的关节软骨细胞凋亡机制仍有待更进一步的研究。 2.4.5 环状RNA参与关节软骨的机械力应激 机械应力的增加是骨关节炎发生发展的重要因素,膝关节内、外侧间室受力不平衡造成的关节不稳可能是膝关节软骨破坏的始发因素[1]。因此,膝关节内侧半月板损伤与前交叉韧带切断常作为诱导小型动物骨关节炎的机械损伤模型[64]。 Liu等[65]分析了膝骨关节炎患者不同损伤程度软骨区域的环状RNA表达谱,有104个环状RNA在轻微损伤和严重损伤的2组间表达差异显著,其中hsa_circRNA_10026在代表受力集中的严重损伤软骨区域显著表达,在GO生物信息分析构建的共表达网络中,它与肿瘤坏死因子α都与miR-875存在潜在结合位点。进一步的体外实验利用FX-5000细胞牵拉系统模拟机械应力刺激,hsa_circRNA_10026在关节软骨的表达随着应力刺激时间延长而显著上调;并且在对软骨细胞的hsa_circRNA_10026敲减后,组织应力反应靶基因肿瘤坏死因子α表达量则随之下调。 相似的,XIAO等[66]对椎间盘终板软骨增加应力负荷后,张力敏感的circRNA0058097表达上调,而敲减该环状RNA后则应力负荷对细胞外基质的影响作用明显被削弱。miR-365a-5p同时具有circRNA0058097和组蛋白去乙酰化酶4的结合位点,而组蛋白去乙酰化酶在应力作用下对软骨细胞增殖和代谢、软骨细胞外基质降解、软骨肥大以及软骨内成骨中有广泛作用[67-68]。这些研究结果都表明环状RNA可能在软骨细胞机械力应激的调控轴中扮演重要角色。 2.5 环状RNA在骨关节炎研究的展望 显然,环状RNA从发现到逐步深入机制,必须需要经历一段积累的时间。骨关节炎相关的环状RNA研究仍处于起步阶段,无论是研究方案还是具体技术工具,都源于miRNA的研究基础。基于环状RNA的基础研究进展,对与骨关节炎相关、有研究价值但仍是未知的领域进行展望。 2.5.1 环状RNA的m6A修饰 m6A修饰在mRNA、tRNA和miRNA等多种RNA中被广泛发现,是影响其剪切、成熟、降解、核质转位、翻译蛋白以及DNA损伤的重要修饰类 型[40,69-71]。2018年MIN等[72]报道了青年与老年人群的外周血单核细胞的m6A修饰谱差异,随着衰老的进展,发生m6A修饰的转录本比例增加。而其中AGO2的丰度与甲基化程度明显相关,这种关系也影响了衰老相关疾病miRNA的表达。LI等[73]利用GSE55235和GSE55457两个GEO数据集的差异甲基化表达基因进行生物信息学分析,发现骨关节炎患者与对照组在炎症信号通路、细胞凋亡、肿瘤坏死因子α信号通路以及成骨分化等多条信号通路上存在较高的甲基化差异水平。这些研究都表明m6A甲基化修饰可能在骨关节炎这类衰老疾病的病理进程中有重要作用。 在软骨发育方面,TENG等[74]通过条件性敲除METTL-3基因的小鼠,发现METTEL-3的缺失降低了骨量,增加了骨髓脂肪。进一步的体外实验表明METTL-3的缺失使m6A甲基化调控甲状旁腺激素基因表达的功能下调,导致间充质干细胞成骨能力下降,成脂分化潜能增强。而实验结果也提示成软骨能力有所下降但并不显著,这也提示了一个可能影响间充质干细胞成软骨分化的研究方向。 环状RNA的m6A修饰在2016年被首次报道,随后的研究表明m6A甲基化修饰可促进环状RNA的蛋白翻译,并且与其出核、降解、免疫等功能状态密切相关[75-77]。到目前为止,仍未见与软骨发育、退变以及骨关节炎发生机制相关环状RNA甲基化修饰的研究报道。鉴于甲基化修饰在衰老性疾病、骨关节炎病理过程中均有体现,而m6A修饰作为研究技术最成熟的甲基化修饰,可能成为未来关于骨关节炎的环状RNA研究的一个热点。 2.5.2 环状RNA的蛋白翻译功能 目前,与骨关节炎相关的环状RNA研究均只涉及miRNA Sponge机制,其他如蛋白互作在肿瘤、心血管疾病中已有少量报道[41,78]。这其中的原因,一方面是Sponge机制的研究技术较成熟,对实验条件的要求较低,并且前期已有大量疾病相关的miRNA被报道,公开的数据库和上、下游靶点预测工具数量众多;而另一方面,环状RNA与蛋白互作的技术方案仍处于起步阶段,当前采用的验证方案依然是抗原-抗体互作、凝胶电泳等传统技术,如RNA pull-down、RNA结合蛋白免疫沉淀技术和电泳迁移率实验等。这些技术无法直接验证二者结合,也未考虑蛋白质或者多肽的三维构象改变[79]。 GARIKIPATI等[78]发现心肌梗死小鼠心脏中circFndc3b的表达显著下调,在对其过表达后能够通过增加RNA结合蛋白与FUS相结合,上调血管内皮生长因子A表达,进而重建梗死心肌、改善心脏功能。这提示研究者骨关节炎相关环状RNA蛋白互作的功能研究同样值得投入关注,可能有助于找到更为直接有效的治疗途径。 2.5.3 环状RNA在间充质干细胞成软骨分化的研究 得益于其获取和培养的方法简便易行、方案成熟,将间充质干细胞应用于骨关节炎、心血管疾病和糖尿病的治疗是近年来干细胞研究的热点,尤其以骨髓间充质干细胞和脂肪干细胞成果显著。相对而言,脂肪干细胞在抗衰老、维持干性特征、高增殖率以及低异质性等方面具有更良好的治疗前景[80]。 环状RNA对干细胞发育潜能调控的机制逐步被揭示,可变剪接和miRNA/RNA结合蛋白 Sponge在其中发挥重要作用。ZHU等[81]报道了hsa_circH19能够与多聚嘧啶序列结合蛋白1直接结合并抑制其功能,下调了甾醇调节元件结合蛋白1的前体剪切,使细胞核(nSREBP1)转位受阻,最终抑制了脂肪干细胞成脂分化。反之,敲除hsa_circH19则可显著增强脂肪干细胞成脂分化能力,可能依此实现靶向治疗脂代谢异常疾病。CHERUBINI等[82]研究证实,circFOXP1通过激活非经典的Wnt信号通路和表皮生长因子受体能够维持骨髓间充质干细胞多能性和分化能力。 目前尚缺乏环状RNA在间充质干细胞成软骨分化过程中作用的相关研究证据,尽管成软骨分化过程已有多种miRNA显示交错复杂的调控机制[13]。是否能够找到决定性作用的环状RNA参与其中,并阐明其功能机制必将是下一阶段的研究热点,这也能够给骨关节炎的软骨组织工程修复提供良好的种子细胞。 2.5.4 环状RNA作为骨关节炎的潜在治疗靶点 骨关节炎最主要的生物治疗策略是针对软骨退变的机械或炎性信号通路进行干预。关节软骨依靠良好的细胞状态和机械应激而维持关节稳态,衰老或损伤的软骨细胞功能则无法提供足够的组织支撑[2]。环状RNA在软骨退变中的机制尚未完全被阐明,但是利用其调控退变进程,改善软骨细胞外基质组分的实验现象值得特别关注。 ZHOU等[51]建立了内侧半月板损伤模型小鼠,从术后第1周开始以包装circRNA.33186的siRNA慢病毒株进行1次/周的关节腔注射,3周后实验组小鼠膝关节软骨细胞外基质降解酶比空白对照组明显降低,而Ⅱ型胶原含量却显著升高。而SHEN等[52]则使用腺病毒包被后的circ0008365/SEPINE2注射到前交叉韧带切断诱导骨关节炎模型兔的膝关节腔,治疗组最终获得的国际骨关节炎协会评分显著低于对照组,而通过Micro-CT三维成像也提示治疗组的骨赘减少。这些实验现象表明环状RNA有机会成为骨关节炎治疗的潜在靶点。理论上,环状RNA在种群进化中序列保守性高、环状结构稳定性好、组织特异性高,能够克服传统非编码RNA干扰(ncRNAi)中运输工具高要求、脱靶效应频繁的难点,相较于miRNA和长链非编码RNA更利于研究人员将其转化为临床治疗工具。"
| [1] HUNTER DJ, BIERMA-ZEINSTRA S. Osteoarthritis. Lancet. 2019; 393 (10182):1745-1759. [2] DIDOMENICO CD, LINTZ M, BONASSAR LJ. Molecular transport in articular cartilage - what have we learned from the past 50 years? Nat Rev Rheumatol. 2018;14(7):393-403. [3] ZHU Z, LI J, RUAN G, et al. Investigational drugs for the treatment of osteoarthritis, an update on recent developments. Exp Opin Invest Drugs. 2018;27(11):881-900. [4] WANG KC, CHANG HY. Epigenomics: Technologies and Applications. Circulation Res.2018;122(9):1191-1199. [5] RAMOS YF, MEULENBELT I. The role of epigenetics in osteoarthritis: current perspective. Curr Opin Rheumatol. 2017; 29(1):119-129. [6] NIGRO JM, CHO KR, FEARON ER, et al. Scrambled exons. Cell. 1991; 64(3):607-613. [7] COCQUERELLE C, MASCREZ B, HÉTUIN D, et al. Mis-splicing yields circular RNA molecules. FASEB J.1993;7(1):155-160. [8] HSU MT, COCA-PRADOS M. Electron microscopic evidence for the circular form of RNA in the cytoplasm of eukaryotic cells. Nature. 1979; 280(5720): 339-340. [9] WANG PL, BAO Y, YEE MC, et al. Circular RNA is expressed across the eukaryotic tree of life. PloS One. 2014;9(6):e90859. [10] JECK WR, SORRENTINO JA, WANG K, et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA (New York, N.Y.). 2013;19(2): 141-157. [11] SALZMAN J, GAWAD C, WANG PL, et al. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PloS One. 2012; 7(2):e30733. [12] SALZMAN J, CHEN RE, OLSEN MN, et al. Cell-type specific features of circular RNA expression. PLoS Genet. 2013; 9(9):e1003777. [13] REYNARD LN, BARTER MJ. Osteoarthritis year in review 2019: genetics, genomics and epigenetics. Osteoarthritis Cartilage. 2020;28(3): 275-284. [14] ZHANG XO, DONG R, ZHANG Y, et al. Diverse alternative back-splicing and alternative splicing landscape of circular RNAs. Genome Res. 2016; 26(9):1277-1287. [15] CONN SJ, PILLMAN KA, TOUBIA J, et al. The RNA binding protein quaking regulates formation of circRNAs. Cell. 2015;160(6):1125-1134. [16] ERRICHELLI L, DINI MODIGLIANI S, LANEVE P, et al. FUS affects circular RNA expression in murine embryonic stem cell-derived motor neurons. Nat Commun. 2017;8:14741. [17] ASHWAL-FLUSS R, MEYER M, PAMUDURTI NR, et al. circRNA biogenesis competes with pre-mRNA splicing. Mol Cell. 2014;56(1):55-66. [18] PAGLIARINI V, JOLLY A, BIELLI P, et al. Sam68 binds Alu-rich introns in SMN and promotes pre-mRNA circularization. Nucl Acids Res. 2020; 48(2):633-645. [19] YU CY, LI TC, WU YY, et al. The circular RNA circBIRC6 participates in the molecular circuitry controlling human pluripotency. Nat Commun. 2017;8(1):1149. [20] LIANG D, TATOMER DC, LUO Z, et al. The output of protein-coding genes shifts to circular rnas when the pre-mrna processing machinery is limiting. Mol Cell. 2017;68(5):940-954.e3. [21] KRAMER MC, LIANG D, TATOMER DC, et al. Combinatorial control of Drosophila circular RNA expression by intronic repeats, hnRNPs, and SR proteins. Genes Dev. 2015;29(20):2168-2182. [22] LI X, LIU CX, XUE W, et al. Coordinated circRNA Biogenesis and Function with NF90/NF110 in Viral Infection. Mol Cell. 2017;67(2): 214-227.e7. [23] EISENBERG E, LEVANON EY. A-to-I RNA editing - immune protector and transcriptome diversifier. Nature reviews. Genetics. 2018;19(8): 473-490. [24] GLAZAR P, PAPAVASILEIOU P, RAJEWSKY N. circBase: a database for circular RNAs. RNA. 2014;20(11):1666-1670. [25] YOU X, VLATKOVIC I, BABIC A, et al. Neural circular RNAs are derived from synaptic genes and regulated by development and plasticity. Nat Neurosci. 2015;18(4):603-610. [26] RYBAK-WOLF A, STOTTMEISTER C, GLAŽAR P, et al. Circular RNAs in the Mammalian Brain Are Highly Abundant, Conserved, and Dynamically Expressed. Mol Cell. 2015;58(5):870-885. [27] DANAN M, SCHWARTZ S, EDELHEIT S, et al. Transcriptome-wide discovery of circular RNAs in Archaea. Nucleic Acids Res. 2012;40(7): 3131-3142. [28] LI Y, ZHENG Q, BAO C, et al. Circular RNA is enriched and stable in exosomes: a promising biomarker for cancer diagnosis. Cell Res. 2015; 25(8): 981-984. [29] BAHN JH, ZHANG Q, LI F, et al. The landscape of microRNA, Piwi-interacting RNA, and circular RNA in human saliva. Clin Chem. 2015; 61(1):221-230. [30] YU F, XIE C, SUN J, et al. Circular RNA expression profiles in synovial fluid: a promising new class of diagnostic biomarkers for osteoarthritis. Int J Clin Exp Pathol. 2018;11(3):1338-1346. [31] SUZUKI H, ZUO Y, WANG J, et al. Characterization of RNase R-digested cellular RNA source that consists of lariat and circular RNAs from pre-mRNA splicing. Nucleic Acids Res. 2006;34(8): e63. [32] NICOLET BP, ENGELS S, AGLIALORO F, et al. Circular RNA expression in human hematopoietic cells is widespread and cell-type specific. Nucleic Acids Res. 2018;46(16):8168-8180. [33] OU R, LV J, ZHANG Q, et al. circAMOTL1 Motivates AMOTL1 Expression to Facilitate Cervical Cancer Growth. Molecular therapy. Nucleic Acids. 2020;19:50-60. [34] YANG Q, DU WW, WU N, et al. A circular RNA promotes tumorigenesis by inducing c-myc nuclear translocation. Cell Death Differ. 2017;24(9): 1609-1620. [35] TAY Y, RINN J, PANDOLFI PP. The multilayered complexity of ceRNA crosstalk and competition. Nature. 2014;505(7483):344-352. [36] KRISTENSEN LS, OKHOLM TLH, VENØ MT, et al. Circular RNAs are abundantly expressed and upregulated during human epidermal stem cell differentiation. RNA Biol. 2018;15(2):280-291. [37] WU N, YUAN Z, DU KY, et al. Translation of yes-associated protein (YAP) was antagonized by its circular RNA via suppressing the assembly of the translation initiation machinery. Cell Death Differ. 2019;26(12): 2758-2773. [38] ZENG Y, DU WW, WU Y, et al. A Circular RNA Binds To and Activates AKT Phosphorylation and Nuclear Localization Reducing Apoptosis and Enhancing Cardiac Repair. Theranostics. 2017;7(16):3842-3855. [39] LI Z, HUANG C, BAO C, et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol. 2015; 22(3): 256-264. [40] YANG Y, FAN X, MAO M, et al. Extensive translation of circular RNAs driven by N(6)-methyladenosine. Cell Res. 2017; 27(5): 626-641. [41] ZHANG M, HUANG N, YANG X, et al. A novel protein encoded by the circular form of the SHPRH gene suppresses glioma tumorigenesis. Oncogene. 2018; 37(13): 1805-1814. [42] HAQUE S, HARRIES LW. Circular RNAs (circRNAs) in health and disease. Genes (Basel). 2017;8(12). pii: E353. [43] HADDAD G, LORENZEN JM. Biogenesis and Function of Circular RNAs in Health and in Disease. Front Pharmacol. 2019;10:428. [44] KRISTENSEN LS, HANSEN TB, VENO MT, et al. Circular RNAs in cancer: opportunities and challenges in the field. Oncogene. 2018; 37(5): 555-565. [45] ALTESHA MA, NI T, KHAN A, et al. Circular RNA in cardiovascular disease. J Cell Physiol. 2019; 234(5): 5588-5600. [46] LU S, YANG X, WANG C, et al. Current status and potential role of circular RNAs in neurological disorders. J Neurochem. 2019;150(3): 237-248. [47] WANG Y, WU C, ZHANG Y, et al. Screening for differentially expressed circRNA between Kashin-Beck disease and osteoarthritis patients based on circRNA chips. Clin Chim Acta. 2020;501:92-101. [48] WANG Y, WU C, YANG Y, et al. Preliminary Exploration of hsa_circ_0032131 Levels in Peripheral Blood as a Potential Diagnostic Biomarker of Osteoarthritis. Genet Test Mol Biomarkers. 2019;23(10): 717-721. [49] SCANZELLO CR. Role of low-grade inflammation in osteoarthritis. Curr Opin Rheumatol. 2017;29(1):79-85. [50] GREENE MA, LOESER RF. Aging-related inflammation in osteoarthritis. Osteoarthritis Cartilage. 2015;23(11):1966-1971. [51] ZHOU ZB, HUANG GX, FU Q, et al. circRNA.33186 Contributes to the Pathogenesis of Osteoarthritis by Sponging miR-127-5p. Mol Ther. 2019;27(3):531-541. [52] SHEN S, WU Y, CHEN J, et al. CircSERPINE2 protects against osteoarthritis by targeting miR-1271 and ETS-related gene. Ann Rheum Dis. 2019;78(6):826-836. [53] STEGEN S, LAPERRE K, EELEN G, et al. HIF-1α metabolically controls collagen synthesis and modification in chondrocytes. Nature. 2019; 565(7740): 511-515. [54] WU Y, ZHANG Y, ZHANG Y, et al. CircRNA hsa_circ_0005105 upregulates NAMPT expression and promotes chondrocyte extracellular matrix degradation by sponging miR-26a. Cell Biol Int. 2017; 41(12): 1283-1289. [55] LIU Q, ZHANG X, HU X, et al. Circular RNA Related to the Chondrocyte ECM Regulates MMP13 Expression by Functioning as a MiR-136 ‘Sponge’ in Human Cartilage Degradation. Sci Rep. 2016; 6:22572. [56] CHENG X, ZHANG L, ZHANG K, et al. Circular RNA VMA21 protects against intervertebral disc degeneration through targeting miR-200c and X linked inhibitor-of-apoptosis protein. Ann Rheum Dis. 2018; 77(5):770-779. [57] HARBO M, DELAISSE JM, KJAERSGAARD-ANDERSEN P, et al. The relationship between ultra-short telomeres, aging of articular cartilage and the development of human hip osteoarthritis. Mech Ageing Dev. 2013; 134(9): 367-372. [58] LIU L, RANDO TA. Manifestations and mechanisms of stem cell aging. J Cell Biol. 2011; 193(2): 257-266. [59] JURK D, WANG C, MIWA S, et al. Postmitotic neurons develop a p21-dependent senescence-like phenotype driven by a DNA damage response. Aging Cell. 2012;11(6):996-1004. [60] DU WW, FANG L, YANG W, et al. Induction of tumor apoptosis through a circular RNA enhancing Foxo3 activity. Cell Death Differ. 2017; 24(2): 357-370. [61] LI T, WANG G. Computer-aided targeting of the PI3K/Akt/mTOR pathway: toxicity reduction and therapeutic opportunities. Int J Mol Sci. 2014;15(10): 18856-18891. [62] JERE SW, HOURELD NN, ABRAHAMSE H. Role of the PI3K/AKT (mTOR and GSK3β) signalling pathway and photobiomodulation in diabetic wound healing. Cytokine Growth Factor Rev. 2019;50:52-59. [63] MI B, XIONG Y, CHEN L, et al. CircRNA AFF4 promotes osteoblast cells proliferation and inhibits apoptosis via the Mir-7223-5p/PIK3R1 axis. Aging. 2019; 11(24):11988-12001. [64] THYSEN S, LUYTEN FP, LORIES RJU. Targets, models and challenges in osteoarthritis research. Dis Model Mech. 2015;8(1):17-30. [65] LIU Q, ZHANG X, HU X, et al. Emerging Roles of circRNA Related to the Mechanical Stress in Human Cartilage Degradation of Osteoarthritis. Mol Ther Nucleic Acids. 2017;7:223-230. [66] XIAO L, DING B, XU S, et al. circRNA_0058097 promotes tension-induced degeneration of endplate chondrocytes by regulating HDAC4 expression through sponge adsorption of miR-365a-5p. J Cell Biochem, 2020;121(1):418-429. [67] NISHIMORI S, LAI F, SHIRAISHI M, et al. PTHrP targets HDAC4 and HDAC5 to repress chondrocyte hypertrophy. JCI Insight. 2019;4(5). pii: 97903. [68] WANG H, ZHANG H, SUN Q, et al. Chondrocyte mTORC1 activation stimulates miR-483-5p via HDAC4 in osteoarthritis progression. J Cell Physiol. 2019;234(3):2730-2740. [69] XIAO W, ADHIKARI S, DAHAL U, et al. Nuclear m(6)A Reader YTHDC1 Regulates mRNA Splicing. Mol Cell. 2016;61(4):507-519. [70] COOTS RA, LIU XM, MAO Y, et al. m6A Facilitates eIF4F-Independent mRNA Translation. Mol Cell. 2017;68(3):504-514.e7. [71] XIAO CL, ZHU S, HE M, et al. N-Methyladenine DNA Modification in the Human Genome. Mol Cell. 2018;71(2):306-318.e7. [72] MIN KW, ZEALY RW, DAVILA S, et al. Profiling of m6A RNA modifications identified an age-associated regulation of AGO2 mRNA stability. Aging Cell. 2018;17(3):e12753. [73] LI Z, ZHANG R, YANG X, et al. Analysis of gene expression and methylation datasets identified ADAMTS9, FKBP5, and PFKBF3 as biomarkers for osteoarthritis. J Cell Physiol. 2019;234(6):8908-8917. [74] TENG H, MAO F, LIANG J, et al. Transcriptomic signature associated with carcinogenesis and aggressiveness of papillary thyroid carcinoma. Theranostics. 2018;8(16):4345-4358. [75] YANG Y, FAN X, MAO M, et al. Extensive translation of circular RNAs driven by N-methyladenosine. Cell research. 2017;27(5):626-641. [76] CHEN YG, CHEN R, AHMAD S, et al. N6-Methyladenosine Modification Controls Circular RNA Immunity. Mol Cell. 2019;76(1):96-109.e9. [77] CHEN RX, CHEN X, XIA LP, et al. N-methyladenosine modification of circNSUN2 facilitates cytoplasmic export and stabilizes HMGA2 to promote colorectal liver metastasis. Nat Commun. 2019;10(1):4695. [78] GARIKIPATI VNS, VERMA SK, CHENG Z, et al. Circular RNA CircFndc3b modulates cardiac repair after myocardial infarction via FUS/VEGF-A axis. Nat Commun. 2019;10(1):4317. [79] HUANG A, ZHENG H, WU Z, et al. Circular RNA-protein interactions: functions, mechanisms, and identification. Theranostics. 2020;10(8): 3503-3517. [80] ZHOU W, LIN J, ZHAO K, et al. Single-cell profiles and clinically useful properties of human mesenchymal stem cells of adipose and bone marrow origin. Am J Sports Med. 2019;47(7):1722-1733. [81] ZHU Y, GUI W, LIN X, et al. Knock-down of circular RNA H19 induces human adipose-derived stem cells adipogenic differentiation via a mechanism involving the polypyrimidine tract-binding protein 1. Exp Cell Res. 2020;387(2):111753. [82] CHERUBINI A, BARILANI M, ROSSI RL, et al. FOXP1 circular RNA sustains mesenchymal stem cell identity via microRNA inhibition. Nucleic Acids Res. 2019;47(10):5325-5340. |
| [1] | Pu Rui, Chen Ziyang, Yuan Lingyan. Characteristics and effects of exosomes from different cell sources in cardioprotection [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(在线): 1-. |
| [2] | Lin Qingfan, Xie Yixin, Chen Wanqing, Ye Zhenzhong, Chen Youfang. Human placenta-derived mesenchymal stem cell conditioned medium can upregulate BeWo cell viability and zonula occludens expression under hypoxia [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(在线): 4970-4975. |
| [3] | Xu Feng, Kang Hui, Wei Tanjun, Xi Jintao. Biomechanical analysis of different fixation methods of pedicle screws for thoracolumbar fracture [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(9): 1313-1317. |
| [4] | Jiang Yong, Luo Yi, Ding Yongli, Zhou Yong, Min Li, Tang Fan, Zhang Wenli, Duan Hong, Tu Chongqi. Von Mises stress on the influence of pelvic stability by precise sacral resection and clinical validation [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(9): 1318-1323. |
| [5] | Zhang Tongtong, Wang Zhonghua, Wen Jie, Song Yuxin, Liu Lin. Application of three-dimensional printing model in surgical resection and reconstruction of cervical tumor [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(9): 1335-1339. |
| [6] | Zhang Yu, Tian Shaoqi, Zeng Guobo, Hu Chuan. Risk factors for myocardial infarction following primary total joint arthroplasty [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(9): 1340-1345. |
| [7] | Li Dadi, Zhu Liang, Zheng Li, Zhao Fengchao. Correlation of total knee arthroplasty efficacy with satisfaction and personality characteristics [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(9): 1346-1350. |
| [8] | Wei Wei, Li Jian, Huang Linhai, Lan Mindong, Lu Xianwei, Huang Shaodong. Factors affecting fall fear in the first movement of elderly patients after total knee or hip arthroplasty [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(9): 1351-1355. |
| [9] | Wang Jinjun, Deng Zengfa, Liu Kang, He Zhiyong, Yu Xinping, Liang Jianji, Li Chen, Guo Zhouyang. Hemostatic effect and safety of intravenous drip of tranexamic acid combined with topical application of cocktail containing tranexamic acid in total knee arthroplasty [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(9): 1356-1361. |
| [10] | Xiao Guoqing, Liu Xuanze, Yan Yuhao, Zhong Xihong. Influencing factors of knee flexion limitation after total knee arthroplasty with posterior stabilized prostheses [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(9): 1362-1367. |
| [11] | Peng Zhihao, Feng Zongquan, Zou Yonggen, Niu Guoqing, Wu Feng. Relationship of lower limb force line and the progression of lateral compartment arthritis after unicompartmental knee arthroplasty with mobile bearing [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(9): 1368-1374. |
| [12] | Huang Zexiao, Yang Mei, Lin Shiwei, He Heyu. Correlation between the level of serum n-3 polyunsaturated fatty acids and quadriceps weakness in the early stage after total knee arthroplasty [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(9): 1375-1380. |
| [13] | Zhang Chong, Liu Zhiang, Yao Shuaihui, Gao Junsheng, Jiang Yan, Zhang Lu. Safety and effectiveness of topical application of tranexamic acid to reduce drainage of elderly femoral neck fractures after total hip arthroplasty [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(9): 1381-1386. |
| [14] | Wang Haiying, Lü Bing, Li Hui, Wang Shunyi. Posterior lumbar interbody fusion for degenerative lumbar spondylolisthesis: prediction of functional prognosis of patients based on spinopelvic parameters [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(9): 1393-1397. |
| [15] | Lü Zhen, Bai Jinzhu. A prospective study on the application of staged lumbar motion chain rehabilitation based on McKenzie’s technique after lumbar percutaneous transforaminal endoscopic discectomy [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(9): 1398-1403. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||